Fast Backcross Breeding for Climate-Resilient Cereals: Integrating Speed Breeding, Marker-Assisted Backcrossing and Genomic Selection
Chenchen Zhao
1
Matthew Tom Harrison
1
Ke Liu
1
Chengdao Li
2
Zhonghua Chen
3
Sergey Shabala
1,4,5
Meixue Zhou
1,*

Received: 30 January 2026 Revised: 24 February 2026 Accepted: 27 February 2026 Published: 05 March 2026
© 2026 The authors. This is an open access article under the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/).
1. Introduction
Climate change intensifies the frequency, duration, and severity of abiotic stresses, which together account for ~50% of global yield losses in major staple crops worldwide [1], leading to over USD$3 trillion loss during the past three decades (FAO, 2025). Abiotic stresses affect crop physiology by multiple means. For example, drought triggers stomatal closure, impairs mesophyll conductance, ultimately restricting carbon assimilation, biomass accumulation, and grain filling [2], whilst heatwaves disrupt pollen viability, destabilize membrane integrity, but accelerate phenology, leading to disruptive reproductive development and therefore irreversible yield penalties, even after short exposure periods [3,4,5,6]. Prolonged waterlogging induces hypoxia/anoxia, limiting mitochondrial respiration and energy availability [7,8], also causing elemental toxicities in plants [9]. Soil salinity is arguably most complex and adversely effects plant growth by causing Na+/K+ disequilibrium and imposing osmotic and oxidative stresses [10]. Nutrient deficiencies, particularly those for phosphorus (P) and nitrogen (N), restrict root development, nutrient acquisition, and photosynthetic capacity [11,12,13]. Underscoring the urgent need for genetically resilient crop varieties that maintain productivity across heterogeneous and stress-prone environments is therefore important [14,15].
Conventional breeding relies heavily on recurrent crossings followed by phenotypic selection across segregating generations and environments. Despite its historical success, this breeding progress is inherently slow, particularly in self-pollinated cereal crops, due to the cumulative duration of crossing, segregation, multi-generation selection, and environmental validation, resulting in the lengthy time (8–10 years or longer) to release a new cultivar [16]. Under climate change, such timelines risk delivering cultivars that are already mismatched to their target environments at the time of release. Second, phenotypic selection for abiotic stress tolerance is intrinsically imprecise and resource intensive. Many stress tolerance traits are quantitative, highly environment-dependent, and strongly influenced by genotype × environment × management (G × E × M) interactions [17]. Reliable phenotyping often requires careful validations across multiple environments and seasons, substantially increasing costs. In practice, however, current phenotyping often relies on simplified or proxy traits that may not capture the physiological complexity of stress tolerance under real field conditions, thereby weakening genotype-to-phenotype translation [17]. For example, reduced shoot Na+ concentration was chased as a primary selection criterion for salinity tolerance; however, this approach inherently favours Na+ exclusion mechanisms while overlooking the tolerance mechanism induced by intracellular Na+ sequestration to maintain metabolic function and yield in crops [7,18]. Consequently, varieties with superior sequestration capacity by locking higher shoot Na+ levels may be prematurely discarded during selection, which has biased breeding outcomes and partially led to the limited success in developing high-yielding, salt-tolerant rice cultivars despite decades of intensive selection. Further, introgression of favourable loci through conventional backcrossing often co-transfers undesirable genomic regions (genetic linkage) that may negatively affect yield. Introgression of the Saltol locus to improve salinity tolerance in rice reduced yield [19]. Similarly, the mlo resistance allele for powdery mildew resistance in barley was initially associated with yield penalties and pleiotropic effects due to linked genomic regions [20]. These cases provide good examples of the trade-offs that can accompany resistance or stress-tolerance breeding. This challenge is especially pronounced for traits governed by loci located in low-recombination regions (e.g., pericentromeric regions) of the chromosome. For example, the introgression of SR2 (for wheat rust resistance) has unintentionally fixed haplotypes in a low-recombination region, resulting in linkage drag that constrained root system development, necessitating additional breeding cycles/efforts to restore elite performance [21,22].
The limitations of conventional and molecular breeding approaches reveal a fundamental gap in current crop improvement pipelines: the lack of an integrated system that simultaneously delivers speed, genetic precision, and consistent field-level impact. A transformative breeding solution is therefore required, one that couples molecular precision with accelerated breeding cycles, while maintaining strong alignment with farmer needs and target populations of environments (TPEs). Fast backcross (FB) framework seeks to overcome the limitations by integrating marker-assisted backcrossing (MABC) for precise introgression of major-effect loci, rapid generation advancement (RGA) to compress breeding timelines, doubled haploid (DH) technology to fix favourable alleles rapidly, and genomic selection (GS) to capture polygenic adaptation and improve selection accuracy for complex traits [23,24]. In this review, we synthesize recent methodological advances, practical case studies, and emerging best practices to address these challenges. By critically evaluating how FB pipelines have been implemented across crops and stress contexts, we aim to identify design principles that enable rapid yet reliable delivery of climate-resilient cultivars, bridging the long-standing divide between laboratory innovation and real-world agricultural gains.
2. The Fast Backcross Breeding Framework Targeting 3–5 Years: Four Core Components with Optimizations
2.1. Rapid Generation Advance (RGA) and Speed Breeding
RGA is most achieved through speed breeding, which accelerates breeding cycles by shortening generation time rather than altering genetic processes [25]. By enabling multiple rounds of crossing and selection within a single year, speed breeding transforms traditionally sequential programs into near-continuous breeding pipelines [26]. In long-day crops such as wheat and barley, speed breeding typically relies on extended photoperiods (~22 h light/2 h dark) combined with controlled temperature and nutrition to accelerate flowering and seed set. Generation time can be reduced to 6–8 weeks without compromising plant development, fertility, or seed viability [25], and subsequent studies have shown it to be reproducible and compatible with breeding, gene discovery, and the production of segregating, doubled-haploid, and backcross populations [25,26,27].
A key operational advantage of RGA is its compatibility with single-seed descent (SSD), which enables efficient advancement of large populations under speed breeding while minimizing labour and space requirements and preserving genetic diversity. Watson et al. (2018) explicitly highlighted SSD as a key, particularly when integrated with high-throughput genotyping, as central to scaling speed breeding for practical breeding programs [25].
Effective RGA implementation emphases sufficient daily light integral and spectral coverage of photosynthetically active radiation, rather than on rigid optimisation of specific wavelength ratios. Practical protocols using LED-based and supplemental lighting systems in growth chambers and glasshouses have demonstrated that flexible lighting configurations can reliably support rapid growth and early flowering under extended photoperiods [26].
For short-day crops such as rice, RGA can be achieved through greenhouse-based rapid cycling and nursery systems with optimised sowing density, temperature control, and early harvest, along with light-emitting diodes (LED) systems, routinely allowing three to five generations per year. Together, these examples illustrate that RGA principles are transferable across crop species when photoperiod sensitivity and developmental requirements are appropriately considered [26,28,29].
2.2. Embryo Culture: Accelerating Generation Turnover and Enabling Cross Compatibility
Within fast breeding pipelines, embryo culture (EC) primarily accelerates generation turnover by shortening the interval between successive selection and crossing steps. In cereals, EC involves isolating and culturing young embryos (e.g., 14–20 days after flowering) on basal media, which can reduce germination time by 2–4 weeks relative to soil-based germination. Immature embryo rescue (IER) enables recovery of otherwise non-viable hybrids, particularly from interspecific or subspecific crosses used to access stress-adaptive alleles [27,30], most commonly Murashige and Skoog (MS) salts supplemented with sucrose as a carbon source [31], allowing germination and shoot emergence typically occur within 7–14 days [27], compared with 21–30 days under conventional conditions. This time saving is particularly valuable in backcross and selfing schemes, where delays due to seed drying, dormancy release, and field or glasshouse establishment otherwise constrain breeding speed, while maintaining genetic fidelity and phenotypic normality [27].
IER plays a complementary role by rescuing embryos that abort due to post-zygotic incompatibility or incomplete endosperm development [30,32], a common limitation in introgressing stress-tolerance alleles from landraces or wild relatives. Embryos typically excised 7–14 days after pollination (DAP), approximately 0.5–2 mm in length, and are still developmentally plastic [27], are cultured on nutrient-rich media to support continued development in vitro, thereby expanding the range of usable crosses without altering genetic selection criteria [33].
2.3. Marker-Assisted Backcrossing (MABC): Theory and Practice
2.3.1. Foreground Selection (FS)
Marker-assisted backcrossing (MABC) is a core precision breeding strategy designed to introgress defined genes or quantitative trait loci (QTLs) into elite genetic backgrounds while minimizing linkage drag and phenotypic uncertainty [34]. Among its three operational components (e.g., foreground, recombinant, and background selection), foreground selection (FS) represents the initial step, ensuring that only individuals carrying the target locus are advanced through successive backcross generations [34,35]. Several general principles exist for FS. First, flanking-marker strategies are preferred over single internal markers, as they reduce false positives arising from recombination events between the marker and the gene [36]. Second, FS should be applied at every backcross generation to ensure the target allele is retained during recurrent parent recovery. Third, FS is particularly powerful for large-effect loci with well-established genotype-phenotype relations, making it an ideal entry point for fast backcross breeding pipelines.
Foreground selection relies on molecular markers tightly linked to, or embedded within, the target genes or QTL, allowing unambiguous identification of favourable alleles regardless of environmental or developmental variation. This is particularly important for abiotic stress tolerance traits, which are often conditional, quantitative, and strongly influenced by G × E × M [37]. By decoupling selection from phenotype, FS dramatically increases selection efficiency and reduces population sizes required at each backcross generation [38]. For instance, the SUB1 region, a ~1.5 Mb on chromosome 9 containing SUB1A, which confers submergence tolerance through elongation growth suppression and carbohydrate reserves conservation [39]. Using flanking simple sequence repeat (SSR) markers (such as RM20584 and RM20585), Xu et al. [39] demonstrated precise stacking of SUB1A across backcross generations, while enabling downstream recombination to reduce donor genome segments.
