Synthetic Biology–Driven Innovation in the Production of Cosmetic Ingredients: From Natural Mimicry to Precision Creation
Jinhang Yi
1
 Mengxi Zhang
1
Fangying Hu
1
Heyun Wu
1,2
Qian Ma
1,2,*
Xixian Xie
1,2,*
Mengxi Zhang
1
Fangying Hu
1
Heyun Wu
1,2
Qian Ma
1,2,*
Xixian Xie
1,2,*
Received: 26 November 2025 Revised: 13 January 2026 Accepted: 30 January 2026 Published: 24 February 2026
© 2026 The authors. This is an open access article under the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/).
Consumers’ threefold demand for naturalness, safety, and efficacy in cosmetic ingredients is compelling the industry to reexamine its sources of raw materials. Traditional chemical synthesis faces environmental burdens and potential toxicity debates, while plant extraction is constrained by resource scarcity and batch variability. Synthetic biology, through the programming of microbial cell factories, paves a third way for the green manufacturing of amino acid derivatives, which is neither pure chemical synthesis nor simple extraction, but a function-driven paradigm of precision creation [1,2]. We define this paradigm as operating at three integrated levels: (i) molecular precision in designing ingredients tailored to skin physiology; (ii) process precision in controlling biosynthetic pathways within microbial cell factories; and (iii) application precision, ultimately extending to personalized formulations. This multi-level framework will serve as the central thread connecting the advances in enzyme engineering, metabolic regulation, and future skincare applications discussed in this Perspective. A milestone in this paradigm shift is that we no longer settle for replicating natural molecules; instead, we can now custom-design and efficiently synthesize derivatives with optimized structures based on skin physiology. In recent years, the production titers of key molecules such as arginine [3], ergothioneine [4], and spermidine [5] have achieved orders-of-magnitude growth.
Compared to traditional approaches, synthetic biology offers distinct advantages in scalability, cost structure, and sustainability. Conventional chemical synthesis, while capable of large-scale production, often relies on petrochemical feedstocks and harsh reaction conditions, resulting in a high environmental footprint and potential safety concerns about residual reagents. Plant extraction, though perceived as “natural”, is constrained by seasonal variability, land-use intensity, and low yields, resulting in unstable supply and high costs. In contrast, synthetic biology-based biomanufacturing can utilize renewable carbon sources, operate under mild conditions, and achieve consistent, high-titer production with a lower lifecycle environmental impact. Collectively, these technological and economic advantages are driving the field’s transition from “proof-of-concept” to the “eve of industrialization”.
Yet, a significant gap persists between technological maturity and commercial success, calling for a proactive assessment of breakthrough directions and key challenges over the next five years.
1. Strategies for the Construction and Optimization of Microbial Cell Factories
Building efficient microbial cell factories for amino acid derivatives demands a systematic design paradigm that encompasses basic molecular parts, intracellular regulation, and system-level evolution. This section elaborates how three technological pillars—precision enzyme engineering, dynamic pathway control, and high throughput screening—act synergistically to overcome the bottlenecks in the biosynthesis of these compounds.
1.1. Rational Design and Optimization of Enzyme Components
The heterologous introduction of natural synthesis modules, pertinent to a vast array of amino acid derivatives and various cosmetic ingredients, into suitable microbial chassis has emerged as a pivotal approach for the construction of efficient cell factories. However, heterologous enzymes often encounter challenges in new hosts—such as low expression, misfolding, inadequate catalytic efficiency, or cofactor mismatch—which can become the bottleneck for efficient functionality [6]. Therefore, optimization of key enzymes constitutes a foundational step in building high-performance cell factories (the ‘Enzyme engineering’ module, Figure 1). Enzyme engineering strategies have evolved from traditional directed evolution, through computationally guided semi-rational design, to the current era of artificial intelligence (AI)-empowered rational design. Leveraging models like AlphaFold2 [7] and ESMFold [8] for structure prediction, along with sequence design tools such as Rosetta [9] and ProteinMPNN [10], researchers can not only de novo design enzymes with desired functions but also accurately predict key mutations that modulate substrate affinity [11], alter cofactor specificity (e.g., switching from NADPH-dependent to NADH-dependent characteristics [12]), or enhance stability. This enables multi-dimensional active-site optimization that surpasses traditional empirical approaches, significantly accelerating the development of high-performance enzymes. For example, to address the rate-limiting histidine methylation step, Li et al. employed a synergistic strategy integrating AI-driven kinetic prediction (CataPro, DLkcat), molecular dynamics simulations, and conformational dynamics analysis to guide site-directed mutagenesis of Mycolicibacrterium smegmatis EgtD. This yielded a triple mutant M3 (L53A/T59S/E282S) with 3.4- and 2.8-fold enhancements in catalytic efficiency (kcat/Km) toward L-histidine and S-adenosyl-L-methionine (SAM), respectively [13]. Despite significant advances, AI-driven enzyme engineering still faces challenges, including a lack of high-quality, standardized data, limited accuracy in predicting complex interactions (e.g., long-range epistasis), and the frequent omission of reaction environment parameters, which necessitate future efforts to establish more comprehensive databases, improve model generalizability, and develop integrated frameworks that incorporate multi-omics and environmental data.
Furthermore, the value of precision-designed enzymes extends beyond intracellular catalysis. When enzymes themselves serve as functional ingredients in the final products, immobilization and delivery technologies (such as nanocarrier encapsulation and microencapsulation) address formulation stability and controlled-release challenges, laying the groundwork for innovative applications like “active-enzyme skincare” [14,15]. Nevertheless, optimizing a single enzyme is only the first step: when multiple engineered enzymes are assembled into a pathway, the adaptation issues between endogenous metabolism and exogenous synthetic routes—such as metabolic competition and host physiological stress—emerge as critical challenges, motivating higher-dimensional, dynamic, and system-level regulation.
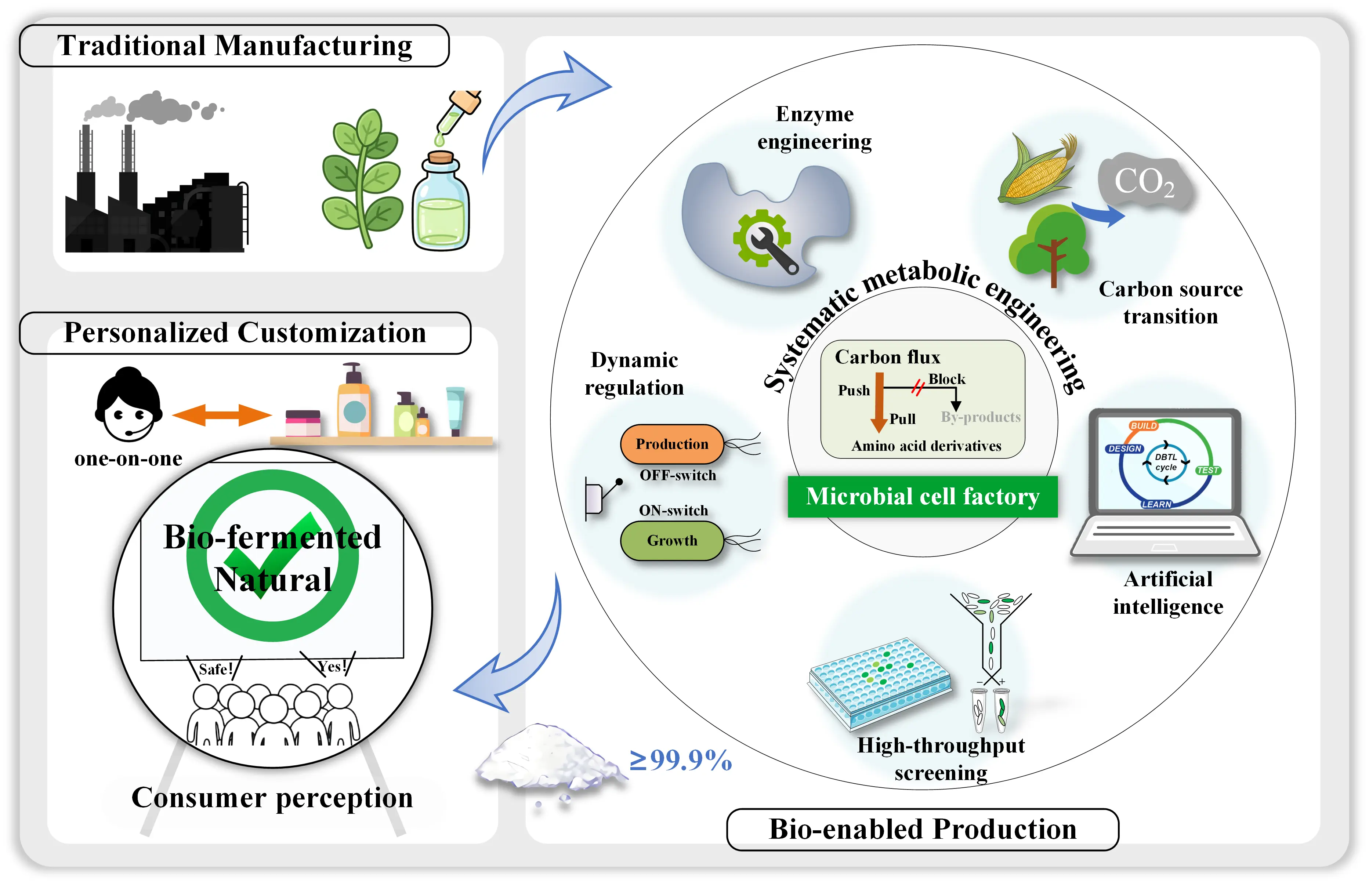
Figure 1. Toward Personalization: a schematic overview of synthetic-biology-driven production of amino acid-derived cosmetic ingredients.
