Assessment of Genetic Diversity and Interspecific Relationships of the Genus Viburnum Inferred from Start Codon Targeted (SCoT) Polymorphism Markers
Received: 21 December 2025 Revised: 28 January 2026 Accepted: 06 March 2026 Published: 16 March 2026
© 2026 The authors. This is an open access article under the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/).
1. Introduction
The genus Viburnum (Adoxaceae) comprises about 150–200 shrub species in the temperate and subtropical regions of the Northern Hemisphere. These are plants of ecological and ornamental importance [1]. In South Korea, 10 native Viburnum taxa have been reported, and several native Viburnum taxa are considered vulnerable [2,3,4]. Among them, V. burejaeticum has been designated and protected as an Endangered Wild Species Class II in South Korea since 2017 [5]. This situation highlights the need for robust genetic information to support conservation and management at the national and institutional levels.
Morphology-based taxonomy alone has often been insufficient to resolve relationships among closely related Viburnum taxa due to phenotypic plasticity and hybridization [6]. Therefore, a molecular approach s required to achieve more reliable identification and to clarify taxonomic boundaries [1,7,8]. Chloroplast DNA (cpDNA) and nuclear markers have been used to examine the phylogenetic and evolutionary characteristics of Viburnum, including East Asian and Korean taxa. However, detailed interspecies relationships are complex to specify, and a method for definitive discrimination of species characteristics for collection is needed [3,9,10,11,12].
Marker selection, which reduces high polymorphism and analysis time in institutions that collect and manage various plant genetic sources, is an attractive tool for identification, redundancy testing, and representative conservation fields [13,14]. Start codon targeting (SCoT) markers are key PCR-based markers that use primers designed around conserved ATG start codons to generate highly polymorphic, reproducible banding profiles without prior sequence information [15,16,17]. These markers have been applied to many crops and wild plants to evaluate genetic diversity and structure. They are simpler and more efficient than SSR markers, which require time-consuming development of species-specific primers [12,13,18]. Markers such as SCoT are handy for resolving relationships in Viburnum, as cpDNA often shows limited sequence divergence between closely related taxa [10].
This study aims to provide a SCoT-based genetic information for the Viburnum Collection at Seoul Botanic Park. This study investigates two main groups: the V. carlesii complex (V. carlesii, V. × burkwoodii, V. × carcalcephalum, V. carlesii var. botchiuense, and V. botchiuense). This complex enables detailed analysis within closely related lineages. It includes a variety of V. plicatum forms and varieties (e.g., ‘Rosacea’, ‘Lanarth’, and ‘Summer Snowflake’) that differ significantly in branch structure and inflorescence morphology. Using SCoT primers, 33 Viburnum species were analyzed with Lonicera japonica as an outgroup, providing basic data for identification, systematic management, and off-site conservation of collection plants.
2. Materials and Methods
2.1. Plant Materials and Genomic DNA Extraction
A total of 33 Viburnum taxa [19,20,21] and L. japonica (Caprifoliaceae) as the outgroup were analysed (Table 1). These 33 taxa were selected to represent the major infrageneric sections of Viburnum occurring in Korea and to span the diversity from wild native species (e.g., V. furcatum, V. wrightii) to two focal taxa of high conservation and horticultural importance, V. burejaeticum and V. setigerum, which are priority targets for ex situ conservation in Korea [16,18]. L. japonica was used as the outgroup because it belongs to the order Dipsacales, as does Viburnum, but is placed outside Adoxaceae, providing a closely related yet distinct lineage for rooting the dendrogram [16].
Table 1. The thirty-three Viburnum Taxa and outgroup (L. japonica) used in the SCoT primer analysis.
|
No. |
Species/Cultivar |
Origin |
Breeding |
History |
Source Code * |
|---|---|---|---|---|---|
|
1 |
Viburnum dilatatum ‘Daruma’ |
Horticultural selection |
unknown |
unknown |
RHS |
|
2 |
V. dilatatum ‘Michael Dodge’ |
Horticultural selection |
unknown |
unknown |
RHS |
|
3 |
V. erosum ‘Mongoloid’ |
Horticultural selection |
unknown |
unknown |
HORT |
|
4 |
V. opulus var. sargentii |
Northeast Asia |
unknown |
unknown |
IPNI |
|
5 |
V. opulus ‘Aureum’ |
Horticultural selection |
unknown |
yellow-leaved cultivar |
RHS |
|
6 |
V. opulus ‘Xanthocarpum’ |
Horticultural selection |
unknown |
yellow-fruited cultivar |
RHS |
|
7 |
V. opulus ‘Hallasan’ |
Korea |
unknown |
Korean selection |
K-HORT |
|
8 |
V. × burkwoodii |
UK |
1920s |
V. carlesii × V. utile |
TSO/RHS |
|
9 |
V. × burkwoodii ‘Mohawk’ |
Horticultural selection |
1960s |
selection from V. × burkwoodii |
RHS |
|
10 |
V. × burkwoodii ‘Conoy’ |
Horticultural selection |
unknown |
compact selection |
RHS |
|
11 |