2.3.2. Background Selection (BGS)
Background selection (BGS) is used to accelerate recovery of the recurrent parent genome (RPG) while retaining the donor allele at the target locus. In a conventional backcross program without marker-assisted background selection, the expected proportion of recurrent parent genome after t backcrosses follows Mendelian expectations:
|
```latexE\left(RPG, \%\right) = \left\{1 - 2^{-(t+1)}\right\} \times 100\%``` |
resulting in approximately 75% RPG in BC1, 87.5% in BC2, 93.75% in BC3, and 96.88% in BC4 [34,35]. Therefore, traditional backcross programs typically require four or more to approach the ≥95–98% RPG levels for elite cultivar development. Marker-assisted background selection uses genome-wide molecular markers to quantify RPG directly in each backcross generation and preferentially selects individuals that most closely resemble the recurrent parent outside the target locus. Modelling and simulation studies have demonstrated that genome-wide BGS can recover ≥95% RPG by BC3, and in some cases even by BC2, thereby saving one or more backcross generations relative to Mendelian expectation [40,41].
The efficiency of BGS depends on marker density, genome size, recombination landscape, and population size, parameters can be optimized through simulation. The software PLABSIM was developed to model marker-assisted backcrossing and evaluate marker densities and selection intensities influence RPG recovery and donor segment length [41]. These simulations showed that moderate population sizes (typically 100–200 individuals per generation) combined with evenly distributed genome-wide markers enable rapid RPG recovery without excessive genotyping costs. Subsequent analyses confirmed that background selection is most effective when applied from BC1 onward, rather than being deferred to later generations [42].
In practice, BGS involves genotyping all candidate plants that pass foreground selection using high-density, genome-wide markers (e.g., SNP arrays, KASP markers, or genotyping-by-sequencing), followed by estimation of recurrent parent genome content, typically based on identity-by-state similarity to the recurrent parent. A protected window around the target locus is excluded to retain the donor haplotype, and individuals with large residual donor segments elsewhere in the genome are deprioritised to reduce linkage drag that can affect yield, quality, or adaptation [34,35].
Operationally, BGS provides clear stop rules for ending backcrossing and proceeding to fixation by selfing or doubled haploids. Typical criteria include achieving ≥95–98% RPG genome-wide (excluding a small, protected window around the target locus), removing large donor segments elsewhere in the genome, and confirming that the donor interval flanking the target gene or QTL has been minimised. Once these conditions are met, further backcrossing yields diminishing returns and delays downstream validation.
Within the FB, BGS converts temporal acceleration into genetic efficiency per unit time. When combined with rapid generation advance and embryo culture, genome-wide BGS ensures that each accelerated generation contributes measurably to elite genome recovery, rather than simply shortening the calendar. This integration is a key reason FB pipelines to reliably deliver near-elite lines within three backcross generations, compared with four or more generations under conventional schemes [35,41].
2.3.3. Marker Set Design, Sampling Intensity, and Scoring in Background Selection
Simulation and empirical studies indicate effective BGS requires well-distributed marker density rather than maximal coverage [43]. A common guideline is to deploy approximately one informative marker per 5–10 cM across the genome [35], with higher density near the target chromosome and in low-recombination regions such as centromeres, where donor segments are more likely to persist [36]. In compact genomes such as rice, ~1000–2000 evenly spaced SNPs are typically sufficient, whereas in larger and more complex genomes such as wheat or maize, 3000–5000 markers are often required to achieve comparable precision [34]. Beyond these ranges, gains diminish relative to genotyping cost, making balanced placement more important than sheer density.
Population size is the second key determinant of BGS efficiency, as it affects the likelihood of recovering individuals with high recurrent parent genome content and favourable recombination. Both simulation and breeding practice support a declining population strategy across backcross generations, typically screening 200–400 individuals at BC1, 100–200 at BC2, and 50–100 at BC3, with increasing selection stringency over time [34,41]. Larger early populations increase the likelihood of identifying individuals with both high genome-wide RPG and minimal donor segments outside the target region, thereby accelerating convergence toward the recurrent parent background.
Accurate scoring of RPG recovery is essential for ranking candidates within each generation. RPG is commonly calculated as identity-by-state (IBS) similarity between each individual and the recurrent parent across genome-wide markers, providing a quantitative estimate of background recovery [34,44]. When ranking individuals, breeders should mask a protected window surrounding the target locus, typically spanning ±10–20 cM in early backcross generations and shrinking thereafter. This practice prevents inadvertent negative selection against the donor haplotype that is intentionally retained and ensures that RPG estimates reflect background recovery rather than recombination events at the target locus [35].
Together, optimized marker density, population size, and scoring convert BGS into a predictable, decision driven process, enabling near elite background recovery within two to three backcross generations, an outcome that is especially critical in fast backcross pipelines where accelerated cycling magnifies both gains and errors.
2.3.4. Where It Plugs into the Pipeline: Integration of Background and Recombinant Selection within the Fast Backcross Pipeline
Within the fast backcross (FB) framework, whole-genome marker-assisted background selection (WG-MABS) is most effective when initiated early and applied continuously rather than used as a late corrective stage. Accordingly, WG-MABS is activated immediately after confirmation of the target allele, typically at the F1 or BC1 generation following foreground selection [34,35].
During Stage B (“cross and compress”), WG-MABS is combined with rapid generation advance to maximise recurrent parent genome recovery while retaining the target locus. Initiating background selection at BC1 exploits the greatest available genetic variance, enabling large gains per generation that are efficiently translated into gains per unit time under speed breeding conditions [34,41]. Foreground selection ensures retention of the target haplotype, while genome-wide markers are used to rank individuals by recurrent parent genome content [25]. As the pipeline progresses through BC2 and BC3, selection intensity increases and recombinant selection is explicitly integrated with WG-MABS using flanking markers to progressively shorten the donor segment around the target locus, thereby minimising linkage drag while continuing genome-wide background recovery [34]. In crops such as maize and wheat, FB pipelines often transition to doubled haploid (DH) production once recurrent parent genome recovery exceeds ~95–98% and the donor interval has been minimised, allowing immediate fixation of the optimized genetic background [35].
The pipeline then enters Stage C (“fix and validate”), where target haplotypes and background recovery are re-verified in BC3F2 families or DH lines prior to stress-specific phenotyping and multi-environment trials (METs) aligned with the target population of environments (TPE), ensuring that accelerated genetic gains translate into stable yield, quality, and agronomic performance under both stress and non-stress conditions [17,45].
2.3.5. Expected Gains and Common Pitfalls
When integrated into fast backcross pipelines, whole-genome background selection typically saves one full backcross generation, enabling ≥95–98% recurrent parent genome recovery by BC3 rather than BC4, without increasing linkage drag when combined with deliberate recombinant selection at the target locus [34,35,41]. This translates to one to two years of time savings, depending on crop and generation turnover.
However, accelerated background selection carries risks. Beneficial donor alleles outside the primary target region may be inadvertently lost, particularly when pyramiding schemes, a risk that can be mitigated by defining additional protected genomic windows during selection [34,36]. A second pitfall is overestimation of recurrent genome recovery due to sparse or uneven marker coverage; robust genome-wide marker distribution and stringent quality control are therefore essential to avoid premature termination of backcrossing [35,40,44].
3. Case Studies: Success Stories in FB Breeding
FB frame has produced several clear translational wins where rapid generation advance and marker-assisted backcrossing converted elite cultivars into stress-resilient versions while maintaining agronomic performance, but it also highlights when “physiologically attractive” loci deliver limited field value. The benchmark case is SUB1 in rice: mapping and cloning resolved SUB1A as an ERF regulator conferring submergence tolerance via a quiescence strategy [39], and field evaluations consistently showed that Sub1 introgression enabled substantial yield recovery after 10–14 days of complete submergence, often restoring 50–70% of yield relative to non-submerged controls, while maintaining agronomic performance under non-flooded conditions [46,47]. Similar FB logic has been effective for Pup1/PSTOL1 under low P (phosphorus) [48]. Pup1 was successfully incorporated into high-yielding germplasm, leading to the development of two irrigated rice varieties and three Indonesian upland varieties [49], which retained elite agronomic characteristics while expressing enhanced P-use efficiency. And for DRO1, Uga et al. [50] demonstrated that introgression of the functional DRO1 allele from the deep-rooting landrace Kinandang Patong into the elite but shallow-rooting cultivar IR64 resulted in markedly steeper root angles and enhanced root penetration into deeper soil layers [50], where NIL-based IR64 improved drought-season yield (up to ~30%) without significant yield costs under well-watered conditions [50]. In contrast, wheat Nax1/Nax2 (from Triticum monococcum) illustrate the limits of Na+-exclusion targets: it is to note that although these loci (linked to HKT-type xylem Na+ retrieval) can reduce leaf Na+ substantially [51,52], field benefits can occur only within a narrow salinity window and remain insufficient to prevent large yield losses in many saline environments [52,53], demonstrating partial trade-offs [54]. Collectively, these cases argue that FB succeeds when loci deliver yield-relevant stress advantages with agronomic neutrality across representative target environments, ideally validated via NIL contrasts and multi-site testing, and that “marker-trackable” mechanisms alone are not adequate justification for rapid deployment.