1.2. Dynamic Regulation of Metabolic Pathways
For most products with short pathways that introduce minor metabolic burden, static engineering strategies remain effective and straightforward. However, for complex amino acid derivative pathways—which often involve metabolic competition or accumulation of toxic intermediates—dynamic regulation systems integrated with “intelligent genetic switches” offer distinct advantages (the ‘Dynamic regulation’ module, Figure 1). These systems usually reprogram metabolic networks in response to physiological states or environmental signals (e.g., cell density, metabolite concentrations). The goal extends beyond simple binary switching to achieving refined control that balances metabolic flux, relieves feedback inhibition, and facilitates timely efflux of toxic intermediates—thereby maximizing target product synthesis [16]. For instance, a quorum-sensing-based regulatory module can automatically activate the expression of the synthesis genes of the target product at high cell density, leading to a several-fold increase in poly-γ-glutamic acid titers [17]. Future development should focus on multi-layer dynamic regulatory networks, for instance, by coupling quorum sensing with metabolite-responsive biosensors to implement “dual-logic-gate” control, which would enhance system scalability and precision. Moreover, the introduction of optogenetic tools enables precise, spatiotemporal activation of specific pathways during fermentation using light, elevating the precision of metabolic control to unprecedented levels [18,19]. In addition to the demand for dynamic regulation of pathways, optimizing the overall performance of cell factories using irrational strategies could further enhance the efficiency and robustness of microbial cell factories in industrial environments.
1.3. Irrational Strategies and High Throughput Screening of Industrial Strains
A fundamental limitation of rational design is its static assumption—it often treats the metabolic network as a predictable equilibrium, overlooking the nonlinear phenotypic impacts of dynamic perturbations during industrial fermentation (e.g., pH shifts, dissolved oxygen gradients, substrate pulses). In contrast, irrational strategies can compensate for these shortcomings by introducing genetic variation and environmental selection, thereby better responding to dynamic disturbances. However, traditional random mutagenesis and passive screening are slow-iterative, creating a “fast design, slow evolution” bottleneck that lags behind the pace of synthetic biology design. The key recent breakthrough lies in introducing programmability and predictability into the entire mutation–screening pipeline, enabling faster and more precise phenotype development.
In mutant library construction, CRISPR-mediated directed evolution has moved beyond simple knockouts/activation. Systems like EvolvR—which fuse a Cas9 nickase with an error-prone DNA polymerase—enable continuous, autonomous mutation–selection–enrichment cycles within gRNA-specified genomic regions, dramatically enhancing the efficiency and precision of directed evolution [20,21]. Following the construction of high-quality mutant libraries, the next critical step is efficient phenotyping, driving innovation in intelligent screening platforms (the ‘High-throughput screening’ module, Figure 1). The core idea is to convert invisible intracellular states into detectable signals for high-throughput sorting. Biosensor-based screening is a prominent high throughput approach in which biosensors, from genetic circuits such as transcription-factor or riboswitch systems to protein, enzymatic and electrochemical sensors, convert metabolite levels into measurable outputs like fluorescence, luminescence, colorimetric or electrochemical signals and selectable growth phenotypes. Coupled with flow cytometry, this enables rapid mining of high-producing strains from million-variant libraries. The revolutionary frontier lies in label-free and separation-free methods. For instance, combining Raman-activated cell sorting (RACS) with metabolite-sensing third-generation riboswitches allows direct sorting based on the unique spectral fingerprints of target metabolites [22]. This approach circumvents the inherent metabolic burden and limited dynamic range of biosensors, shifting the screening paradigm from an indirect phenotype–reporter–fluorescence cascade back to the direct metabolite-to-spectral-signature readout, representing a major direction for next-generation high-throughput screening.
2. Star Molecules: Redefining Anti-Aging Efficacy
Anti-aging has become a central goal in biomedical and functional formulation development. This section focuses on ergothioneine, spermidine, carnosine, and glutathione—star molecules with demonstrated or potential anti-aging efficacy—and reviews their mechanisms of action, key industrialization bottlenecks, and corresponding engineering strategies and formulation approaches. The relevant research progress is summarized in Table 1.
2.1. Ergothioneine
Ergothioneine, a rare sulfur-containing L-histidine derivative, holds unique value in the high-end anti-aging skincare market. Its core bioactivity arises from a dual mechanism: exceptional intrinsic antioxidant capacity [23] and targeted enrichment mediated by the specific transporter OCTN1 [24]. This “intelligent targeting” property underpins its molecular rationale for use in UV protection and photoaging repair [25]. However, achieving industrial-scale production necessitates systematically overcoming two critical bottlenecks: insufficient enzymatic catalytic efficiency and prohibitively high precursor costs.
Heterologous expression of ergothioneine biosynthetic modules is often constrained by low catalytic efficiency. To overcome this limitation, enzyme engineering strategies have been employed to discover and optimize ergothioneine synthases, enhancing both their expression and activity [26,27]. For instance, Zhang et al. applied structure-guided semi-rational design and directed evolution to the Neurospora crassa-derived sulfoxide synthase (NcEgt1), obtaining a G2S/D109A double mutant with significantly enhanced catalytic efficiency. Further optimization through saturation mutagenesis yielded the methyltransferase (EgtD) mutant I50A. When combined, these engineered enzymes enabled a titer of 5.4 g/L in a 96-h fed batch fermentation, corresponding to a productivity of 56.3 mg/(L·h) [28]. These results underscore that enhancing enzyme stability and expression levels constitutes an effective approach to alleviating bottlenecks in metabolic pathways.
The exogenous supply of precursor amino acids (L-histidine, L-cysteine, L-methionine) constitutes over 60% of the total raw material cost, making the development of autonomous precursor supply systems essential for industrialization. To address this, Wu et al. began with an engineered histidine-high-producing strain and systematically engineered both the ergothioneine synthesis module and the precursor supply module. Coupled with fermentation process optimization, this integrated strategy achieved ergothioneine production without exogenous supplementation with L-histidine or L-cysteine (Figure 2a) [29]. A further innovative approach bypasses costly L-cysteine altogether. Yan et al. incorporated a rhodanese domain-containing protein (EanB) from Chlorobium limicola to construct a synthesis route that utilizes thiosulfate as an inorganic sulfur source. By further fusing betaine homocysteine methyltransferase (BHMT), they established a non-SAM-dependent methyl-donor cycle. This integrated design achieved a high titer of 7.2 g/L in a 5-L fermenter, highlighting the potential of pathway redesigning to overcome intrinsic metabolic and economic constraints [4].