V. × burkwoodii ‘Park Farm Hybrid’ |
Horticultural selection |
unknown |
unknown |
HORT |
|
12 |
V. carlesii Hemsl. |
Korea |
1888 |
wild species |
IPNI |
|
13 |
V. erubescens ‘Eskimo’ |
unknown |
unknown |
unknown |
UNK |
|
14 |
V. opulus f. hydrangeoides |
Japan |
unknown |
morphological form |
IPNI |
|
15 |
V. lantana ‘Variegatum’ |
Europe |
unknown |
variegated cultivar |
RHS |
|
16 |
V. lentago |
Canada |
1753 |
wild species |
IPNI |
|
17 |
V. lobophyllum Gräbn |
China |
1901 |
wild species |
IPNI |
|
18 |
V. bitchiuense Makino |
Japan |
1902 |
wild species |
IPNI |
|
19 |
V. cotinifolium D. Don |
Himalayan region |
1825 |
wild species |
IPNI |
|
20 |
V. burejaeticum |
China |
1862 |
wild species |
IPNI |
|
21 |
V. plicatum ‘Rosacea’ |
Horticultural selection |
unknown |
unknown |
HORT |
|
22 |
V. plicatum f. plicatum ‘Pink Sensation’ |
Horticultural selection |
unknown |
unknown |
HORT |
|
23 |
V. carlesii var. bitchiuense |
Korea/Japan |
unknown |
wild form |
IPNI |
|
24 |
V. × bodnantense |
UK |
1930s |
V. farreri × V. grandiflorum |
TSO/RHS |
|
25 |
V. × bodnantense ‘Dawn’ |
UK |
1935 |
selection: ‘Dawn’ |
RHS |
|
26 |
V. rhytidophyllum |
China |
1888 |
wild species |
IPNI |
|
27 |
V. rhytidophyllum ‘Variegatum’ |
Horticultural selection |
unknown |
variegated cultivar |
RHS |
|
28 |
V. macrocephalum |
China |
1847 |
wild species |
IPNI |
|
29 |
V. setigerum Hance |
China |
1882 |
wild species |
IPNI |
|
30 |
V. plicatum f. tomentosum ‘Lanarth’ |
UK |
unknown |
selection |
HORT |
|
31 |
V. plicatum f. tomentosum ‘Summer Snowflake’ |
Horticultural selection |
unknown |
unknown |
RHS |
|
32 |
V. japonicum |
Japan/Korea |
1824 |
wild species |
IPNI |
|
33 |
V. × carlcephalum |
USA |
1950s |
V. carlesii × V. macrocephalum |
TSO/RHS |
|
Out group |
L. japonica |
East Asia |
1784 |
wild species |
IPNI |
* Source codes: IPNI = International Plant Names Index; RHS = Royal Horticultural Society Plant Finder; TSO = Trees and Shrubs Online; HORT = general horticultural sources; K-HORT = Korean horticultural sources; UNK = unknown.
Plant materials were sampled from accessions preserved at Seoul Botanic Park. Genomic DNA was extracted from 0.2 g of fresh leaf material using the i-genomic Plant DNA Extraction Kit (iNtRon Biotechnology, Seongnam, Republic of Korea). DNA was diluted to 20 ng/µL for subsequent PCR reactions. The sampling design emphasizes two groups: the V. carlesii complex (including V. carlesii, V. × burkwoodii, V. × carlcephalum, V. carlesii var. bitchiuense, and V. bitchiuense), which allows fine-scale resolution of relationships within a closely related lineage, and a diverse set of V. plicatum (e.g., ‘Rosacea’, ‘Lanarth’, ‘Summer Snowflake’) that differ markedly in branching architecture and inflorescence morphology.
2.2. SCoT Marker Analysis
In this study, we initially screened 60 SCoT primers and selected 17 highly polymorphic and reproducible primers for genetic analysis (Table S1), following previously described SCoT protocols with minor modifications [15,16]. PCR amplification was performed in a total volume of 20 µL containing 20 ng genomic DNA, 1× PCR buffer, 2.5 mM MgCl2, 0.2 mM of each dNTP, 0.5 µM primer, and 1 U Taq DNA polymerase. PCR conditions were as follows: an initial denaturation at 95 °C for 5 min; followed by 35 cycles of 95 °C for 1 min, 50 °C for 1 min, and 72 °C for 2 min; and a final extension at 72 °C for 10 min.
2.3. Gel Electrophoresis and Data Analysis
The amplified products were separated on 1.5% agarose gels at 130 V for 120 min. Bands were scored in a binary format (1 = present, 0 = absent). Specifically, PCR products were separated on 1.5% agarose gels, stained with Dyne Loading Star (Dyne Bio, Seongnam, Republic of Korea), and visualized under UV light using a Gel Doc XR+ imaging system (Bio-Rad, Hercules, CA, USA). A 100 bp DNA ladder (NanoHelix, Daejeon, Republic of Korea) was used as a molecular size standard. All PCR amplifications were performed in triplicate independently to confirm the reproducibility of the banding patterns.
2.4. Statistical Analysis
Pairwise Jaccard distance coefficients were calculated from the binary matrix (Table S2) using NTSYSpc version 2.2 (Exeter Software, Setauket, NY, USA) and used to construct a UPGMA dendrogram. [22]. Principal component analysis (PCA) was performed on the Jaccard distance matrix to visualize multilocus patterns of variation.
3. Results
3.1. SCoT Marker Polymorphism
A total of 60 SCoT primers were initially screened, of which 17 highly polymorphic and reproducible primers were selected for genetic analysis, generating 489 polymorphic bands. All screened primers are listed in Table S1, and the key characteristics of the 17 selected primers are summarized in Table 2. The number of polymorphic bands per selected primer ranged from 19 (SCoT 45) to 35 (SCoT 18), with an average of 28.76 bands per primer. Among these, SCoT34 and SCoT48 showed relatively high polymorphism information content (PIC) and clear, reproducible banding patterns. They were therefore chosen as representative examples for the gel image in Figure 1.