Specifically, waterlogging and soil acidity frequently co-occur in high-rainfall environments, imposing constraints on crop root function through hypoxia and aluminium (Al3+) toxicity. Genetic studies in barley have identified two major loci on chromosome 4H: a waterlogging-tolerance QTL (QTL_AER-4H) controlling root aerenchyma formation associated with improved internal oxygen diffusion and performance under waterlogged conditions [55], and the acid-soil tolerance locus HvAACT1 (HvMATE), which mediates citrate efflux from root apices to alleviate Al3+ toxicity [56,57,58]. Using the proposed fast backcross (FB) framework, the waterlogging tolerance locus QTL_AER-4H was successfully introgressed into a commercial variety, RGT Planet, resulting in the derived variety RGT Atlantis. Under non-waterlogged conditions, RGT Atlantis exhibits yield and grain quality comparable to RGT Planet, while achieving more than a 20% yield advantage under waterlogged conditions. In parallel, the acid soil tolerance locus HvAACT1 was introgressed into RGT Planet to generate the breeding line P33-1, which displayed yields comparable to or exceeding those of RGT Planet on non-acidic soils, together with markedly improved performance under acidic soil conditions [59]. Subsequent crossing of RGT Atlantis with P33-1 enabled pyramiding of waterlogging tolerance (QTL_AER-4H) and acid soil tolerance (HvAACT1). The resulting line, P80-2, combined tolerance to both stresses while maintaining the yield and grain quality of the recurrent parent under non-stress environments (Figure 1).
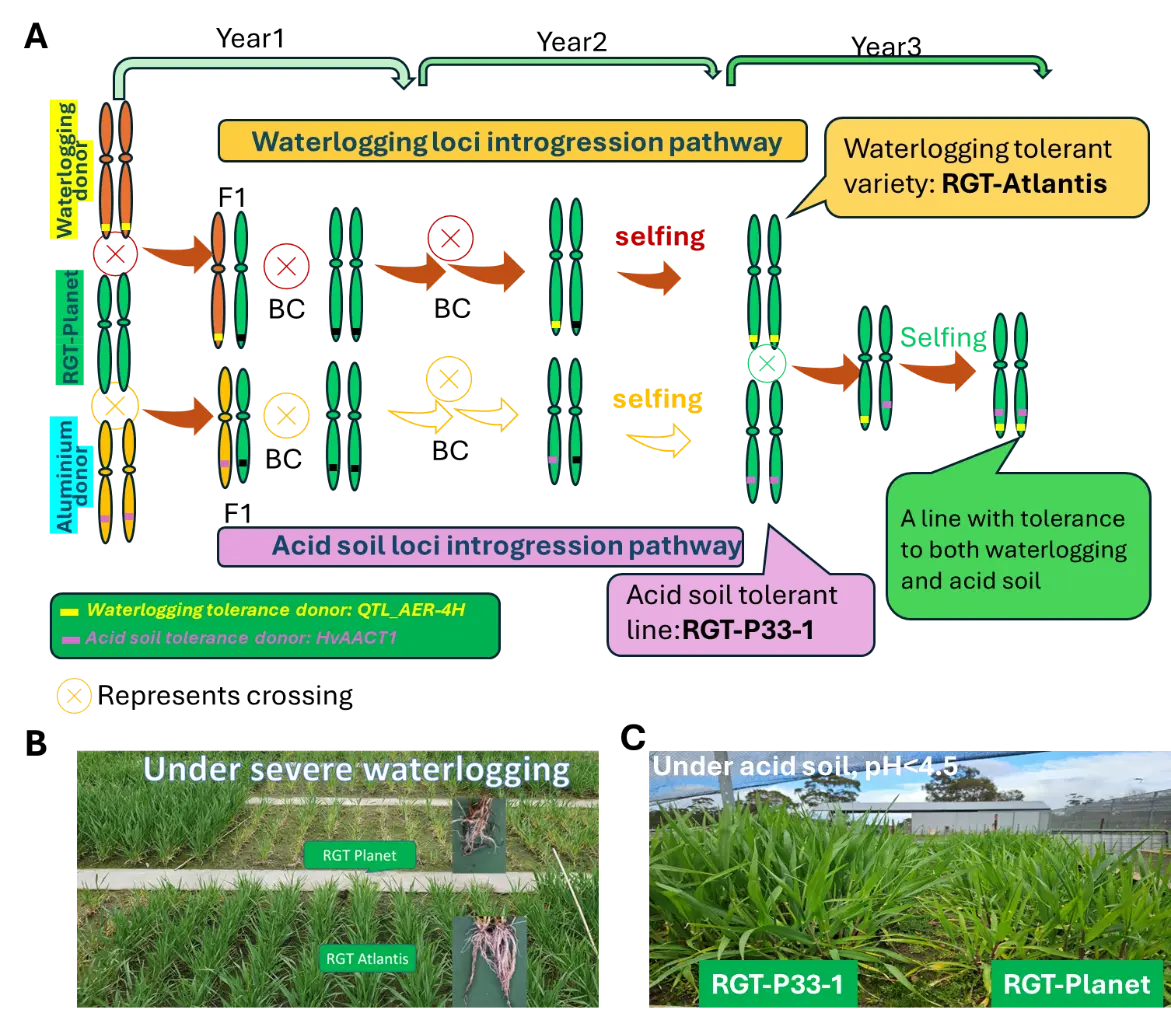
Figure 1. Pyramiding of waterlogging- and acid soil–tolerant loci into the elite cultivar RGT-Planet using the FB. (A) Schematic of breeding pipeline enabling the sequential introgression and pyramiding of two independent stress-tolerance loci into the elite barley cultivar RGT Planet within three years. A waterlogging-tolerant gene donor (TAM7227, wild barley) and an acid soil (aluminium) tolerant gene donor (Brindabella) were each crossed with RGT Planet (Australian elite cultivar), followed by repeated backcrossing (BC) combined with rapid generation advance (3–5 generations per year and whole genome selection, resulting in one waterlogging tolerant variety, RGT Atlantis, one acid soil tolerant line, P33-1 and one line with tolerance to both stresses. (B) Representative field performance of derived varieties under severe waterlogging conditions; (C) Representative field performance of derived varieties under acid soil conditions (pH < 4.5).
4. Complexity of Abiotic Stress Tolerance and Genes, QTLs, and Alleles with Minimal Yield and Quality Penalty
Abiotic stress tolerance in crops is inherently complex, reflecting the integration of multiple molecular, physiological, and developmental processes that operate across spatial and temporal scales. Unlike monogenic traits, tolerance to stresses such as drought, heat, salinity, nutrient deficiency, and submergence is typically governed by polygenic loci that collectively determine plant performance [17,60].
4.1. Combined Mechanisms Underpinning Abiotic Stress Tolerance
Biological defence mechanisms underlying abiotic stress tolerance are usually mediated by osmotic adjustment, ion and redox homeostasis, and hormonal signalling, which are tightly coupled and rarely act independently, particularly in drought and salinity tolerance [61,62]. For example, osmotic adjustment is a central component of tolerance to drought, salinity, and cold stress. Plants accumulate compatible solutes, such as proline and soluble sugars (e.g., sucrose, trehalose), which help maintain cell turgor, stabilize proteins and membranes, and protect cellular structures under water deficit or osmotic stress induced by cold [63]. Salinity tolerance depends on the plant’s capacity to limit cytosolic Na+ accumulation while maintaining adequate K+ levels for enzyme activity and membrane potential regulation [7,18,64]. This balance is achieved through coordinated activities of ion transporters, channels, and compartmentation mechanisms, including Na+ exclusion at the root-soil interface and vacuolar sequestration in shoot tissues [7,18].
Hormonal regulation coordinates physiological, molecular, and developmental adjustments. For instance, abscisic acid (ABA) is a key regulator of drought and salinity tolerance, mediating stomatal closure [65,66], root system architecture [67], and stress-responsive gene expression [68]. Ethylene signalling is central to submergence and waterlogging tolerance, particularly through its role in regulating shoot elongation [69,70], aerenchyma formation [71], and metabolic acclimation under hypoxic conditions [72]. Importantly, hormonal signalling pathways do not operate in isolation but interact extensively with other regulatory networks, including reactive oxygen species (ROS), calcium signalling, and transcriptional control, adding further layers of complexity [73]. To add more complexity to this process, ROS and Ca2+ signalling are tightly interconnected by the presence of so-called NADPH-Ca “hub” at the plasma membrane [74] that amplifies both signals to trigger a cascade of downstream responses.
Morphological and developmental adaptations provide longer-term strategies for coping with abiotic stress. Examples include deeper or more plastic root systems to enhance water and nutrient acquisition under drought or low-phosphorus conditions [75,76], reduced plant stature or altered phenology to escape heat stress [77], and enhanced aerenchyma formation to facilitate oxygen diffusion under waterlogging [78]. These traits are often highly environment-specific and developmentally regulated [17], contributing strongly to genotype by environment interactions observed in field trials. From a genetic perspective, these molecular and physiological mechanisms are controlled by multiple quantitative trait loci (QTLs) with small to moderate individual effects rather than single major genes [17]. Many QTLs exhibit pleiotropy or context-dependent expression, and their effects can vary across environments, genetic backgrounds, and developmental stages. Consequently, introgression of abiotic stress tolerance through conventional breeding is challenging, as favourable alleles must be assembled and maintained across multiple loci while avoiding negative impacts on yield and adaptation.
Abiotic stress tolerance ultimately arises from allelic variation at key loci, where specific alleles or allele combinations confer superior physiological advantages under stress conditions. In contrast to simple presence-absence of genes, natural variation in coding sequences, regulatory regions, and gene expression dynamics often determines the magnitude and stability of stress tolerance across environments [14,79]. Classic examples illustrate the power of large-effect adaptive alleles, for instance, as in rice, the SUBMERGENCE 1A (SUB1A) allele confers strong tolerance to transient submergence by suppressing excessive shoot elongation, reducing carbohydrate consumption, and enhancing post-submergence recovery [8,39]. Introgression of SUB1A into elite rice cultivars through marker-assisted backcrossing has resulted in varieties with minimal yield penalty under non-flooded conditions [39], demonstrating the successful translation of gene discovery into farmer impact. However, such examples remain relatively rare, as most abiotic stress traits lack single dominant loci with comparable effect sizes. For most stresses, including drought, heat, salinity, and nutrient deficiency, variation is distributed across multiple loci, each contributing incrementally to the phenotype. In these cases, haplotypes, defined as specific combinations of linked alleles across a genomic region, often provide a more accurate representation of functional genetic variation than individual markers [80,81]. Haplotype structure captures both causal variants and their local linkage context, enabling finer discrimination between favourable and unfavourable genetic backgrounds [81].