The key to future research lies in integrating the aforementioned high-efficiency enzyme elements with alternative precursor supply routes to construct a “fully integrated” cell factory. Simultaneously, it is essential to optimize scaled-up fermentation processes and develop downstream extraction technologies to assess their overall cost advantage. Ultimately, this highly engineered platform aims not only to sustainably produce ergothioneine but also holds the potential to be expanded for the synthesis of other rare sulfur-containing natural products, thereby opening up a broader new field of “sulfur metabolism” biomanufacturing.
2.2. Spermidine
Spermidine, a natural polyamine, exhibits considerable potential in anti-aging and neuroprotection, primarily through its role in activating TFEB nuclear translocation to induce autophagy [30,31]. Nevertheless, conventional chemical synthesis of spermidine faces several limitations, including low conversion efficiency, residual putrescine contamination, and elevated purification costs, all of which impede its large-scale production. Consequently, the development of efficient biotechnological production platforms represents a crucial step toward enabling the broader commercial utilization of spermidine.
Current biological synthesis of spermidine primarily involves two enzymatic routes: the SAM and ASA pathways. The SAM pathway utilizes a dual-enzyme cascade involving S-adenosylmethionine decarboxylase (SAMDC) and spermidine synthase (SPDS). For example, Liu et al. co-expressed high-activity SAMDC from Thermotoga maritima and SPDS from Escherichia coli in an engineered E. coli host. Under precisely controlled reaction conditions, their system produced 3.7 g/L spermidine within 8 h [5]. The ASA pathway similarly relies on two key enzymes: carboxyspermidine dehydrogenase (CASDH), which catalyzes the aminopropyl transfer, and carboxyspermidine decarboxylase (CASDC), responsible for the subsequent decarboxylation. Liang et al. demonstrated the potential of this route by expressing CASDH from Agrobacterium fabrum str. C58 and coupling it with an NADPH regeneration system in a whole-cell biocatalyst, achieving a 91% molar conversion from aspartate semialdehyde to carboxyspermidine [32]. Despite these advances, both strategies suffer from a reliance on costly substrate supplementation. Furthermore, the SAM pathway is particularly energy-intensive, consuming three ATP molecules per spermidine synthesized, which poses a significant economic drawback for scalable production.
Xi et al. identified a previously uncharacterized spermidine biosynthetic pathway in the model cyanobacterium Synechocystis sp. PCC 6803, which employs carboxyaminopropylagmatine (CAPA) as a key intermediate and involves three distinct enzymes: carboxyaminopropylagmatine dehydrogenase (CAPADH), carboxyaminopropylagmatine decarboxylase (CAPADC), and aminopropylagmatine ureohydrolase (APAUH) [33]. This newly elucidated, SAM-independent pathway offers a promising alternative route for spermidine production. Building on this discovery, Guo et al. introduced the CAPADH/CAPADC/APAUH into the robust industrial host Serratia marcescens HBQA7. Through directed evolution, they further optimized the pathway by screening a DfCAPADC variant from Desulfovibrio desulfuricans, which displayed a six-fold enhancement in catalytic efficiency over the parental enzyme. Integrated with multi-strategy metabolic reprogramming, this engineered system produced 5.8 g/L spermidine within 72 h without requiring costly SAM precursors (Figure 2b) [34]. Together, these advances establish a scalable and economically viable platform for the sustainable microbial production of spermidine and related polyamines.
2.3. L-Carnosine
L-Carnosine, a biologically active dipeptide composed of β-alanine and L-histidine, exerts skin anti-aging effects through multiple mechanisms, including attenuating telomere shortening, exerting antioxidant and anti-glycation activities, and modulating autophagy in senescent cells. Conventional chemical synthesis routes, such as the phthalic anhydride method, are hampered by low efficiency and a significant environmental footprint, conflicting with green chemistry principles. Consequently, enzyme-catalyzed green synthesis has emerged as an increasingly attractive alternative for L-carnosine production.
Currently, enzymes capable of catalyzing L-carnosine synthesis are primarily divided into two groups: ATP-dependent carnosine synthase and the hydrolase-type enzymes, such as aminopeptidases (Pep) and dipeptidases (PepD). The latter catalyzes reversible reactions without the need for energy cofactors, offering distinct economic advantages for industrial applications and thus constituting the current research focus in this field. Recent advances delineate a clear technological trajectory extending from enzyme discovery to integrated system engineering.
-
-
Mining and Characterization of Novel Enzyme Elements: Researchers have identified highly active enzymes from diverse biological sources—including Serratia marcescens [35], Bacillus megaterium [36], Trichoderma virens Gv29-8 [37], and deep-sea metagenomes [38]—thereby expanding the available catalytic toolbox. These efforts are directed toward obtaining robust natural catalysts that exhibit both high activity and stability.
-
-
Precise Tuning of Catalytic Performance through Protein Engineering: Through rational design and site-directed modification of key residues—exemplified by the T168S/G148D mutation in SmPepD [39] and the T171A mutation in BmPepD [36]—the catalytic efficiency and stability of these enzymes have been substantially improved, an engineering advance that addresses the core bottleneck of insufficient activity in their native counterparts.
-
-
Metabolic and Process Integration from Enzyme to System: Research has progressed beyond isolated enzyme engineering to systems-level optimization of synthesis efficiency. This is achieved through strategies such as knocking out host degradation pathways, enhancing intracellular substrate accumulation, and implementing cell-free synthesis systems (Figure 2c). These integrated approaches have enabled high-yield production, achieving titers exceeding 30.3 g/L [39] or 18.6 g/L [37] in scaled 5-L fermenters.
Currently, an efficient L-carnosine synthesis system based on non-ATP-dependent hydrolases has been successfully established. However, its transition to industrialization is still hindered by two core challenges: the high cost of substrates, especially L-histidine, and the thermodynamic barrier imposed by the reversible reaction equilibrium inherent to hydrolytic catalysis. The key to future breakthroughs, therefore, lies in constructing an integrated next generation biomanufacturing system. This requires enhancing the host’s autonomous precursor synthesis to reduce costs, coupled with employing reaction engineering strategies (e.g., in-situ product separation) to drive the equilibrium toward synthesis. Ultimately, this green synthesis pathway is anticipated to evolve into a modular platform for the production of other high value dipeptides.
2.4. Glutathione
Glutathione, one of the most critical low molecular weight thiol compounds in living organisms, serves as a central molecule in maintaining cellular homeostasis and delaying aging. Its broad application in cosmetics has driven substantial market demand [40]. Saccharomyces cerevisiae, owing to its inherent capacity for efficient intracellular glutathione accumulation, remains the predominant industrial host for glutathione production [41,42]. Numerous efforts have therefore been directed toward developing high yielding strains through genetic engineering and cultivation optimization [43,44,45]. Nevertheless, the yeast-based process faces two major constraints: the requirement for expensive L-cysteine supplementation and the high extraction costs associated with intracellular product accumulation. These factors collectively result in a product titer that remains comparatively low (~6 g/L), presenting a significant economic ceiling.
To address these limitations, research efforts have pivoted toward E. coli, a host naturally equipped with product efflux machinery [46,47]. This host not only facilitates product secretion into the culture medium—thereby simplifying downstream recovery—but also enables more systematic engineering to mitigate precursor dependency. For instance, heterologous expression of high-efficiency glutathione synthetases has enabled intracellular glutathione titers of up to 15.21 g/L [48]. In another key advance, Mori et al. employed metabolomics to pinpoint the conversion of O-acetylserine to L-cysteine as the central metabolic bottleneck. By innovatively substituting thiosulfate as an alternative sulfur source, the group devised a novel biosynthetic route that eliminates reliance on exogenous L-cysteine (Figure 2d) [49]. Following comprehensive optimization, this engineered pathway attained a titer of 22 g/L—the highest reported in the literature—underscoring its considerable promise as an industrially scalable alternative.