Table 2. Sequences and polymorphism statistics of the 17 SCoT primers used for genetic analysis of Viburnum taxa.
|
No. |
Primer |
Sequence (5′-3′) |
Melting Tem. (°C) |
GC Content (%) |
Polymorphic Bands |
PIC |
|---|---|---|---|---|---|---|
|
1 |
SCoT 12 |
ACGACATGGCGACCACCG |
59.8 |
61.1 |
33 |
0.38 |
|
2 |
SCoT 13 |
ACGACATGGCGACCATCG |
59.3 |
61.1 |
31 |
0.41 |
|
3 |
SCoT 14 |
CAACAATGGCTACCACCT |
62.5 |
66.7 |
33 |
0.36 |
|
4 |
SCoT 16 |
CAACAATGGCTACCACGC |
58.1 |
61.1 |
33 |
0.35 |
|
5 |
SCoT 18 |
CAACAATGGCTACCACGT |
61.1 |
66.7 |
35 |
0.35 |
|
6 |
SCoT 34 |
ACCATGGCTACCACCGCA |
60.3 |
61.1 |
31 |
0.44 |
|
7 |
SCoT 45 |
ACAATGGCTACCACTGAC |
53.2 |
50.0 |
19 |
0.38 |
|
8 |
SCoT 47 |
ACAATGGCTACCACTGCC |
56.3 |
55.6 |
22 |
0.38 |
|
9 |
SCoT 48 |
ACAATGGCTACCACTGGC |
56.3 |
55.6 |
21 |
0.41 |
|
10 |
SCoT 50 |
ACAATGGCTACCACTGGG |
55.5 |
55.6 |
34 |
0.33 |
|
11 |
SCoT 51 |
ACAATGGCTACCACTGTC |
53.2 |
50.0 |
33 |
0.33 |
|
12 |
SCoT 52 |
ACAATGGCTACCACTGCA |
55.5 |
50.0 |
28 |
0.36 |
|
13 |
SCoT 53 |
ACAATGGCTACCACCGAC |
56.0 |
55.6 |
29 |
0.39 |
|
14 |
SCoT 54 |
ACAATGGCTACCACCAGC |
56.3 |
55.6 |
29 |
0.33 |
|
15 |
SCoT 55 |
ACAATGGCTACCACTACC |
52.5 |
50.0 |
26 |
0.37 |
|
16 |
SCoT 59 |
ACAATGGCTACCACCATC |
53.2 |
50.0 |
23 |
0.26 |
|
17 |
SCoT 60 |
ACAATGGCTACCACCACA |
55.0 |
50.0 |
27 |
0.38 |
|
Total Mean |
489 28.76 |
|
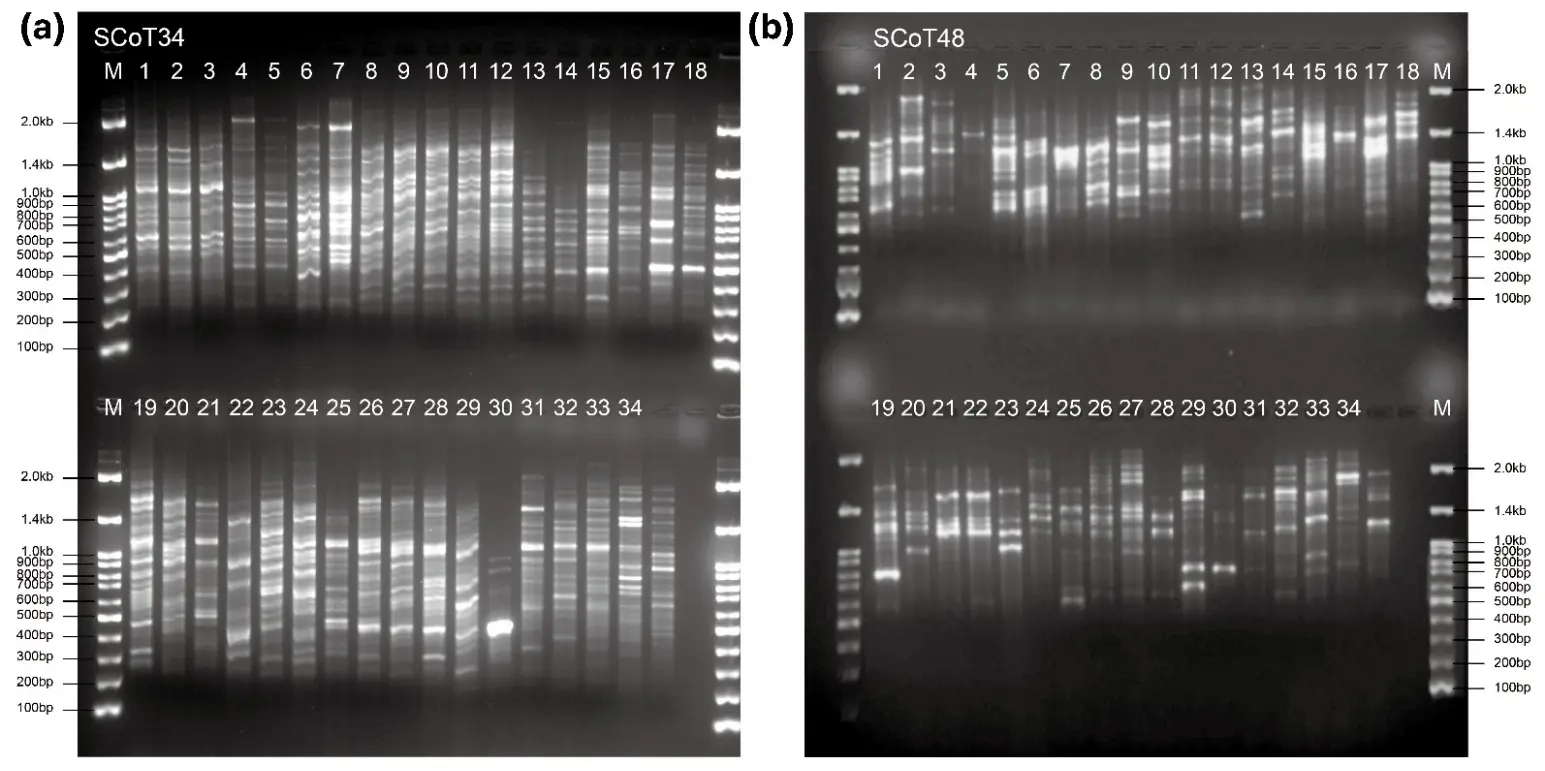
Figure 1. SCoT banding patterns of 33 Viburnum accessions and the outgroup were generated with primers SCoT 34 (a) and SCoT 48 (b). M, 100 bp DNA ladder with size markers from 100 to 2000 bp indicated on the left (a) and right (b); lanes 1–33 correspond to the Viburnum accessions listed in Table 1, lane 34 represents the outgroup, and lane 35 was not used in the analyses.