Emergence and application of genome-wide association studies (GWAS) have accelerated the identification of stress-associated loci and haplotypes across diverse germplasm panels [82]. GWAS-based analyses have revealed that favourable haplotypes often differ in frequency among landraces [83,84], modern cultivars [83], and wild relatives, reflecting historical selection pressures and breeding bottlenecks [85]. Importantly, haplotype-based analyses frequently explain a greater proportion of phenotypic variance than single-SNP models, because haplotypes integrate the combined effects of multiple linked variants and more effectively capture local linkage disequilibrium, regulatory variation, and allele interactions underlying complex traits, as particularly demonstrated in barley for complex traits influenced by regulatory variation and gene-gene interactions [86]. From a breeding perspective, haplotype-based selection offers several advantages over single-marker approaches. It improves selection precision by reducing false positives arising from weak marker-trait associations [87,88]. Also, it facilitates the stacking of favourable alleles within genomic regions while minimizing linkage drag, particularly when combined with recombinant selection strategies. Third, haplotypes provide a natural interface with genomic selection, where genome-wide marker information is used to predict breeding values that implicitly capture favourable allele combinations across loci [88]. The effects of many haplotypes are context-dependent, varying across environments, developmental stages, and genetic backgrounds, and are often modulated by G × E × M interactions [37].
4.2. Energetic Costs, Pleiotropy Co-Benefits, and Trade-Offs on Yield: A Case Study for Salinity
Understanding and managing trade-offs, e.g., yield penalties, is central to the successful deployment of stress-tolerant alleles in elite cultivars. However, when energetic costs and pleiotropic effects are well aligned with prevailing stress conditions, such alleles can also confer co-benefits by improving yield stability or performance under stress.
Salinity tolerance illustrates the energetic trade-offs and pleiotropic constraints associated with stress adaptive mechanisms. Na+ exclusion from the shoot, mediated by HKT1;5-type Na+ transporters, has historically been used as a main target for improved crop survival [51,89]. However, this process is energetically costly, as such a strategy requires plants to heavily invest in the production of compatible organic solutes to osmotically adjust to hypersaline soil conditions—a process that requires 10-fold more ATP than vacuolar Na+ sequestration [53]. Therefore, constitutive or excessive activation of Na+ exclusion pathways imposes growth penalties under non-saline or moderately saline conditions, highlighting important trade-offs between tolerance and productivity.
Vacuolar sequestration of Na+ represents an alternative or complementary strategy, allowing plants to use Na+ as a “cheap osmoticum”. This process is mediated by tonoplast based Na+/H+ exchangers energized by vacuolar proton pumps, such as H+-ATPases and H+-pyrophosphatases (PPases) [90]. As ATP pool is drastically reduced due to negative impact of salinity on leaf photochemistry [91] and NaCl-induced stomata closure. The H+-PPase AVP1, however, has potential for offering a more energy-efficient means of sustaining ion compartmentation. Field and physiological studies demonstrated that enhanced AVP1 activity can improve salinity tolerance and, in some cases, biomass production [92]. However, AVP1 is a multifunctional protein involved in growth, development, and phloem loading, and both overexpression and misregulation have been associated with pleiotropic effects under non-stress conditions [92,93]. Recent evidence for dynamic regulation and dual localization of AVP1 further suggests that spatial and temporal control of activity is critical for balancing tolerance and growth [92].
Pleiotropy at the developmental level represents another major source of yield penalty. A prominent example is the rice gene DRO1, which enhances drought tolerance by promoting deeper root growth [50]. While advantageous under water-limited conditions, DRO1 introgression has been shown to reduce harvest index and yield under well-watered environments due to altered biomass allocation [50]. Recent advances suggest that regulated, rather than constitutive, stress tolerance mechanisms may offer a pathway to minimizing yield penalties. For example, natural variation at regulatory loci such as TaSPL6-D, which modulates expression of TaHKT1;5-D, allows fine-tuning of Na+ exclusion capacity in wheat [94]. Such allelic variants provide sufficient exclusion under salinity stress while avoiding excessive energetic costs under low-salinity conditions, highlighting the importance of quantitative regulation rather than maximal flux [95]. From a breeding perspective, these insights underscore a critical challenge: maximizing stress tolerance without sacrificing yield stability across environments.
Many stress-associated loci identified through mapping or transcriptomic studies fail at later breeding stages because their effects are either context-dependent, energetically costly, pleiotropic [17], or not even evaluated in situ within typical cropping systems [96]. We propose a structured approach centred on the development and evaluation of near-isogenic lines (NILs) in elite genetic backgrounds, combined with distinct phenotypic contrasts across stress and non-stress environments, allowing causal attribution of phenotypic effects to specific loci or haplotypes, while minimizing confounding effects from background variation.
5. Genetic Validation and Agronomic Relevance of Stress Tolerance
5.1. Technical Approaches: NIL-Based Validation Using Distinct Genetic Contrasts
NILs are generated by introgressing a target gene or QTL into an elite recurrent parent through marker-assisted backcrossing, followed by selfing or doubled haploid production [97]. Foreground markers ensure retention of the target locus, while genome-wide background selection accelerates recovery of the recurrent parent genome, typically achieving >95% recurrent parent genome (RPG) within two to three backcross generations [97,98]. NILs provide a clean genetic contrast, enabling direct comparison between lines that differ primarily at the locus of interest. This is particularly critical for abiotic stress traits, where background genetic noise and genotype × environment interactions can otherwise obscure true allelic effects [37]. To quantify the yield penalties, paired NILs are evaluated across multi-environment trials (METs) that deliberately span both benign (non-stress) and stress-prone conditions (e.g., salinity, drought, heat, and submergence), enabling rigorous quantification of the net agronomic value of a target locus by simultaneously assessing its contribution to stress resilience and its impact on performance under normal conditions. Assessments of both agronomic and phenological traits, including flowering time, plant height, and tillering dynamics, which can indirectly influence yield stability, adaptability, and farmer acceptance, should be carefully conducted [17,99]. Furthermore, stress-inducible, tissue-specific, or developmental-stage-specific alleles may reduce unnecessary energetic expenditure under non-stress conditions and minimize pleiotropic growth penalties [18,53]. Natural regulatory variation, such as promoter polymorphisms or transcriptional modulators, often provides this fine-tuning and is recognised as a rich source of breeding-relevant diversity [100,101].
5.2. Demonstrated Example of Low to No Cost Stress Tolerance
A flagship example for this case is the rice SUB1A locus, which confers submergence tolerance via a “quiescence” strategy that restricts shoot elongation, conserves carbohydrates during flooding, and promotes rapid recovery after de-submergence [47,102]. SUB1A has been introgressed into farmer adopted varieties (e.g., IR64, Swarna) with large post-flood yield benefits, while showing little or no yield penalty under non-submerged conditions critical for large-scale deployment [47,103].
Analogous “low-cost” exemplars are from drought stress, where rice qDTY loci controlling grain yield under reproductive-stage drought were identified. While pyramiding strategies can be effective, several studies show that evaluating component loci as NILs (or NIL-like contrasts) is essential, as individual qDTY regions differ in their ability to deliver stress yield gains without compromising performance under irrigated conditions [104,105]. This reinforces the broader principle that “no/low-cost” tolerance is most reliably identified when stress performance is explicitly benchmarked against non-stress yield stability across environments.
HKT1;5-mediated Na+ exclusion represents one of the most mechanistically resolved and agronomically validated tolerance mechanisms. In wheat, the major Na+-exclusion loci Nax2/Kna1 are strongly associated with HKT1;5-like transporters that retrieve Na+ from the xylem and reduce shoot Na+ accumulation, enabling substantial yield protection on saline soils [51,52]. However, as described above, energetic and regulatory analyses highlight that the yield neutrality of Na+ exclusion under non-saline conditions can be genetic background-dependent, particularly if exclusion is constitutively high rather than stress-regulated [53,95]. Also, field trials showed that beneficial effects of Nax were reported for only one (out of three) field sites [106]; even there the crop grain yield was ⁓50% of that in the non-saline control. This aligns with broader arguments that “cost-efficient” tolerance often comes from tuned or conditional activity of transport processes rather than maximal flux under all conditions [18,53].
Barley offers an additional example through the boron-toxicity tolerance gene HvBot1, where reduced boron accumulation in shoots improves performance on high-boron soils; when introgression is conducted carefully to minimize linkage drag, such ion-toxicity tolerance mechanisms can be deployed with limited penalty under benign conditions [107]. Collectively, these examples converge on a pragmatic lesson for FB: successful acceleration begins not with rapid cycling itself, but with prioritising loci that show a robust stress advantage while remaining agronomically neutral under favourable conditions, ideally confirmed through NIL contrasts and TPE-aligned multi-environment testing.
To operationalise this principle, loci can be “green-lit” for fast breeding deployment using transparent acceptance thresholds: a ≥10–20% grain yield gain under the target stress, combined with ≤±2–3% yield shift under non-stress environments across at least two seasons and/or representative sites. Equally, loci should show no unacceptable movement in key quality KPIs (e.g., grain protein, test weight, malt extract/enzymes in malting barley) and no undesirable phenology or height shifts unless these are intentional breeding targets. Finally, for prioritisation across multiple candidates, loci can be evaluated using a marginal trade-off framework, in which incremental gains in stress-associated yield are weighed against corresponding marginal penalties in yield under non-stress conditions; the optimal deployment point is reached when additional stress tolerance no longer delivers proportional yield stability or gain [53].