While the E. coli platform offers clear advantages in both titer and downstream processing, its path to industrial application remains challenging. Precise control over the intracellular redox balance—essential for maintaining a high ratio of reduced glutathione to its oxidized form—remains an unresolved engineering hurdle. Moreover, the robustness and stability of the process under scaled up fermentation conditions await further demonstration. To truly enable a host platform shift, future work must focus on deciphering and reprogramming the redox regulatory network of the cell, coupled with rigorous scale up validation from bench to pilot scale production.
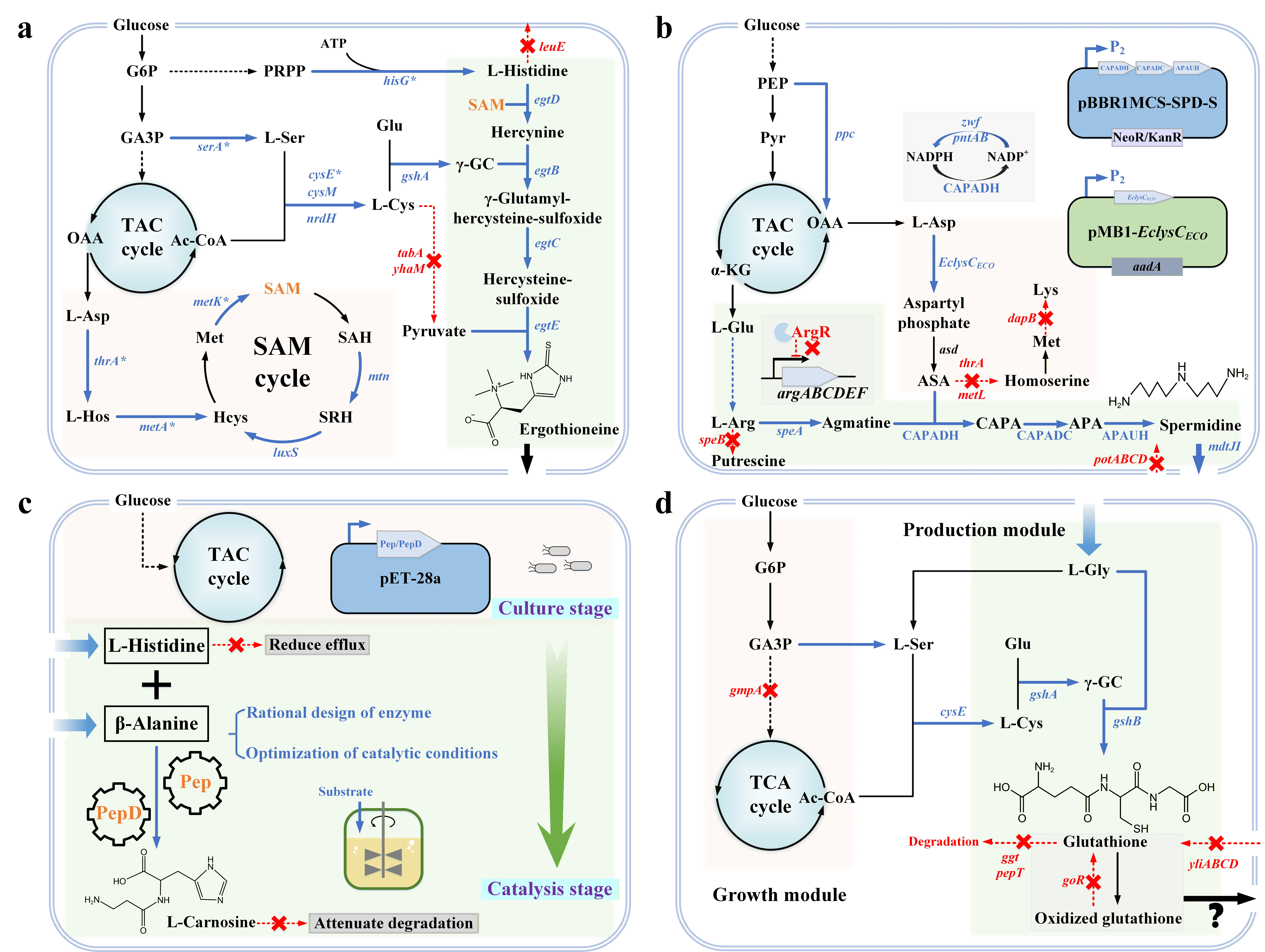
Figure 2. Representative studies on the synthesis of ergothioneine, spermidine, L-carnosine, and glutathione. (a) Metabolic engineering of E. coli for producing ergothioneine [29]; (b) Overall metabolic engineering of S. marcescens HBQA7 for producing spermidine via CAPA pathway [34]; (c) Systematic optimization of the enzymatic system for efficient L-carnosine synthesis [37,39]; (d) Engineering E. coli for efficient glutathione production without adding L-cysteine [49]; The schematic uses the following conventions: blue arrows for overexpression, red crosses for knockout, asterisks (*) for mutations, and a question mark (?) for an unknown efflux transporter.
Table 1. Recent advances in the biosynthesis of ergothioneine, spermidine, L-carnosine, and glutathione.
|
Product |
Strains |
Production Mode |
Substrate |
Key Strategies |
Titer |
Productivity |
Culture Scale |
References |
|---|---|---|---|---|---|---|---|---|
|
Ergothioneine |
E. coli BW25113 |
Fermentation |
Glucose; L-histidine, L-methionine |
Overexpression of egtDEMs, egtBMp, cysE*, serA* and ydeD; ΔmetJ |
657 mg/L |
3.42 mg/L/h |
Shake flask |
[26] |
|
E. coli BW25113 |
Fermentation |
Glucose; L-histidine, L-cysteine, L-methionine |
Overexpression of tregt1 and tregt2 |
4.34 g/L |
30.35 mg/L/h |
2-L bioreactor |
[27] |
|
|
E. coli BL21 (DE3) |
Fermentation |
Glycerol; L-histidine, L-cysteine, L-methionine |
Overexpression of egtE, egtD and TNcEgt1 with semirational design and random mutations |
5.4 g/L |
56.3 mg/L/h |
5-L bioreactor |
[28] |
|
|
E. coli MG1655 |
Fermentation |
Glucose; L-methionine |
Overexpression of egtDMs, egtBMp, Ncegt2, hisG*cg, hisDBCHAFI, cysE*, serA* and metK*; ΔtnaA |
7.2 g/L |
120 mg/L/h |
2-L bioreactor |
[29] |
|
|
E. coli BL21 (DE3) |
Fermentation |
Glycerol; Betaine, Na2S2O3 |
Overexpression of egtDEMs, Ncegt1, hisABCDFHI, glpF, dhak, gldA and mfsT1Mn; expression of hisG*Cg; ΔpurF and ΔmetJ; integration of betaine-driven methyl supply and inorganic sulfur modules |
7.2 g/L |
75 mg/L/h |
5-L bioreactor |
[4] |
|
|
Spermidine |
E. coli BL21 (DE3) |
Whole-cell catalysis |
1,3-diaminopropane, putrescine |
BvHSSE232D/D361E |
933.5 mg/L |
155.6 mg/L/h |
Shake flask |
[50] |
|
E. coli BL21 (DE3) |
Whole-cell catalysis |
Putrescine, S-adenosylmethionine |
SpeE and SpeDZm; two-enzyme cascade system in vitro |
3.7 g/L |
460 mg/L/h |
Shake flask |
[5] |
|
|
E. coli BL21 (DE3) |
Fermentation |
Glucose |
Overexpression of slr0049-sll0873-sll0228, ppc, speA and amvA; ΔthrA, ΔmetL, ΔdapA, ΔargR and ΔspeB |
1164.22 mg/L |
12.13 mg/L/h |
5-L bioreactor |
[51] |
|
|
Serratia marcescens HBQA7 |
Fermentation |
Glucose |
Overexpression of slr0049-DfCAPADC-sll0228, speA, and lysC*eco; P2-driven overexpression of ppc, zwf, pntAB, and mdtJI; ΔspeG, ΔthrA, ΔmetL, ΔdapA, ΔargR, ΔspeB and ΔpotA |
5.8 g/L |
80.6 mg/L/h |
5-L bioreactor |
[34] |
|
|
L-Carnosine |
- |
Enzymatic catalysis |
β-alanine, L-histidine |
Identification of SmPepD; NF membrane separation |
72.3 g/L |
18.08 g/L/d |
15-L bioreactor |
[35] |
|
- |
Enzymatic catalysis |
β-alanine, L-histidine |
BmPepDT171N |
7.08 g/L |
885 mg/L/h |
Shake flask |
[36] |
|
|
E. coli BL21 (DE3) |
Whole-cell catalysis |
β-alanine methyl ester, L-histidine |
Identification of the mutation in the deep-sea aminopeptidase |
2.29 g/L |
382 mg/L/h |
Shake flask |
[38] |
|
|
E. coli BL21 (DE3) |
Whole-cell catalysis |
β-alanine, L-histidine |
SmPepDT168S/G148D, ΔyeaS |
30.3 g/L |
631 mg/L/h |
5-L bioreactor |
[39] |
|
|
C. glutamicum |
Fermentation |
Glucose |
Overexpression of CARNS1, hisG, rel, and panD; enhancing the PPP and attenuating pgi |
323.26 mg/L |
6.73 mg/L/h |
2-L bioreactor |
[52] |
|
|
E. coli BL21 (DE3) |
Enzymatic catalysis |
β-alanine methyl ester, L-histidine |
Identification of TrvPep; ΔpepA |
18.6 g/L |
6.2 g/L/h |
5-L bioreactor |
[37] |
|
|
Glutathione |
S. cerevisiae HBSD-W01-013 |
Fermentation |
Glucose; L-glutamic acid |
Screening of high glutathione-producing strains and fermentation medium optimization |
3.70 g/L |
77 mg/L/h |
Shake flask |
[44] |
|
S. cerevisiae CGMCC 2842 |
Fermentation |
Corn steep liquor, Molasses |
Stepwise control based on oxidative stress and energy metabolism |