The PIC values ranged from 0.26 to 0.44, with a mean of 0.36. According to commonly used criteria, markers with PIC values between 0.25 and 0.50 are considered moderately informative, whereas values below 0.25 indicate only slightly informative markers [23]. Thus, the SCoT primers used in this study are moderately informative and provide sufficient discriminatory power for diversity and relationship analyses in this collection. The high number of polymorphic bands per primer indicates that a compact SCoT primer set can effectively capture genetic variation across the 33 Viburnum taxa.
3.2. Genetic Similarity and Phylogenetic Analysis (UPGMA)
The UPGMA dendrogram separated the Viburnum taxa from the outgroup, L. japonica (Figure 2). Cluster analysis indicated substantial genetic differentiation, with four distinct clusters being recognized at a similarity coefficient of 0.46: Cluster I contained 23 accessions, Cluster II 3, Cluster III 2, and Cluster IV 5 accessions. The largest group (Cluster I) encompassed the majority of species, whereas Clusters II–IV represented more restricted assemblages. The clustering pattern broadly corresponded to traditional infrageneric sections (Tomentosa, Odontotinus, Lentago, Euviburnum, and Opulus), although several taxa showed placements inconsistent with morphology-based classification. Pairwise Jaccard distances among the 33 Viburnum taxa (and L. japonica as outgroup) are given in Table S2.
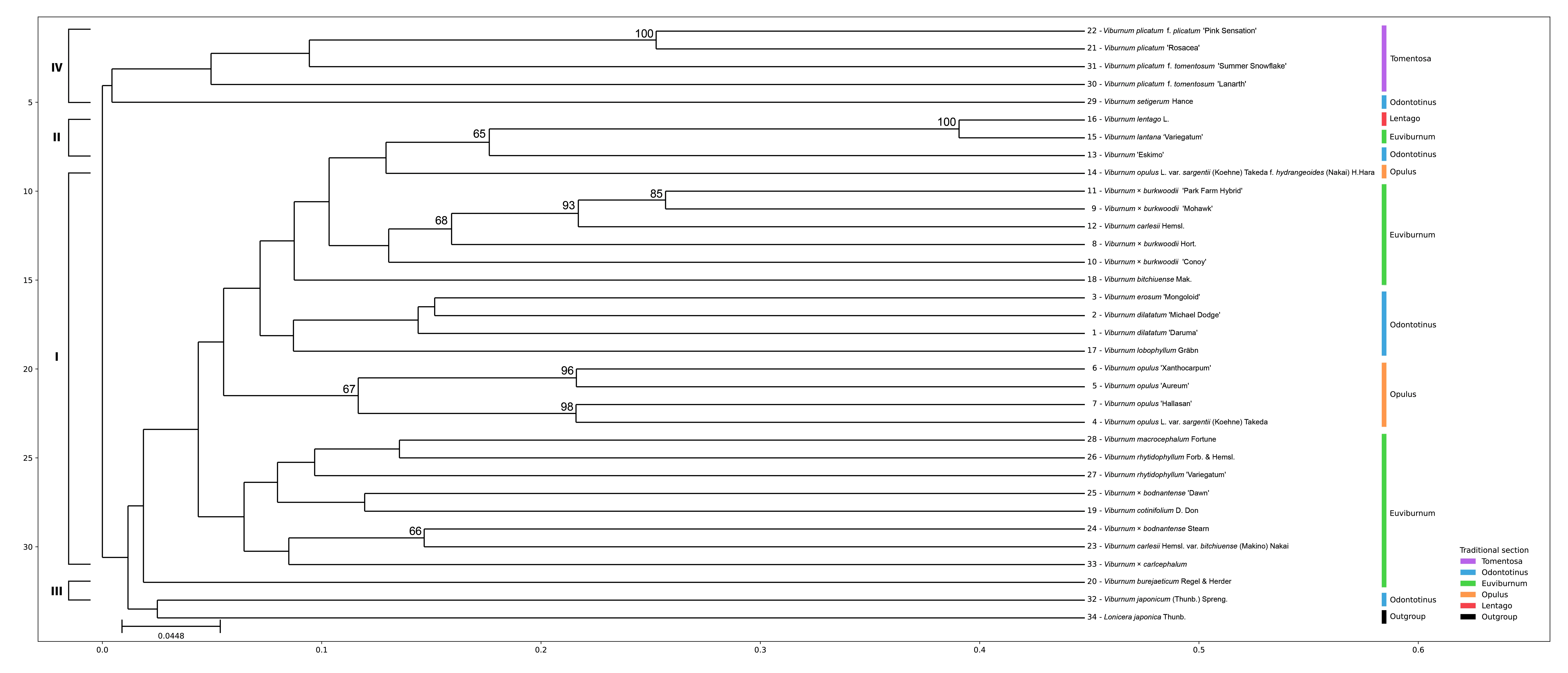
Figure 2. UPGMA phylogram inferred from SCoT primer data for 33 Viburnum taxa, with L. japonica as the outgroup. Branch lengths are proportional to Jaccard distance among taxa (scale bar). Taxon numbers correspond to those listed in Table 1, and taxa are color-coded according to traditional section classification. Numbers above branches indicate bootstrap support values (%) based on 1000 replicates (only values ≥50% are shown).