6. Pyramiding Complementary Mechanisms for Stable Gains
6.1. Design Rules for Pyramiding Complementary Stress-Tolerance Mechanisms
Simply stacking “stress loci” does not reliably translate into stable yield gains and can increase the risk of yield or quality penalties under benign conditions. This is unsurprising because most stress resilience traits are governed by complex genetic architectures, strong genotype × environment (G × E) interactions, and frequent trade-offs that become more visible when multiple loci are combined [17,37].
A first principle is to combine orthogonal mechanisms, meaning loci that operate through distinct physiological pathways and ideally act at different time points or tissue domains of the stress response. Examples include combining loci that improve ion homeostasis with loci conferring stress avoidance or quiescence (e.g., submergence quiescence strategies) and with reproductive-stage yield QTLs that stabilize grain set and grain filling under stress [39]. Such orthogonality reduces redundancy and increases the probability of additive or synergistic outcomes, while lowering the likelihood that the pyramid simply amplifies the energetic costs of a single pathway [17,37].
Further, pyramiding should be built from vetted NILs rather than directly stacking loci identified only through discovery mapping or GWAS. NIL-based validation provides cleaner causal inference and allows breeders to quantify the stress benefit relative to non-stress neutrality before loci are combined, which is essential for managing trade-offs and avoiding hidden penalties [17]. Operationally, this means progressing from validated NILs to NIL pyramids (typically two to four loci) and, only then, converting the best-performing pyramids into deployment pipelines using marker-assisted selection (MAS) and, where appropriate, genomic selection (GS) for background recovery and polygenic performance [24,103]. This staged approach is consistent with the broader need to align genetic gains with the target population of environments (TPE) rather than pursuing broad but untested generality.
Lastly, because multi-locus combinations frequently generate epistasis, pyramids must be evaluated explicitly for interaction effects on both yield and non-yield traits. Epistatic interactions can produce desirable synergy, but they can also lead to unexpected penalties in growth, phenology, or end-use quality that were not apparent in single-locus lines [108]. Consequently, pyramids should be assessed not only for yield stability across representative stress and non-stress environments, but also for quality neutrality, using crop-relevant panels (e.g., malting quality metrics in barley; baking/breadmaking traits in wheat) and grain ionomic analyses when the target loci are associated with ion transport or homeostasis, where unintended changes in elemental composition can occur. This emphasis on multi-trait validation reflects a central lesson from stress phenotyping: robust translation depends on testing the right traits, at the right developmental stage, across the right environments [17].
6.2. Potential Stacks: Complementary Pyramids (Representative Valuable Genetic Resources for Gene Pyramiding in Major Cereals)
Evidence from several crop systems demonstrates that carefully designed pyramids of complementary stress-tolerance loci can deliver larger and more stable yield gains than single-locus introgressions, provided that component loci are mechanistically orthogonal and individually validated for minimal penalty under non-stress conditions.
In rice, some of the clearest designs come from pyramids combining submergence, salinity, and drought tolerance mechanisms. The SUB1A locus provides tolerance to transient flooding through a quiescence strategy that conserves carbohydrates and supports post-submergence recovery [39,47], while the Saltol region enhances seedling-stage salinity tolerance by maintaining K+/Na+ homeostasis [19]. Pyramiding two loci with reproductive-stage drought yield QTLs, such as qDTY2.2, qDTY4.1 have shown improved yield stability across drought [104].
In wheat, pyramiding strategies have tended to be more site-specific, reflecting strong environmental heterogeneity and sensitivity of quality traits. A potential combination involves HKT1;5-mediated Na+ exclusion for salinity tolerance [51], together with TaALMT1 [109], which confers aluminium tolerance on acid soils through malate exudation. This pairing targets two co-occurring constraints in many rainfed systems, salinity and soil acidity, and has been shown to broaden adaptation when deployed in appropriate environments.
In barley, emerging evidence supports the value of pyramiding tolerance to waterlogging and acid soils, since anaerobic soil reduces soil redox potential, leading to soil elemental toxicity [70], including Al [110], which is also the primary toxicity from acid soils [59], stresses that frequently co-occur in high-rainfall production zones. Representative QTL loci underlying aerenchyma development conferring to waterlogging tolerance have been identified from a wild type of barley [55], multiple filed trials have validated its roles in waterlogging tolerance [111]. Meanwhile, a QTL on chromosome 4, named ‘QTL5’ has been identified [57], containing a HVACCT1 [58], responsible for producing citric acid, chelating Al alleviating acid soil toxicity. Combining these two traits will greatly alleviate both waterlogging and acid soil effects on crop growth. Below, we present the successful practice of breeding barley cultivars with these two traits, adopted by farmers in this timeframe.
6.3. Three Gates Validation for Stress-Tolerance Pyramids
Pyramiding amplifies both genetic gains and risks, multi-locus stacks must pass through validation gates before being adopted in fast-breeding pipelines or considered for variety release. These gates ensure that pyramids deliver genuine, stable agronomic value rather than short-term stress performance at the expense of yield stability, quality, or adaptation (Figure 2).
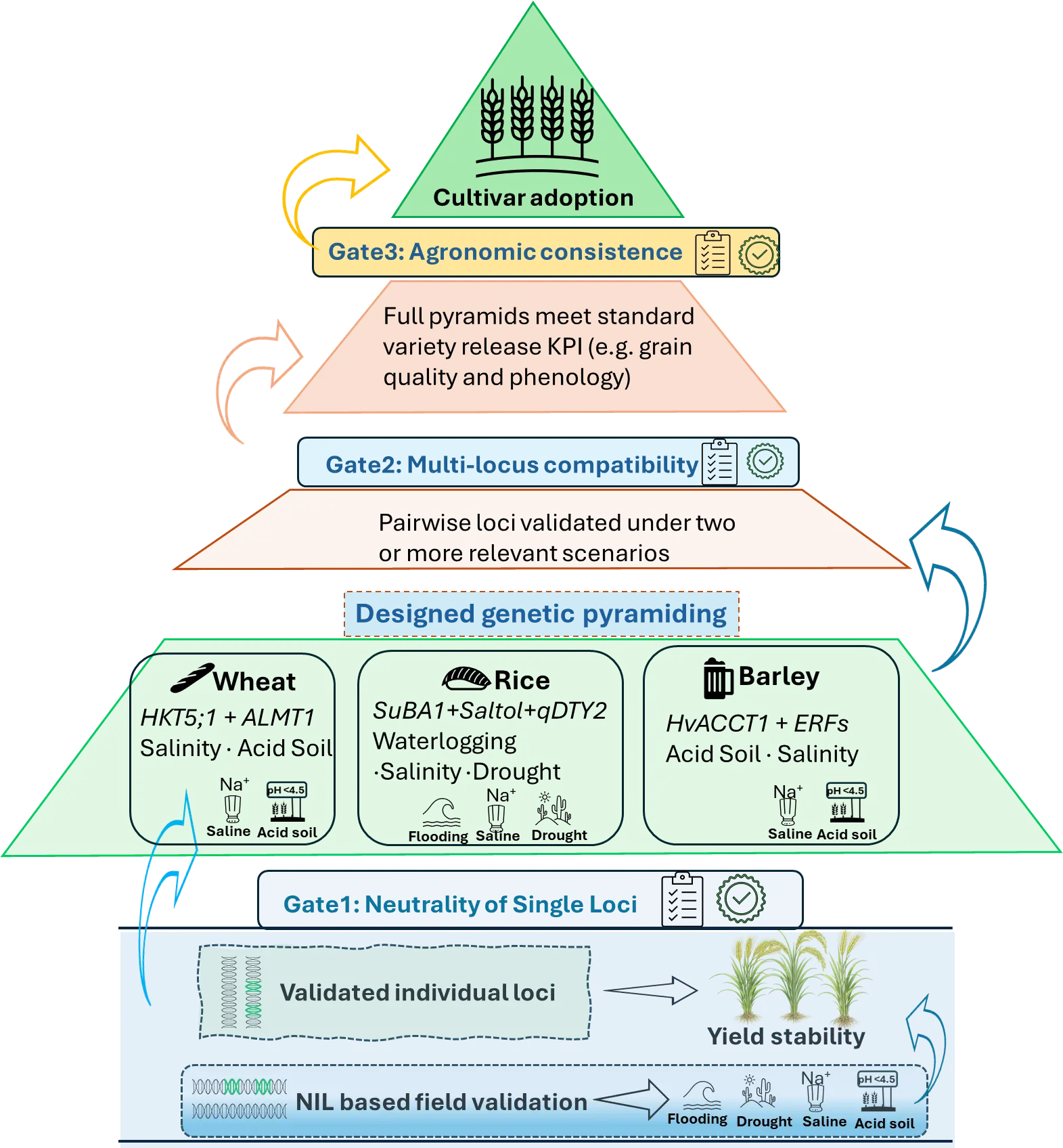
Figure 2. Genetic pyramiding framework for deploying validated loci for resilient cereal cultivars. Individual loci with demonstrated stress benefits are first confirmed through near-isogenic line (NIL) based field validation to ensure yield stability under both stress and non-stress conditions (Gate 1: neutrality of single loci). Validated loci are then combined through designed genetic pyramiding, where compatibility among multiple loci is evaluated to avoid negative epistasis or agronomic penalties (Gate 2: multi-locus compatibility under multiple stress scenarios). Crop-specific examples illustrate pyramiding strategies in wheat (e.g., HKT1;5 + ALMT1 for salinity and acid soils), rice (e.g., SUB1 + Saltol + qDTY2 for waterlogging, salinity, and drought), and barley (e.g., HvACCT1 + ERFs for acid soil and waterlogging tolerance). Only pyramids that achieve consistent agronomic performance and trait expression across environments advance to Gate 3, culminating in cultivar adoption. This gate-based framework operationalises genetic stacking by converting locus discovery into predictable, adoption-ready outcomes for resilient crop breeding.