5.76 g/L |
53 mg/L/h |
10-L bioreactor |
[45] |
|
|
E. coli BL21 (DE3) |
Fermentation |
Glucose, L-glutamic acid, L-cysteine, Glycine |
Overexpression of gshFSt |
15.21 g/L |
0.82 g/L/h |
5-L bioreactor |
[48] |
|
|
E. coli BW25113 |
Fermentation |
Glucose; Glycine, (NH4)2SO4 |
Overexpression of gshA, gshB; PompF-driven overexpression of cysE; Δggt, ΔpepT, Δgor, and ΔyliABCD; supplementation of glycine and sodium thiosulfate |
22 g/L |
0.41 g/L/h |
5-L bioreactor |
[49] |
3. Industrialization Bottlenecks: Challenges Beyond Technology
3.1. The Cost Paradox: High Titer ≠ Low Cost
Although strain improvements have substantially increased the fermentation titers of peptides, ergothioneine, and other target products, the complexity and high cost of downstream separation and purification still account for roughly half of total production costs, creating an industry-level bottleneck where high titer does not equate to low cost. This impasse may be alleviated by innovations on two fronts: simplifying the production process and rethinking product formats.
Process simplification centers on the principle of “designing downstream processes upstream”—considering extraction and separation requirements at the strain-design stage. By engineering cellular membrane transport systems (e.g., efflux pumps) to promote the secretion of target products into the extracellular medium [53], we can fundamentally avoid complex cell disruption and intracellular impurity removal steps. Coupling this secretion strategy with in situ technologies such as fermentation–extraction coupling could enable continuous, intensified production, significantly reducing downstream purification costs [54]. Product innovation involves the direct use of fermentation crude extracts. Using sterilized whole fermentation broth or simply clarified filtrate as “bio-active complexes” rich in active ingredients saves purification costs while preserving the natural synergistic components present in the fermentation system—aligning well with “clean beauty” market trend, which emphasizes ingredient transparency, safety, and sustainability. However, the industrialization of this pathway hinges on establishing a stringent quality standard system for these crude extracts to ensure batch-to-batch consistency, microbial safety, and efficacy uniformity.
3.2. Regulation and Certification: Rethinking the Definition of “Natural”
Currently, the Cosmetics Organic and Natural Standard (COSMOS) accepts “natural origin” ingredients derived from traditional fermentation, but only when non-genetically modified organism (non-GMO) microbial strains and substrates are used; any organisms modified through synthetic biology or gene editing are treated as GMOs and are therefore excluded from natural and organic categories [55]. To overcome this regulatory barrier, the industry could pioneer an independent “Bio-fermented Natural” label (or certification) for products derived from sustainable substrates and gene-edited strains, emphasizing their sustainability and carbon footprint advantages while clearly differentiating them from established “natural/organic” claims. Complementary efforts must include the systematic development of toxicology databases that quantify clearance efficiency and provide safety assessments for residuals such as engineered DNA, host proteins, and antibiotic resistance genes, thereby preparing for potential future quantitative evaluations by EU scientific bodies like the SCCS and EFSA.
3.3. Consumer Perception: The Art of “Scientific Storytelling”
Public concerns about “genetic modification” stand in contrast to their enthusiasm for “fermentation products”, a disparity rooted in narrative framing. Effective market strategies must reframe the discourse: translating “gene editing” into an accessible narrative such as “cell factories mimicking plant biosynthesis”, and ensuring transparency throughout the entire process from “sugar to active ingredient” to build trust. Technically, micro-dosing strategies (using high-purity active ingredients at ppm levels) ensure efficacy while significantly reducing perceived risk. For building trust, publishing independent third-party test results and investing in targeted science communication are crucial to reframing synthetic biology as a “precision skincare” value proposition.
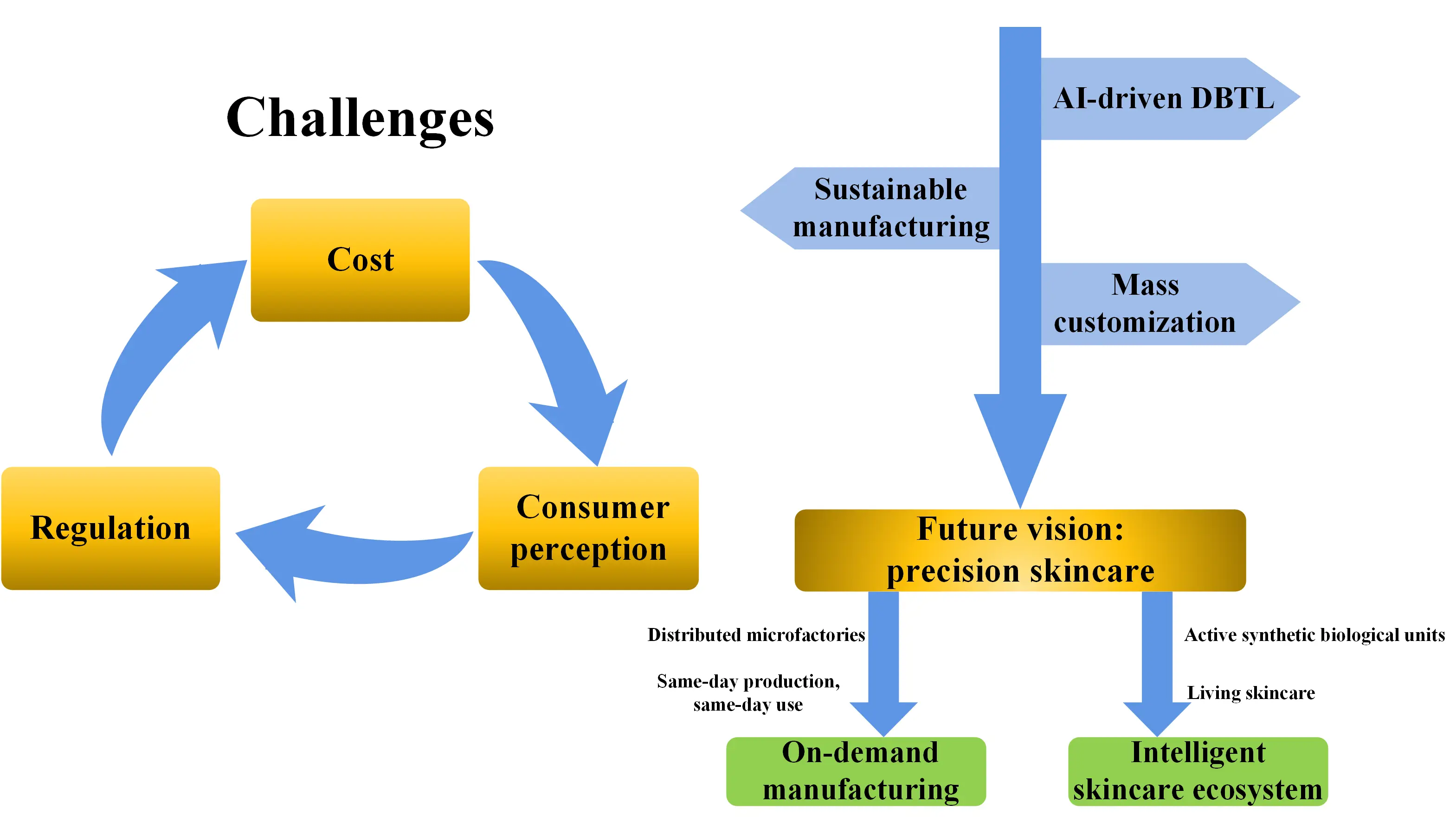
In summary, the industrialization of synthetic biology-driven cosmetic ingredients still faces a triple bottleneck: production costs, regulatory certification, and consumer perception. To systematically navigate these challenges and outline viable future pathways, we propose an integrated development roadmap (Figure 3), which aims to progress from current industrial bottlenecks towards establishing an efficient, sustainable, and personalized next-generation manufacturing system for cosmetic ingredients.