Within this structure, the V. carlesii complex (including V. carlesii, V. × burkwoodii, and V. bitchiuense) consistently formed a subgroup in Cluster I. In contrast, V. plicatum and its forms and cultivars were placed together in Cluster IV as a clearly separated group, emphasizing how divergent they are from the remaining taxa.
Several species, such as V. dilatatum, V. erosum, and V. wrightii, showed intermediate positions within Cluster I, suggesting partial admixture within this cluster. V. japonicum grouped most closely with V. burejaeticum in Cluster III.
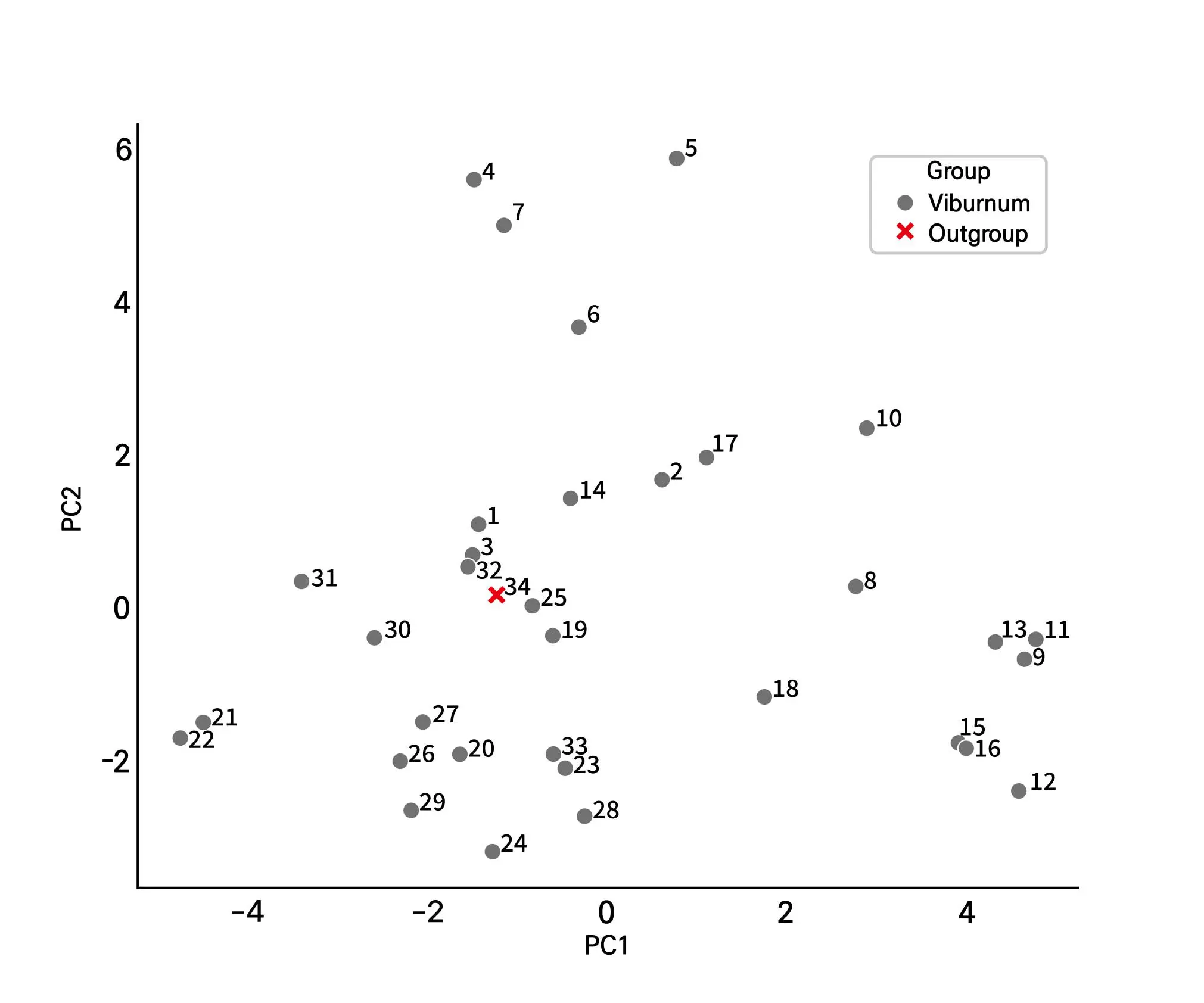
3.3. Principal Component Analysis (PCA)
The first two principal components explained a substantial proportion of the total variance (PC1: 26.5%, PC2: 15.8%), allowing a clear visualization of genetic relationships among the accessions (Figure 3). Most Viburnum taxa were widely scattered along the two PCA axes, indicating substantial genetic diversity within the genus. In contrast, members of the V. carlesii complex were clustered closely together, whereas V. plicatum occupied a more distant position. This distribution pattern is consistent with the relationships inferred from the UPGMA dendrogram.