The first gate requires that each component locus demonstrate neutrality under non-stress conditions when evaluated individually. This criterion reflects a central lesson from abiotic stress breeding: alleles that confer stress tolerance but impose constitutive growth or quality penalties rarely translate into farmer adoption [45], even if stress benefits are large. Validation at this stage is typically achieved using near-isogenic lines or clean contrasts in elite backgrounds, assessed across representative benign environments [97].
The second gate focuses on pairwise combinations of loci, which serve as an early test of genetic interaction and functional complementarity. At this stage, pyramids should demonstrate additive or complementary effects under at least two relevant stress scenarios, such as salinity plus drought, or waterlogging plus soil acidity. This requirement acknowledges that epistasis is common in complex traits and that favourable single-locus effects do not guarantee favourable multilocus outcomes [112]. Pairwise evaluation helps identify antagonistic interactions early, before additional loci are added, and provides mechanistic insight into whether combined pathways reinforce or interfere with one another [113].
Only pyramids that pass these initial gates should proceed to full stacking and advanced testing. The final gate requires that the complete pyramid meet standard variety-release key performance indicators (KPIs) across multi-environment trials aligned with the target population of environments (TPE). These KPIs extend beyond stress yield advantage and include yield stability under non-stress conditions, maintenance of end-use quality parameters (e.g., grain protein, malting or baking performance), and acceptable agronomic characteristics such as plant height, lodging resistance, and phenological fit. This holistic assessment reflects the reality that cultivar success depends on system-level performance rather than stress tolerance in isolation [17].
Importantly, these validation gates are not intended to slow breeding progress, but to prevent the accelerated advancement of suboptimal genetic combinations in fast-breeding pipelines. By enforcing neutrality at the component level, additivity at the pairwise level, and agronomic completeness at the final stack level, breeders can balance speed with reliability. This gate-based approach aligns pyramiding with modern breeding principles, where disciplined decision-making is essential to translate molecular innovation into durable, farmer-relevant outcomes.
7. Conclusions and Future Perspectives
7.1. Key Success Factors
Successful FB depends on three interlinked factors. First, explicit alignment with the target population of environments (TPEs) is foundational. Defining dominant production contexts (e.g., rainfed vs. irrigated, acidic high-rainfall soils vs. saline lowlands) constrains trait prioritisation, locus selection, and validation strategy, and avoids selecting traits that perform in managed trials but fail in farmers’ fields. Empirical and theoretical work show that matching selection environments to TPEs improves yield stability and adoption, particularly for abiotic stress traits with strong genotype × environment × management interactions [14,37]. The success of SUB1, Pup1, and DRO1 illustrates how FB pipelines anchored to clearly defined stress profiles can deliver robust gains under target conditions while remaining agronomically neutral elsewhere. Second, pipeline integration, rather than any single technology, underpins FB efficiency. Rapid generation advance (RGA) compresses time, marker-assisted backcrossing (MABC) ensures genetic precision [38], doubled haploids (DHs) enable rapid fixation [80], and genomic selection (GS) captures residual polygenic variation [114]. When orchestrated as a coherent system, these tools convert accelerated cycling into predictable genetic gain per unit time, whereas isolated deployment yields only incremental benefits [25,35,115]. Third, farmer engagement and product relevance are critical for impact at scale. Stress-tolerant alleles are most readily adopted when introgressed into farmer-preferred backgrounds that retain established agronomic and quality traits. Participatory testing, on-farm validation, and alignment with local management practices markedly increase uptake, as exemplified by the widespread adoption of SUB1 rice [17,45]. Collectively, these factors position FB breeding not as a faster version of conventional backcrossing but as a design-driven framework integrating environmental targeting, technological coordination, and stakeholder relevance.
7.2. Cost-Benefit Trade-Offs
The economic efficiency of FB breeding is strongly shaped by genetic architecture and deployment scale. For traits governed by large-effect loci, FB pipelines are highly cost-effective: MABC reduces population sizes and phenotyping demands, while RGA shortens program duration. Comparative analyses in rice indicate that integrating MABC with RGA can reduce total breeding costs by ~30–40% relative to conventional backcrossing, largely by eliminating one or more backcross generations and reducing field cycles [25,35]. These efficiencies are maximised when a validated locus is deployed across multiple elite backgrounds. In contrast, FB breeding becomes more resource-intensive for polygenic traits, where gains rely on GS rather than discrete loci. GS requires large training populations, extensive multi-environment phenotyping, and repeated genotyping, increasing upfront costs before returns are realised [17,37]. While justified for complex traits such as yield stability, GS is less efficient for narrowly defined stress traits with major loci. A pragmatic strategy therefore emerges: deploy FB with MABC to target large-effect loci first, and layer GS selectively where residual polygenic variation constrains performance. Strategic matching tools to trait architecture is essential to ensure that acceleration delivers both genetic and economic returns.
7.3. Limitations and Mitigations
Despite its advantages, FB breeding requires disciplined implementation to manage key risks. Linkage drag remains a persistent challenge, particularly when introgressing alleles from landraces or wild relatives [21,22]. However, rigorous recombinant selection using flanking markers, combined with genome-wide background selection (BGS), can substantially reduce donor segment size and eliminate residual donor blocks, mitigating unintended trait introgression [22,34,35]. A second limitation arises from G × E interactions, which are pronounced for abiotic stress tolerance traits. Gains observed under managed or single-location trials may not translate consistently across heterogeneous production systems [17,37,42]. Consequently, FB-derived lines must be validated through multi-environment trials spanning representative stress and non-stress conditions within the TPE, typically across 3–5 sites, to distinguish robust effects from context-specific responses [14,23]. Finally, the effectiveness of BGS depends on adequate marker density and genome coverage. Sparse or uneven marker sets can lead to overestimation of recurrent parent genome recovery and premature termination of backcrossing. Advances in high-throughput genotyping platforms, including SNP arrays and genotyping-by-sequencing, have largely alleviated this constraint by enabling dense genome-wide coverage at declining cost. When combined with appropriate quality control and sampling strategies, these tools enhance the reliability and predictability of FB outcomes.
7.4. Future Directions
Emerging technologies are poised to further amplify FB breeding by reducing reliance on recurrent crossing and improving decision quality. CRISPR-based genome editing offers a direct route to recreate favourable alleles in elite backgrounds, bypassing linkage drag and multiple backcross generations when causal variants are well defined [39,50]. In practice, genome editing is likely to complement rather than replace FB breeding, providing an efficient alternative where regulatory frameworks permit.
A second frontier lies in AI-assisted decision support, where machine-learning models integrate genomic, phenotypic, and environmental data to optimise cross design, marker deployment, and advancement strategies. Such approaches have already improved prediction accuracy for complex traits and resource allocation, particularly when combined with GS and high-throughput phenotyping [116,117].
Finally, extending FB breeding to orphan and underutilised crops, including sorghum and millets, offers substantial opportunities for climate resilience in semi-arid regions. The modular nature of FB pipelines makes them well suited for deployment as genomic resources expand, provided that investment in reference genomes, marker platforms, and participatory testing accompanies technological adoption [118]. Together, these directions suggest that the future of FB breeding will be shaped less by incremental acceleration and more by strategic integration, aligning genetics, analytics, and users’ needs to deliver resilient cultivars at scale.
7.5. Conclusions
FB framework (Figure 3) practically closes the gap between gene discovery and farmer adoption by integrating genetic precision, accelerated generation turnover, and disciplined deployment. Across crops and stress contexts, the cases reviewed here show that FB pipelines can convert validated loci into elite genetic backgrounds within ~3–5 years while limiting linkage drag and preserving yield and quality under non-stress conditions. The central conceptual advance is that FB functions as a design-driven framework rather than a collection of acceleration tools, where genetic gain per unit time rather than generations per year is the relevant metric of progress. This requires explicit alignment with target populations of environments (TPEs), early and continuous application of foreground, recombinant, and genome-wide background selection, and clear “green-light” criteria for locus prioritisation, favouring alleles that deliver yield-relevant stress benefits while remaining agronomically neutral under favourable conditions. The framework further shows that linkage drag is a controllable design variable, not an inevitable cost of speed, when donor segments are proactively minimised, and that validation and adoption are integral components of FB efficiency, achieved through TPE-aligned multi-environment testing and participatory evaluation. Finally, FB delivers its greatest impact when validated loci are deployed across multiple farmer-preferred backgrounds, enabling rapid scaling and cost amortisation. Together, these insights position FB breeding as a robust, scalable strategy for delivering timely and durable genetic gains as climate variability intensifies and demand for resilient crop varieties accelerates.