4. Future Outlook: AI Empowerment and Sustainable Manufacturing
4.1. AI Reshapes the R&D Paradigm
Next-generation computational tools for protein structure prediction and design are fundamentally reshaping traditional approaches to enzyme design and metabolic engineering. AI-driven design–build–test–learn (DBTL) cycles are expected to accelerate strain development efficiency by an order of magnitude within five years [56]. By leveraging machine learning to mine multi-omics data, these paradigms reveal hidden cross-pathway connections and directly predict non-intuitive targets beyond human experience, thereby driving the field from trial-and-error to predictive design, and from local optimization to global design [57]. It is crucial to stress that the reliability of these models ultimately depends on the continuous accumulation and validation of high-quality experimental data.
4.2. Carbon Source Revolution: From Food to Waste
Current biomanufacturing relies heavily on edible sugars like glucose, posing ethical competition with the food supply chain. To build a resilient and sustainable industrial future, we must adopt a tiered carbon source strategy: in the short term, prioritize second-generation feedstocks such as lignocellulosic hydrolysates and industrial by-products (e.g., glycerol) [58]; in the long term, explore autotrophic fermentation using syngas (CO/H2) or CO2, which not only liberates the industry from agricultural carbon dependence but also enables a “carbon-negative manufacturing” paradigm [59]. Life cycle assessment must guide this transition, systematically evaluating technological maturity and economics to prioritize industrially viable routes. This represents not merely a technical upgrade, but a responsible evolution of industrial ethics.
4.3. Precision Skincare: From “One-Size-Fits-All” to Personalized Customization
The ultimate manifestation of the precision creation paradigm is its translation into personalized customization. The value of synthetic biology lies in on-demand manufacturing. While the vision of fully personalized, on-site production represents a long-term frontier, a series of feasible intermediate milestones is already emerging.
In the near to medium term, personalization will likely evolve through scalable, yet increasingly tailored, biomanufacturing. Initially, modular cell factories could be designed to produce a library of pre-defined active ingredient variants (e.g., a suite of ergothioneine analogs with different stability or penetration profiles). Consumer skin type or concern data, gathered through digital questionnaires or basic imaging, could then guide the selection and blending of these pre-manufactured actives into customized formulations. Concurrently, advances in rapid, low-cost microbiome or biomarker profiling could enable the precise categorization of consumers based on skin status [60], for which centralized facilities would produce optimized active ingredients with specific efficacy profiles for different clusters (e.g., ergothioneine derivatives with enhanced antioxidant and anti-inflammatory properties for sensitive skin clusters, or specific signal peptides for aging skin clusters). This “mass customization” model leverages synthetic biology’s flexibility for producing diversity while relying on established supply chains for distribution, representing a critical and pragmatic step toward full personalization.
Looking further ahead, the integration of real-time skin microbiome and genetic analysis with AI-driven formulation engines will enable truly dynamic personalization. Modular cell factories could then rapidly switch to produce different derivatives on demand, and distributed micro-factories, such as on-site fermenters in retail or clinical settings, would realize the concept of “same-day production, same-day use” for maximal freshness and efficacy. This progression is poised to fundamentally transform cosmetic supply chains—shifting gradually from large-scale mass production to agile, precise, and ultimately personal response.
5. Conclusions
Amino acid derivative biosynthesis has evolved from “substituting traditional methods” to “creating novel functions”. We stand at a tipping point: future cosmetics will not be mere chemical blends but intelligent systems containing “active synthetic biological units”. Microencapsulated engineered probiotics could continuously secrete actives on skin, dynamically responding to environmental stress. While this “living skincare” concept seems futuristic, the technical roadmap is emerging. Realizing this vision requires interdisciplinary collaboration: synthetic biologists provide parts, fermentation engineers optimize processes, dermatologists validate efficacy, and toxicologists assess safety. Only by building a full-chain innovation ecosystem can the cosmetics industry evolve from “manufacturing” to “intelligent creation”, delivering truly safe, effective, and sustainable beauty solutions.
Statement of the Use of Generative AI and AI-Assisted Technologies in the Writing Process
During the preparation of this manuscript, the author(s) used DeepL and ChatGPT in order to improve readability and language editing. After using these tools, the author(s) reviewed and edited the content as needed and take(s) full responsibility for the content of the published article.
Acknowledgments
We thank members of the laboratory for helpful discussions.
Author Contributions
Writing—Original Draft Preparation, J.Y.; Visualization, M.Z. and F.H.; Writing—Review & Editing, H.W. and Q.M.; Project Administration, X.X.; Funding Acquisition, X.X.
Ethics Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
No datasets were generated or analyzed during the current study.
Funding
This research received funding from the National Key Research and Development Program of China (2022YFA0911800), Key Research and Development Program of Ningxia Hui Autonomous Region (2025BEE02021), National Natural Science Foundation of China (22378315, 32200038), Natural Science Foundation of Tianjin (24JCYBJC00830).