Within the V. carlesii complex, putative hybrids such as V. × burkwoodii and V. × carlcephalum occupied intermediate positions between their parental taxa, consistent with their hybrid origin. In contrast, V. plicatum accessions were more widely distributed along the PC1 axis, indicating clear separation from the V. carlesii complex but relatively weak structure within the V. plicatum group itself. This distribution pattern was similar to the clustering pattern observed in the UPGMA dendrogram.
4. Discussion
4.1. Effectiveness of SCoT Markers
The high level of polymorphism observed in this study (489 polymorphic bands; mean PIC of 0.36) indicates that the selected SCoT primers are highly informative and reliable for assessing genetic diversity within Viburnum. These values lie at the upper end of the ranges reported for SCoT markers in crop and woody species, supporting the strong discriminating power of this marker system for germplasm characterization [15,16].
4.2. Interspecific Relationships and Taxonomic Implications
The UPGMA dendrograms and PCA arrays revealed well-structured interspecific relationship patterns that are mainly consistent with the proposed morphology and DNA-based classification of Viburnum species [1,7,9,10]. The four genetic clusters (I–IV) showed broad concordance with traditional infrageneric sections, as Cluster IV (V. plicatum group) corresponds to section Tomentosa, and Cluster II (V. lentago, V. lantana, V. erabescens) to section Lentago [10]. The tight clustering of the V. carlesii complex confirms its close taxonomic affinity. Indicates that relatively few, well-defined lineages can capture much of the genetic and morphological variation represented by these taxa, which is advantageous for targeted breeding and ex situ conservation planning [1,9,11,12,18]. By contrast, East Asian species such as V. dilatatum, V. erosum, and V. wrightii showed partially intermixed positions within the main cluster, whereas V. japonicum grouped with V. burejaeticum in Cluster III. These relationships are not entirely consistent with current morphological classifications. Together, suggesting a complex evolutionary history and possible historical hybridization among East Asian Viburnum lineages. These patterns indicate that multiple, partially diverged lineages coexist within East Asian Viburnum, and that conservation and taxonomic work in this group should explicitly account for this hidden evolutionary complexity [3,4,8,9,10].
The genetic isolation of V. plicatum (Cluster IV) is particularly noteworthy. Although morphologically classified within section Tomentosa, V. plicatum and its cultivars formed a strongly supported monophyletic group, markedly distant from all other clusters. These patterns support a unique systematic position and are consistent with the characteristic stratified horizontal branch described in taxonomic and horticultural treatments, as well as with striking sterile marginal flowers [7,10,24]. The strong genetic isolation of the V. plicatum group, together with its distinctive branching architecture and showy sterile marginal flowers, suggests a long-term divergence from other Viburnum lineages and supports its recognition as a distinct conservation unit and a valuable source of genetically divergent parents for breeding [1,10].
The close association of V. burejaeticum with V. japonicum in Cluster III likewise highlights their genetic differentiation from other East Asian taxa. It suggests that this lineage represents an evolutionarily and conservationally important component of Viburnum diversity that has so far received relatively little detailed genetic study [1,3,4,9].
4.3. Implications for Conservation and Breeding
The genetic marker information selected here may provide basic information for the discrimination, conservation, and management of plants of the genus Viburnum maintained at the Seoul Botanic Park [9,10,24]. In particular, identifying genetically distant lineages such as V. plicatum and V. burejaeticum can help minimize redundancy within the collection [1,3,4,10]. These patterns indicate that multiple, partially diverged lineages coexist within East Asian Viburnum. They also suggest that conservation and taxonomic work in this group should explicitly account for this hidden evolutionary complexity [9,25].
The substantial genetic distances observed between clusters (Table S2) provide a framework for selecting genetically diverse parental combinations in future breeding programs. For example, some of the highest pairwise Jaccard distances in Table S2 occur between members of the V. plicatum group (Cluster IV; e.g., V. plicatum f. tomentosum ‘Lanarth’, no. 30) and taxa in the V. carlesii complex (Cluster I; e.g., V. carlesii, no. 12, and V. burkwoodii, nos. 8–11), indicating that such inter-cluster combinations would maximize nuclear genetic divergence in breeding crosses. Inter-cluster crosses, particularly those involving the genetically isolated V. plicatum lineage (Cluster IV), may maximize genetic diversity in hybrid offspring [1,7,10,12]. In particular, genetically isolated lineages such as the V. plicatum group and the V. burejaeticum-V. japonicum lineage should be given high priority in conservation programmes. They can serve as strategically important parents for generating hybrids with novel combinations of ornamental and adaptation-related traits [11,12,26].
Moreover, V. burejaeticum (no. 20) shows consistently high genetic differentiation from the main East Asian cluster, reinforcing its status as a genetically unique lineage and a priority candidate for conservation-oriented breeding together with V. japonicum (no. 32) in Cluster III [3,4,9]. Distinct genetic differences between major clusters may indicate new allelic combinations that could be beneficial for resource conservation. Furthermore, crossbreeding between genetically distant taxa may facilitate the development of ornamental cultivars with novel or improved traits [12,26,27,28].