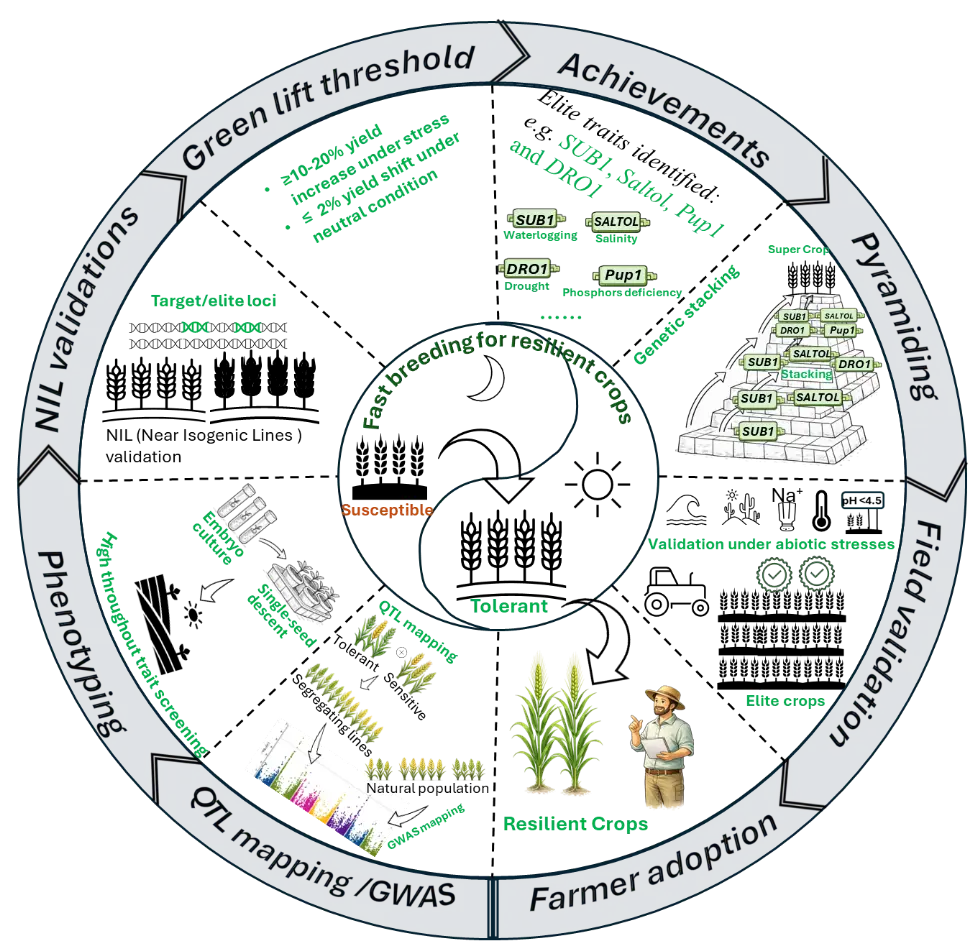
Figure 3. The conceptional framework of the proposed fast backcross (FB) breeding strategy for developing resilient crops. Elite loci identified through QTL mapping and genome-wide association studies (GWAS) are rapidly introgressed into elite genetic backgrounds using marker-assisted backcrossing, accelerated by embryo culture to enable rapid single-seed descent. The resulting near-isogenic lines (NILs) are evaluated across multiple environments under relevant abiotic stress conditions. Breeding success is defined by exceeding a “green lift” threshold, achieving substantial yield improvement under stress without penalties under non-stress conditions, thereby ensuring agronomic relevance. Subsequently, deliberate genetic pyramiding integrates complementary stress-tolerance mechanisms (e.g., waterlogging, drought, salinity, heat, and soil acidity). Multi-environment field validation under diverse abiotic stresses provides rigorous confirmation of locus effects while minimizing linkage drag. The framework culminates in farmer adoption and deployment of resilient cultivars, highlighting FB breeding as a systematic pipeline that bridges gene discovery and real-world impact, rather than merely accelerating conventional backcrossing.
Statement of the Use of Generative AI and AI-Assisted Technologies in the Writing Process
During the preparation of this manuscript, the authors used chatbot by OpenAI for English improvement. The authors reviewed and edited the content as needed and take full responsibility for the content of the published article.
Author Contributions
Conceptualization, M.Z.; Writing—Original Draft Preparation, C.Z.; Writing—Review & Editing, M.T.H., K.L., C.L., Z.C., S.S. and M.Z.; Funding Acquisition, M.Z.
Ethics Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Funding
This research was funded by the Grains Research & Development Corporation of Australia.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
-
Lesk C, Rowhani P, Ramankutty N. Influence of extreme weather disasters on global crop production. Nature 2016, 529, 84–87. DOI:10.1038/nature16467 [Google Scholar]
-
Flexas J, Carriquí M, Coopman RE, Gago J, Galmés J, Martorell S, et al. Stomatal and mesophyll conductances to CO2 in different plant groups: Underrated factors for predicting leaf photosynthesis responses to climate change? Plant Sci. 2014, 226, 41–48. DOI:10.1016/j.plantsci.2014.06.011 [Google Scholar]
-
Bita C, Gerats T. Plant tolerance to high temperature in a changing environment: Scientific fundamentals and production of heat stress-tolerant crops. Front. Plant Sci. 2013, 4, 273. DOI:10.3389/fpls.2013.00273 [Google Scholar]
-
Zhao C, Siddique AB, Guo C, Shabala S, Li C, Chen Z, et al. A high-throughput protocol for testing heat-stress tolerance in pollen. aBIOTECH 2025, 6, 63–71. DOI:10.1007/s42994-024-00183-3 [Google Scholar]
-
Zhao C, Liu B, Piao S, Wang X, Lobell DB, Huang Y, et al. Temperature increase reduces global yields of major crops in four independent estimates. Proc. Natl. Acad. Sci. USA 2017, 114, 9326–9331. DOI:10.1073/pnas.1701762114 [Google Scholar]
-
Muleke A, Harrison MT, de Voil P, Hunt I, Liu K, Yanotti M, et al. Earlier crop flowering caused by global warming alleviated by irrigation. Environ. Res. Lett. 2022, 17, 044032. DOI:10.1088/1748-9326/ac5a66 [Google Scholar]
-
Munns R, Tester M. Mechanisms of Salinity Tolerance. Annu. Rev. Plant Biol. 2008, 59, 651–681. DOI:10.1146/annurev.arplant.59.032607.092911 [Google Scholar]
-
Bailey-Serres J, Lee SC, Brinton E. Waterproofing Crops: Effective Flooding Survival Strategies. Plant Physiol. 2012, 160, 1698–1709. DOI:10.1104/pp.112.208173 [Google Scholar]
-
Yun P, Shabala S. Optimizing plant nutrient acquisition under hypoxia: Likely trade-offs, implications for breeders, and lessons from wetland species. J. Exp. Bot. 2026, erag093. DOI:10.1093/jxb/erag093 [Google Scholar]
-
Zhao C, Zhang H, Song C, Zhu J-K, Shabala S. Mechanisms of Plant Responses and Adaptation to Soil Salinity. Innov. 2020, 1, 100017. DOI:10.1016/j.xinn.2020.100017 [Google Scholar]
-
Vance CP, Uhde-Stone C, Allan DL. Phosphorus acquisition and use: Critical adaptations by plants for securing a nonrenewable resource. New Phytol. 2003, 157, 423–447. DOI:10.1046/j.1469-8137.2003.00695.x [Google Scholar]
-
Lambers H. Phosphorus Acquisition and Utilization in Plants. Annu. Rev. Plant Biol. 2022, 73, 17–42. DOI:10.1146/annurev-arplant-102720-125738 [Google Scholar]
-
Lopez G, Ahmadi SH, Amelung W, Athmann M, Ewert F, Gaiser T, et al. Nutrient deficiency effects on root architecture and root-to-shoot ratio in arable crops. Front. Plant Sci. 2023, 13, 1067498. DOI:10.3389/fpls.2022.1067498 [Google Scholar]
-
Ceccarelli S, Grando S. Diversity as a Plant Breeding Objective. Agronomy 2024, 14, 550. DOI:10.3390/agronomy14030550 [Google Scholar]
-
Challinor AJ, Watson J, Lobell DB, Howden SM, Smith DR, Chhetri N. A meta-analysis of crop yield under climate change and adaptation. Nat. Clim. Change 2014, 4, 287–291. DOI:10.1038/nclimate2153 [Google Scholar]
-
Carey-Fung O, Johnson AAT. From convention to innovation: The role of genetic modification and genome editing in Australian wheat breeding. AoB PLANTS 2025, 17, plaf040. DOI:10.1093/aobpla/plaf040 [Google Scholar]
-
Araus JL, Kefauver SC, Zaman-Allah M, Olsen MS, Cairns JE. Translating High-Throughput Phenotyping into Genetic Gain. Trends Plant Sci. 2018, 23, 451–466. DOI:10.1016/j.tplants.2018.02.001 [Google Scholar]
-
Shabala S, Chen X, Yun P, Zhou M. Salinity tolerance in wheat: Rethinking the targets. J. Exp. Bot. 2025, eraf152. DOI:10.1093/jxb/eraf152 [Google Scholar]
-
Thomson MJ, de Ocampo M, Egdane J, Rahman MA, Sajise AG, Adorada DL, et al. Characterizing the Saltol Quantitative Trait Locus for Salinity Tolerance in Rice. Rice 2010, 3, 148–160. DOI:10.1007/s12284-010-9053-8 [Google Scholar]
-
Jørgensen IH. Discovery, characterization and exploitation of Mlo powdery mildew resistance in barley. Euphytica 1992, 63, 141–152. DOI:10.1007/BF00023919 [Google Scholar]
-
Voss-Fels KP, Qian L, Parra-Londono S, Uptmoor R, Frisch M, Keeble-Gagnère G, et al. Linkage drag constrains the roots of modern wheat. Plant Cell Environ. 2017, 40, 717–725. DOI:10.1111/pce.12888 [Google Scholar]
-
Peng T, Sun X, Mumm RH. Optimized breeding strategies for multiple trait integration: I. Minimizing linkage drag in single event introgression. Mol. Breed. 2014, 33, 89–104. DOI:10.1007/s11032-013-9936-7 [Google Scholar]
-
Cobb JN, Juma RU, Biswas PS, Arbelaez JD, Rutkoski J, Atlin G, et al. Enhancing the rate of genetic gain in public-sector plant breeding programs: Lessons from the breeder’s equation. Theor. Appl. Genet. 2019, 132, 627–645. DOI:10.1007/s00122-019-03317-0 [Google Scholar]
-
Gaynor RC, Gorjanc G, Bentley AR, Ober ES, Howell P, Jackson R, et al. A Two-Part Strategy for Using Genomic Selection to Develop Inbred Lines. Crop Sci. 2017, 57, 2372–2386. DOI:10.2135/cropsci2016.09.0742 [Google Scholar]
-
Watson A, Ghosh S, Williams MJ, Cuddy WS, Simmonds J, Rey M-D, et al. Speed breeding is a powerful tool to accelerate crop research and breeding. Nat. Plants 2018, 4, 23–29. DOI:10.1038/s41477-017-0083-8 [Google Scholar]
-
Ghosh S, Watson A, Gonzalez-Navarro OE, Ramirez-Gonzalez RH, Yanes L, Mendoza-Suárez M, et al. Speed breeding in growth chambers and glasshouses for crop breeding and model plant research. Nat. Protoc. 2018, 13, 2944–2963. DOI:10.1038/s41596-018-0072-z [Google Scholar]