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- Benner SA, Sismour AM. Synthetic biology. Nat. Rev. Genet. 2005, 6, 533–543. DOI:10.1038/nrg1637 [Google Scholar]
- Auslander S, Auslander D, Fussenegger M. Synthetic biology—The synthesis of biology. Angew. Chem. Int. Ed. 2017, 56, 6396–6419. DOI:10.1002/anie.201609229 [Google Scholar]
- Jiang S, Wang RR, Wang DH, Zhao CG, Ma Q, Wu HY, et al. Metabolic reprogramming and biosensor-assisted mutagenesis screening for high-level production of L-arginine in Escherichia coli. Metab. Eng. 2023, 76, 146–157. DOI:10.1016/j.ymben.2023.02.003 [Google Scholar]
- Yan J, Chen Y, Ma Y, Yang X, Yao Y, Zhao G, et al. Efficient ergothioneine production through reconstruction of the methyl and sulfur supply systems in Escherichia coli. Trends Biotechnol. 2026, 44, 477–495. DOI:10.1016/j.tibtech.2025.09.014 [Google Scholar]
- Liu Y, Guo X, Wang X, Chen KQ, Ouyang PK. A two-enzyme cascade system for the bio-production of spermidine from putrescine. Mol. Catal. 2021, 504, 111439. DOI:10.1016/j.mcat.2021.111439 [Google Scholar]
- Jiang R, Yuan S, Zhou Y, Wei Y, Li F, Wang M, et al. Strategies to overcome the challenges of low or no expression of heterologous proteins in Escherichia coli. Biotechnol. Adv. 2024, 75, 108417. DOI:10.1016/j.biotechadv.2024.108417 [Google Scholar]
- Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. DOI:10.1038/s41586-021-03819-2 [Google Scholar]
- Lin Z, Akin H, Rao R, Hie B, Zhu Z, Lu W, et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science 2023, 379, 1123–1130. DOI:10.1126/science.ade2574 [Google Scholar]
- Leman JK, Weitzner BD, Lewis SM, Adolf-Bryfogle J, Alam N, Alford RF, et al. Macromolecular modeling and design in Rosetta: Recent methods and frameworks. Nat. Methods 2020, 17, 665–680. DOI:10.1038/s41592-020-0848-2 [Google Scholar]
- Dauparas J, Anishchenko I, Bennett N, Bai H, Ragotte RJ, Milles LF, et al. Robust deep learning–based protein sequence design using ProteinMPNN. Science 2022, 378, 49–56. DOI:10.1126/science.add2187 [Google Scholar]
- Kroll A, Ranjan S, Engqvist MK, Lercher MJ. A general model to predict small molecule substrates of enzymes based on machine and deep learning. Nat. Commun. 2023, 14, 2787. DOI:10.1038/s41467-023-38347-2 [Google Scholar]
- Kim J, Woo J, Park JY, Kim K-J, Kim D. Deep learning for NAD/NADP cofactor prediction and engineering using transformer attention analysis in enzymes. Metab. Eng. 2025, 87, 86–94. DOI:10.1016/j.ymben.2024.11.007 [Google Scholar]
- Li Y, Rao J, Jiang Z, Zhang X, Zhu Q, Zhu C, et al. AI-driven engineering of EgtD enabling high-efficiency ergothioneine synthesis with a multi-enzyme cascade. Int. J. Biol. Macromol. 2025, 327, 147266. DOI:10.1016/j.ijbiomac.2025.147266 [Google Scholar]
- Zhou H, Luo D, Chen D, Tan X, Bai X, Liu Z, et al. Current advances of nanocarrier technology-based active cosmetic ingredients for beauty applications. Clin. Cosmet. Investig. Dermatol. 2021, 14, 867–887. DOI:10.2147/CCID.S313429 [Google Scholar]
- Hao X, Liu P, Chu X. Recent advances in the strategies of simultaneous enzyme immobilization accompanied by nanocarrier synthesis. Appl. Sci. 2024, 14, 3702. DOI:10.3390/app14093702 [Google Scholar]
- Qin H, Zhou J, Pang A, Huang L, Liu Z, Zheng Y. Microbial dynamic regulatory tools: Design, applications, and prospects. ACS Synth. Biol. 2025, 14, 2418–2432. DOI:10.1021/acssynbio.5c00219 [Google Scholar]
- Hu LX, Zhao M, Hu WS, Zhou MJ, Huang JB, Huang XL, et al. Poly-γ-glutamic acid production by engineering a degU quorum-sensing circuit in Bacillus subtilis. ACS Synth. Biol. 2022, 11, 4156–4170. DOI:10.1021/acssynbio.2c00464 [Google Scholar]
- Figueroa D, Ruiz D, Tellini N, De Chiara M, Kessi-Pérez EI, Martínez C, et al. Optogenetic control of horizontally acquired genes prevent stuck fermentations in yeast. Microbiol. Spectr. 2025, 13, e01794-24. DOI:10.1128/spectrum.01794-24 [Google Scholar]
- Gao C, Song W, Ye C, Ding Q, Wei W, Hu G, et al. Bifunctional optogenetic switch powered NADPH availability for improving l-valine production in Escherichia coli. ACS Sustain. Chem. Eng. 2024, 12, 15103–15113. DOI:10.1021/acssuschemeng.4c04806 [Google Scholar]
- Halperin SO, Tou CJ, Wong EB, Modavi C, Schaffer DV, Dueber JE. CRISPR-guided DNA polymerases enable diversification of all nucleotides in a tunable window. Nature 2018, 560, 248–252. DOI:10.1038/s41586-018-0384-8 [Google Scholar]
- Sadanand S. EvolvR-ing to targeted mutagenesis. Nat. Biotechnol. 2018, 36, 819. DOI:10.1038/nbt.4247 [Google Scholar]
- Wang X, Sun Q, Diao Z, Xu J, Ma B. Advances with applications of Raman spectroscopy in single-cell phenotype sorting and analysis. Synth. Biol. J. 2023, 4, 204–224. DOI:10.12211/2096-8280.2022-043 [Google Scholar]
- Cheah IK, Halliwell B. Ergothioneine; antioxidant potential, physiological function and role in disease. Biochim. Et Biophys. Acta (BBA)-Mol. Basis Dis. 2012, 1822, 784–793. DOI:10.1016/j.bbadis.2011.09.017 [Google Scholar]
- Gründemann D, Harlfinger S, Golz S, Geerts A, Lazar A, Berkels R, et al. Discovery of the ergothioneine transporter. Proc. Natl. Acad. Sci. USA 2005, 102, 5256–5261. DOI:10.1073/pnas.0408624102 [Google Scholar]
- Hseu Y-C, Lo H-W, Korivi M, Tsai Y-C, Tang M-J, Yang H-L. Dermato-protective properties of ergothioneine through induction of Nrf2/ARE-mediated antioxidant genes in UVA-irradiated Human keratinocytes. Free Radic. Biol. Med. 2015, 86, 102–117. DOI:10.1016/j.freeradbiomed.2015.05.026 [Google Scholar]
- Kamide T, Takusagawa S, Tanaka N, Ogasawara Y, Kawano Y, Ohtsu I, et al. High production of ergothioneine in Escherichia coli using the sulfoxide synthase from Methylobacterium strains. J. Agric. Food Chem. 2020, 68, 6390–6394. DOI:10.1021/acs.jafc.0c01846 [Google Scholar]
- Chen Z, He Y, Wu X, Wang L, Dong Z, Chen X. Toward more efficient ergothioneine production using the fungal ergothioneine biosynthetic pathway. Microb. Cell Factories 2022, 21, 76. DOI:10.1186/s12934-022-01807-3 [Google Scholar]
- Zhang L, Tang J, Feng M, Chen S. Engineering methyltransferase and sulfoxide synthase for high-yield production of ergothioneine. J. Agric. Food Chem. 2022, 71, 671–679. DOI:10.1021/acs.jafc.2c07859 [Google Scholar]
- Wu HY, Liu WC, Xie XX, Jiang S, Ma Q, Shuai ZL. A Genetic Engineering Bacteria Producing Ergothioneine and Its Construction Method and Application. CN116121161A, 16 May 2023. [Google Scholar]
- Madeo F, Eisenberg T, Pietrocola F, Kroemer G. Spermidine in health and disease. Science 2018, 359, eaan2788. DOI:10.1126/science.aan2788 [Google Scholar]
- Zhang H, Alsaleh G, Feltham J, Sun Y, Napolitano G, Riffelmacher T, et al. Polyamines control eIF5A hypusination, TFEB translation, and autophagy to reverse B cell senescence. Mol. Cell 2019, 76, 110–125.e9. DOI:10.1016/j.molcel.2019.08.005 [Google Scholar]