4.4. Future Perspectives
Future research should integrate these SCoT-based results with additional molecular approaches, such as ISSR, SSR, and SNP markers, which have already been developed or applied in Viburnum and provide complementary, codominant information on allelic and lineage variation [12,18,26]. Combining such multilocus molecular data with detailed morphological, anatomical, and ecological trait analyses is expected to yield a more highly resolved phylogenetic framework and help clarify remaining taxonomic uncertainties within the genus [1,7,9,10]. In addition, plastid phylogenomic work on Adoxaceae suggests that including additional outgroups from closely related genera would improve root placement and allow more rigorous tests of Viburnum monophyly in future phylogenetic analyses. In this context, Sambucus (Adoxaceae) could serve as an appropriate outgroup candidate [3].
Integrating these SCoT data with chloroplast sequences and newly developed SSR markers from the Viburnum collection at Seoul Botanic Park will enable finer-scale inference of demographic history, gene flow, and population structure, thereby strengthening both taxonomic and conservation applications in Viburnum [3,12,18,29].
5. Conclusions
This study provides the first systematic application of SCoT markers to assess genetic diversity and interspecific relationships in the genus Viburnum. Using 17 SCoT primers, we detected 489 polymorphic bands across 33 Viburnum taxa, including several endangered and vulnerable native species. We resolved four distinct genetic clusters that clearly separated Viburnum from the outgroup Lonicera japonica.
The cluster pattern well reflected the independent systematic location of V. plicatum and the close relationships in the V. carlesii complex. While revealing partially mixed groups of East Asian taxa and distinct V. japonicum and V. burejaeticum taxa in cluster III, consistent with complex evolutionary histories in these taxa. These results provide practical genetic markers that can be utilized for taxonomic identification and help refine ex-situ strategies by selectively obtaining genetically unique lineages and contributing to reducing intra-cluster duplication.
The pronounced genetic differentiation between the major clusters indicates that significant genetic variation exists that has not yet been fully utilized in ornamental breeding. Crossbreeding between genetically distant taxa may promote the development of varieties with new combinations of adaptation-related traits and horticultural/corrosive traits. Overall, the SCoT-based framework developed here will support long-term conservation planning, germplasm management, and the strategic selection of breeding parents in Viburnum. Furthermore, this study highlights the value of SCoT markers as an efficient tool for managing woody ornamental plant germplasm. It provides basic data to support long-term conservation plans for some Viburnum resources, including endangered taxa.
Supplementary Materials
The following supporting information can be found at: https://www.sciepublish.com/article/pii/910, Table S1: List of all 60 SCoT primers screened in this study and their basic characteristics; Table S2: Jaccard distance matrix calculated from the binary SCoT dataset for the 33 Viburnum accessions.
Statement of the Use of Generative AI and AI-Assisted Technologies in the Writing Process
During the preparation of this work, the authors used Grammarly and Perplexity (AI-assisted tools) to improve readability and language. After using these tools, the authors reviewed and edited the content as needed and took full responsibility for the publication’s content.
Acknowledgments
The authors thank Wan-Hee Lee, Head of the Plant Management & Research Division, and Su-Mi Park, Director of Seoul Botanic Park, for administrative support and facilitation of this project. The authors also thank the staff of Seoul Botanic Park for their assistance with field sampling and cultivation of Viburnum accessions.
Author Contributions
I.-J.C.: conceptualization, data curation, formal analysis, supervision, writing—original draft, writing—review & editing. H.-M.P.: investigation, long-term collection, and ex situ conservation of Viburnum germplasm, establishment and management of the Viburnum display garden at Seoul Botanic Park. H.-K.K.: investigation, monitoring of growth performance, and ongoing propagation and maintenance of Viburnum accessions.
Ethics Statement
Not applicable. This study did not involve humans or animals. The plant materials used in this study are not listed in the IUCN Red List, and no endangered or protected species were harmed during sample collection. All plant sampling and subsequent experiments were conducted in accordance with the guidelines and regulations of Seoul Botanic Park.
Informed Consent Statement
Not applicable. This study did not involve humans.
Data Availability Statement
Processed data supporting the findings of this study are provided in the Supplementary Materials. The raw data (including gel images and the binary SCoT marker matrix) are available from the corresponding author upon reasonable request.
Funding
This research was funded by the Seoul Metropolitan Government.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have influenced the work reported in this paper.
References
- Spriggs EL, Clement WL, Sweeney PW, Madrinán S, Edwards EJ, Donoghue MJ. Temperate radiations and dying embers of a tropical past: The diversification of Viburnum. New Phytol. 2015, 207, 340–354. DOI:10.1111/nph.13305. [Google Scholar]
- Choi YG, Oh SH. A comparative morphological study of Viburnum (Adoxaceae) in Korea. Korean J. Plant Taxon. 2019, 49, 107–117. DOI:10.11110/kjpt.2019.49.2.107. [Google Scholar]
- Ran H, Liu Y, Wu C, Cao Y. Phylogenetic and comparative analyses of complete chloroplast genomes of Chinese Viburnum and Sambucus (Adoxaceae). Plants 2020, 9, 1143. DOI:10.3390/plants9091143. [Google Scholar]
- Zhu H, Liu J, Li H, Yue C, Gao M. Complete chloroplast genome structural characterization and comparative analysis of Viburnum japonicum (Adoxaceae). Forests 2023, 14, 1819. DOI:10.3390/f14091819. [Google Scholar]
- National Institute of Biological Resources. V. burejaeticum Species Information. Available online: https://species.nibr.go.kr (accessed on 18 December 2025).
- Moeglein MK. The Roles of Polyploidy, Climate, and Genetic Architecture in the Evolution of Leaf Form in Viburnum (Adoxaceae). Ph.D. Thesis, Yale University, New Haven, CT, USA, 2021. [Google Scholar]
- Ghimire B, Son DC, Park BK, Oh SH. Comparative wood anatomy of Korean Viburnum L. (Adoxaceae) and its taxonomic implication. PhytoKeys 2020, 156, 27–46. DOI:10.3897/phytokeys.156.52031. [Google Scholar]
- Zhao L, Wang Y, Lyu W, Tang Z, Qiu L, Tang M, et al. A new synonym for Viburnum erosum (Viburnaceae) in East China, based on morphological and molecular evidence. PLoS ONE 2025, 20, e0312920. DOI:10.1371/journal.pone.0312920. [Google Scholar]
- Choi YG, Youm JW, Lim CE, Oh SH. Phylogenetic analysis of Viburnum (Adoxaceae) in Korea using DNA sequences. Korean J. Plant Taxon. 2018, 48, 206–217. DOI:10.11110/kjpt.2018.48.3.206. [Google Scholar]
- Clement WL, Arakaki M, Sweeney PW, Edwards EJ, Donoghue MJ. A chloroplast tree for Viburnum (Adoxaceae) and its implications for phylogenetic classification and character evolution. Am. J. Bot. 2014, 101, 1029–1049. DOI:10.3732/ajb.1400015. [Google Scholar]
- Dean D. Assessing the Genetic Diversity of the Genus Viburnum Using Simple Sequence Repeats. Ph.D. Thesis, University of Tennessee, Knoxville, TN, USA, 2014. [Google Scholar]
- Hamm TP, Nowicki M, Boggess SL, Ranney TG, Trigiano RN. A set of SSR markers to characterize genetic diversity in all Viburnum species. Sci. Rep. 2023, 13, 5343. DOI:10.1038/s41598-023-31878-0. [Google Scholar]
- Hamm TP, Nowicki M, Boggess SL, Klingeman WE, Hadziabdic D, Huff ML, et al. Development and characterization of 15 novel genomic SSRs for Viburnum farreri. Plants 2021, 10, 487. DOI:10.3390/plants10030487. [Google Scholar]
- Yun YE, Yu JN, Lee BY, Kwak M. An introduction to microsatellite development and analysis. Korean J. Plant Taxon. 2011, 41, 299–314. DOI:10.11110/kjpt.2011.41.4.299. [Google Scholar]
- Ahmad FK, Noori IM. Phytochemical analysis and SCoT molecular marker used to determine genetic diversity of many fig accessions distributed in various locations of Sulaymaniyah province. Kufa J. Agric. Sci. 2024, 16, 86–118. DOI:10.36077/kjas/2024/v16i4.12074. [Google Scholar]
- Altaf MT, Nadeem MA, Ali A, Liaqat W, Bedir M, Baran N, et al. Applicability of start codon targeted (SCoT) markers for the assessment of genetic diversity in bread wheat germplasm. Genet. Resour. Crop. Evol. 2025, 72, 1205–1218. DOI:10.1007/s10722-024-02016-0. [Google Scholar]
- Collard BCY, Mackill DJ. Start codon targeted (SCoT) polymorphism: A simple, novel DNA marker technique for generating gene-targeted markers in plants. Plant Mol. Biol. Rep. 2009, 27, 86–93. DOI:10.1007/s11105-008-0060-5. [Google Scholar]
- Barish S, Arakaki M, Edwards EJ, Donoghue MJ, Clement WL. Characterization of 16 microsatellite markers for the Oreinotinus clade of Viburnum (Adoxaceae). Appl. Plant Sci. 2016, 4, 1600103. DOI:10.3732/apps.1600103. [Google Scholar]
- International Plant Names Index. Available online: https://www.ipni.org (accessed on 15 February 2025).
- Korea National Arboretum Plant List. Available online: http://www.nature.go.kr (accessed on 15 February 2025).
- Trees and Shrubs Online. Available online: https://treesandshrubsonline.org (accessed on 15 February 2025).
- Rohlf FJ. NTSYSpc: Numerical Taxonomy and Multivariate Analysis System, version 2.2; Exeter Software: Setauket, NY, USA, 2009. [Google Scholar]
- Botstein D, White RL, Skolnick M, Davis RW. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am. J. Hum. Genet. 1980, 32, 314–331. [Google Scholar]
- Royal Horticultural Society Plant Finder. Available online: https://www.rhs.org.uk/plants (accessed on 15 February 2025).
- Winkworth RC, Donoghue MJ. Viburnum phylogeny based on combined molecular data: Implications for taxonomy and biogeography. Am. J. Bot. 2025, 92, 653–666. DOI:10.3732/ajb.92.4.653. [Google Scholar]
- Hasan N, Choudhary S, Naaz N, Sharma N, Laskar RA. Recent advancements in molecular marker-assisted selection and applications in plant breeding programs. J. Genet. Eng. Biotechnol. 2021, 19, 128. DOI:10.1186/s43141-021-00231-1. [Google Scholar]
- Hong JH, Shim EJ, Park WH, Soh EH. Construction of SSR profile database for variety identification of plum collected in Korea. Korean J. Breed. Sci. 2015, 47, 97–104. DOI:10.9787/KJBS.2015.47.2.97. [Google Scholar]
- Ran H, Liu Y, Wu C, Cao Y. Phylogenetic and comparative analyses of complete chloroplast genomes of Chinese Viburnum and Sambucus (Adoxaceae). Plants 2020, 9, 1143. DOI:10.3390/plants9091143. [Google Scholar]
- Le TY, Park J. The complete chloroplast genome sequence of Viburnum odoratissimum and phylogenetic relationship with other close species in the Adoxaceae family. Plant 2021, 9, 28–35. DOI:10.11648/j.plant.20210902.12. [Google Scholar]