- Liang X, Deng H, Xiong T, Bai Y, Fan TP, Zheng X, et al. Overexpression and biochemical characterization of a carboxyspermidine dehydrogenase from Agrobacterium fabrum str. C58 and its application to carboxyspermidine production. J. Sci. Food Agric. 2022, 102, 3858–3868. DOI:10.1002/jsfa.11735 [Google Scholar]
- Xi H, Nie X, Gao F, Liang X, Li H, Zhou H, et al. A bacterial spermidine biosynthetic pathway via carboxyaminopropylagmatine. Sci. Adv. 2023, 9, eadj9075. DOI:10.1126/sciadv.adj9075 [Google Scholar]
- Gou L, Wang J, Liu D, Wu S, Zhou X, Fan T-P, et al. Multi-strategy metabolic reprogramming of Serratia marcescens HBQA7 for high-efficiency spermidine biosynthesis via the carboxyaminopropylagmatine pathway. Chem. Eng. J. 2025, 522, 167413. DOI:10.1016/j.cej.2025.167413 [Google Scholar]
- Yin DY, Pan J, Zhu J, Liu YY, Xu JH. A green-by-design bioprocess for L-carnosine production integrating enzymatic synthesis with membrane separation. Catal. Sci. Technol. 2019, 9, 5971–5978. DOI:10.1039/C9CY01622H [Google Scholar]
- Guan BH, Yin WT, Cao BH, Zhao C, Pan J, Xu JH. Characterization and mutagenesis of a high-activity and highly substrate-tolerant dipeptidase for L-carnosine biosynthesis via reversed hydrolysis. Mol. Catal. 2023, 549, 113500. DOI:10.1016/j.mcat.2023.113500 [Google Scholar]
- Zhu Z, Han H, Zhao Y, Wang Y, Zhang Z, Huang W, et al. Heterologous expression of an aminopeptidase in Escherichia coli for cell-free biosynthesis of l-carnosine. Int. J. Biol. Macromol. 2025, 332, 148703. DOI:10.1016/j.ijbiomac.2025.148703 [Google Scholar]
- She JJ, Fu LH, Zheng XW, Li J, Wang LM, Yu B, et al. Characterization of a new L-carnosine synthase mined from deep-sea sediment metagenome. Microb. Cell Factories 2022, 21, 129. DOI:10.1186/s12934-022-01854-w [Google Scholar]
- Liu YR, Pan XW, Zhang HW, Zhao ZQ, Teng ZX, Rao ZM. Combinatorial protein engineering and transporter engineering for efficient synthesis of L-Carnosine in Escherichia coli. Bioresour. Technol. 2023, 387, 129628. DOI:10.1016/j.biortech.2023.129628 [Google Scholar]
- Al-Temimi AA, Al-Mossawi AEB, Al-Hilifi SA, Korma SA, Esatbeyoglu T, Rocha JM, et al. Glutathione for food and health applications with emphasis on extraction, identification, and quantification methods: A review. Metabolites 2023, 13, 465. DOI:10.3390/metabo13040465 [Google Scholar]
- Malairuang K, Krajang M, Sukna J, Rattanapradit K, Chamsart S. High cell density cultivation of Saccharomyces cerevisiae with intensive multiple sequential batches together with a novel technique of fed-batch at cell level (FBC). Processes 2020, 8, 1321. DOI:10.3390/pr8101321 [Google Scholar]
- Santos LO, Silva PGP, Lemos Junior WJF, De Oliveira VS, Anschau A. Glutathione production by Saccharomyces cerevisiae: Current state and perspectives. Appl. Microbiol. Biotechnol. 2022, 106, 1879–1894. DOI:10.1007/s00253-022-11826-0 [Google Scholar]
- Zhu Y, Sun J, Zhu Y, Wang L, Qi B. Endogenic oxidative stress response contributes to glutathione over-accumulation in mutant Saccharomyces cerevisiae Y518. Appl. Microbiol. Biotechnol. 2015, 99, 7069–7078. DOI:10.1007/s00253-015-6629-7 [Google Scholar]
- Hu XY, Shen XL, Zhu S, Zeng HW, Shuai YY. Optimization of glutathione production in Saccharomyces cerevisiae HBSD-W08 using Plackett–Burman and central composite rotatable designs. BMC Microbiol. 2023, 23, 11. DOI:10.1186/s12866-022-02742-4 [Google Scholar]
- Chen HL, Cao XT, Zhu NQ, Jiang LH, Zhang XG, He QM, et al. A stepwise control strategy for glutathione synthesis in Saccharomyces cerevisiae based on oxidative stress and energy metabolism. World J. Microbiol. Biotechnol. 2020, 36, 1–10. DOI:10.1007/s11274-020-02895-2 [Google Scholar]
- Smirnova GV, Muzyka NG, Ushakov VY, Tyulenev AV, Oktyabrsky ON. Extracellular superoxide provokes glutathione efflux from Escherichia coli cells. Res. Microbiol. 2015, 166, 609–617. DOI:10.1016/j.resmic.2015.07.007 [Google Scholar]
- Poole RK, Cozens AG, Shepherd M. The CydDC family of transporters. Res. Microbiol. 2019, 170, 407–416. DOI:10.1016/j.resmic.2019.06.003 [Google Scholar]
- Wang C, Zhang J, Wu H, Li ZM, Ye Q. Heterologous gshF gene expression in various vector systems in Escherichia coli for enhanced glutathione production. J. Biotechnol. 2015, 214, 63–68. DOI:10.1016/j.jbiotec.2015.09.004 [Google Scholar]
- Mori H, Matsui M, Bamba T, Hori Y, Kitamura S, Toya Y, et al. Engineering Escherichia coli for efficient glutathione production. Metab. Eng. 2024, 84, 180–190. DOI:10.1016/j.ymben.2024.07.001 [Google Scholar]
- Liu WJ, Hu XX, Yan Y, Cai YJ. Rational engineering of homospermidine synthase for enhanced catalytic efficiency toward spermidine synthesis. Synth. Syst. Biotechnol. 2024, 9, 549–557. DOI:10.1016/j.synbio.2024.04.012 [Google Scholar]
- Wang J, Gou L, Liu D, Wu S, Zhou X, Fan T-P, et al. Highly efficient spermidine production system in Escherichia coli BL21 (DE3) based on precursor metabolic modules optimization and carboxyaminopropylagmatine pathway construction. Enzym. Microb. Technol. 2025, 193, 110763. DOI:10.1016/j.enzmictec.2025.110763 [Google Scholar]
- Kim M, Ko YJ, Jeong DW, Jeong W-Y, Han SO. Ecofriendly synthesis of L-carnosine in metabolically engineered Corynebacterium glutamicum by reinforcing precursor accumulation. ACS Synth. Biol. 2021, 10, 1553–1562 DOI:10.1021/acssynbio.1c00168 [Google Scholar]
- Jones CM, Hernández Lozada NJ, Pfleger BF. Efflux systems in bacteria and their metabolic engineering applications. Appl. Microbiol. Biotechnol. 2015, 99, 9381–9393. DOI:10.1007/s00253-015-6963-9 [Google Scholar]
- Tönjes S, Uitterhaegen E, De Winter K, Soetaert W. Reactive extraction technologies for organic acids in industrial fermentation processes–A review. Sep. Purif. Technol. 2025, 356, 129881. DOI:10.1016/j.seppur.2024.129881 [Google Scholar]
- COSMOS Standard: Version 4.2; Organic and Natural Cosmetic Standards. COSMOS-Standard AISBL: Brussels, Belgium, 2025. [Google Scholar]
- Liao L, Xie M, Zheng X, Zhou Z, Deng Z, Gao J. Molecular insights fast-tracked: AI in biosynthetic pathway research. Nat. Prod. Rep. 2025, 42, 911–936. DOI:10.1039/D4NP00003J [Google Scholar]
- Kim GB, Kim WJ, Kim HU, Lee SY. Machine learning applications in systems metabolic engineering. Curr. Opin. Biotechnol. 2020, 64, 1–9. DOI:10.1016/j.copbio.2019.08.010 [Google Scholar]
- Dharmaraja J, Shobana S, Arvindnarayan S, Francis RR, Jeyakumar RB, Saratale RG, et al. Lignocellulosic biomass conversion via greener pretreatment methods towards biorefinery applications. Bioresour. Technol. 2023, 369, 128328. DOI:10.1016/j.biortech.2022.128328 [Google Scholar]
- Liu Z, Wang K, Chen Y, Tan T, Nielsen J. Third-generation biorefineries as the means to produce fuels and chemicals from CO2. Nat. Catal. 2020, 3, 274–288. DOI:10.1038/s41929-019-0421-5 [Google Scholar]
- Long B, Pan W, Wu S, Nong Q, Li W, Chen S, et al. Advances in the application of multi-omics analysis in skin aging. Front. Aging 2025, 6, 1596050. DOI:10.3389/fragi.2025.1596050 [Google Scholar]