-
Zheng Z, Gao S, Wang H, Liu C. Shortening generation times for winter cereals by vernalizing seedlings from young embryos at 10 degree Celsius. Plant Breed. 2023, 142, 202–210. DOI:10.1111/pbr.13074 [Google Scholar]
-
Sandhu N, Singh J, Pruthi G, Verma VK, Raigar OP, Bains NS, et al. SpeedyPaddy: A revolutionized cost-effective protocol for large scale offseason advancement of rice germplasm. Plant Methods 2024, 20, 109. DOI:10.1186/s13007-024-01235-x [Google Scholar]
-
Jähne F, Hahn V, Würschum T, Leiser WL. Speed breeding short-day crops by LED-controlled light schemes. Theor. Appl. Genet. 2020, 133, 2335–2342. DOI:10.1007/s00122-020-03601-4 [Google Scholar]
-
Manzur JP, Oliva-Alarcón M, Rodríguez-Burruezo A. In vitro germination of immature embryos for accelerating generation advancement in peppers (Capsicum annuum L.). Sci. Hortic. 2014, 170, 203–210. DOI:10.1016/j.scienta.2014.03.015 [Google Scholar]
-
de Paiva Neto VB, Otoni WC. Carbon sources and their osmotic potential in plant tissue culture: Does it matter? Sci. Hortic. 2003, 97, 193–202. DOI:10.1016/S0304-4238(02)00231-5 [Google Scholar]
-
Zheng Z, Wang HB, Chen GD, Yan GJ, Liu CJ. A procedure allowing up to eight generations of wheat and nine generations of barley per annum. Euphytica 2013, 191, 311–316. DOI:10.1007/s10681-013-0909-z [Google Scholar]
-
Rogo U, Fambrini M, Pugliesi C. Embryo Rescue in Plant Breeding. Plants 2023, 12, 3106. DOI:10.3390/plants12173106 [Google Scholar]
-
Hospital F. Selection in backcross programmes. Philos. Trans. R. Soc. B Biol. Sci. 2005, 360, 1503–1511. DOI:10.1098/rstb.2005.1670 [Google Scholar]
-
Collard BCY, Mackill DJ. Marker-assisted selection: An approach for precision plant breeding in the twenty-first century. Philos. Trans. R. Soc. B Biol. Sci. 2007, 363, 557–572. DOI:10.1098/rstb.2007.2170 [Google Scholar]
-
Frisch M, Bohn M, Melchinger AA. Minimum Sample Size and Optimal Positioning of Flanking Markers in Marker-Assisted Backcrossing for Transfer of a Target Gene. Crop Sci. 1999, 39, 967–975. DOI:10.2135/cropsci1999.0011183X003900040003x [Google Scholar]
-
Raffo MA, Jensen J. Gene × gene and genotype × environment interactions in wheat. Crop Sci. 2023, 63, 1779–1793. DOI:10.1002/csc2.20986 [Google Scholar]
-
Xu Y, Crouch JH. Marker-Assisted Selection in Plant Breeding: From Publications to Practice. Crop Sci. 2008, 48, 391–407. DOI:10.2135/cropsci2007.04.0191 [Google Scholar]
-
Xu K, Xu X, Fukao T, Canlas P, Maghirang-Rodriguez R, Heuer S, et al. Sub1A is an ethylene-response-factor-like gene that confers submergence tolerance to rice. Nature 2006, 442, 705–708. DOI:10.1038/nature04920 [Google Scholar]
-
Frisch M, Bohn M, Melchinger AE. Comparison of Selection Strategies for Marker-Assisted Backcrossing of a Gene. Crop Sci. 1999, 39, 1295–1301. DOI:10.2135/cropsci1999.3951295x [Google Scholar]
-
Frisch M, Bohn M, Melchinger AE. Computer note. PLABSIM: Software for simulation of marker-assisted backcrossing. J. Hered. 2000, 91, 86–87. DOI:10.1093/jhered/91.1.86 [Google Scholar]
-
Ribaut JM, Jiang C, Hoisington D. Simulation Experiments on Efficiencies of Gene Introgression by Backcrossing. Crop Sci. 2002, 42, 557–565. DOI:10.2135/cropsci2002.5570 [Google Scholar]
-
Kalpana MP, Ramesh S, Siddu CB, Basanagouda G, Madhusudan K, Sathish H, et al. Low density marker-based effectiveness and efficiency of early-generation genomic selection relative to phenotype-based selection in dolichos bean (Lablab purpureus L. Sweet). Plant Genome 2025, 18, e70039. DOI:10.1002/tpg2.70039 [Google Scholar]
-
Chukwu SC, Rafii MY, Ramlee SI, Ismail SI, Oladosu Y, Muhammad I, et al. Recovery of Recurrent Parent Genome in a Marker-Assisted Backcrossing Against Rice Blast and Blight Infections Using Functional Markers and SSRs. Plants 2020, 9, 1411. DOI:10.3390/plants9111411 [Google Scholar]
-
Ganie SA, Azevedo RA. Why stress-resistant crops remain a scientific promise rather than a farming reality? Bridging the gap between genetic discovery and agricultural impact. Ann. Appl. Biol. 2026, 188, 6–10. DOI:10.1111/aab.70050 [Google Scholar]
-
Neeraja CN, Maghirang-Rodriguez R, Pamplona A, Heuer S, Collard BCY, Septiningsih EM, et al. A marker-assisted backcross approach for developing submergence-tolerant rice cultivars. Theor. Appl. Genet. 2007, 115, 767–776. DOI:10.1007/s00122-007-0607-0 [Google Scholar]
-
Septiningsih EM, Pamplona AM, Sanchez DL, Neeraja CN, Vergara GV, Heuer S, et al. Development of submergence-tolerant rice cultivars: The Sub1 locus and beyond. Ann. Bot. 2009, 103, 151–160. DOI:10.1093/aob/mcn206 [Google Scholar]
-
Heuer S, Lu X, Chin JH, Tanaka JP, Kanamori H, Matsumoto T, et al. Comparative sequence analyses of the major quantitative trait locus phosphorus uptake 1 (Pup1) reveal a complex genetic structure. Plant Biotechnol. J. 2009, 7, 456–471. DOI:10.1111/j.1467-7652.2009.00415.x [Google Scholar]
-
Chin JH, Gamuyao R, Dalid C, Bustamam M, Prasetiyono J, Moeljopawiro S, et al. Developing Rice with High Yield under Phosphorus Deficiency: Pup1 Sequence to Application. Plant Physiol. 2011, 156, 1202–1216. DOI:10.1104/pp.111.175471 [Google Scholar]
-
Uga Y, Sugimoto K, Ogawa S, Rane J, Ishitani M, Hara N, et al. Control of root system architecture by DEEPER ROOTING 1 increases rice yield under drought conditions. Nat. Genet. 2013, 45, 1097–1102. DOI:10.1038/ng.2725 [Google Scholar]
-
Byrt CS, Platten JD, Spielmeyer W, James RA, Lagudah ES, Dennis ES, et al. HKT1;5-Like Cation Transporters Linked to Na+ Exclusion Loci in Wheat, Nax2 and Kna1. Plant Physiol. 2007, 143, 1918–1928. DOI:10.1104/pp.106.093476 [Google Scholar]
-
James RA, Blake C, Byrt CS, Munns R. Major genes for Na+ exclusion, Nax1 and Nax2 (wheat HKT1;4 and HKT1;5), decrease Na+ accumulation in bread wheat leaves under saline and waterlogged conditions. J. Exp. Bot. 2011, 62, 2939–2947. DOI:10.1093/jxb/err003 [Google Scholar]
-
Munns R, Day DA, Fricke W, Watt M, Arsova B, Barkla BJ, et al. Energy costs of salt tolerance in crop plants. New Phytol. 2020, 225, 1072–1090. DOI:10.1111/nph.15864 [Google Scholar]
-
Munns R, Passioura JB, Colmer TD, Byrt CS. Osmotic adjustment and energy limitations to plant growth in saline soil. New Phytol. 2020, 225, 1091–1096. DOI:10.1111/nph.15862 [Google Scholar]
-
Zhang X, Zhou G, Shabala S, Koutoulis A, Shabala L, Johnson P, et al. Identification of aerenchyma formation-related QTL in barley that can be effective in breeding for waterlogging tolerance. Theor. Appl. Genet. 2016, 129, 1167–1177. DOI:10.1007/s00122-016-2693-3 [Google Scholar]
-
Ma Y, Li C, Ryan PR, Shabala S, You J, Liu J, et al. A new allele for aluminium tolerance gene in barley (Hordeum vulgare L.). BMC Genom. 2016, 17, 186. DOI:10.1186/s12864-016-2551-3 [Google Scholar]
-
Zhou G, Broughton S, Zhang X-Q, Ma Y, Zhou M, Li C. Genome-Wide Association Mapping of Acid Soil Resistance in Barley (Hordeum vulgare L.). Front. Plant Sci. 2016, 7, 406. DOI:10.3389/fpls.2016.00406 [Google Scholar]
-
Zhou G, Delhaize E, Zhou M, Ryan PR. The barley MATE gene, HvAACT1, increases citrate efflux and Al3+ tolerance when expressed in wheat and barley. Ann. Bot. 2013, 112, 603–612. DOI:10.1093/aob/mct135 [Google Scholar]
-
Guo C, Shabala S, Liu K, Zhou M, Zhao C. The impact of HvAACT1 gene conferring aluminium toxicity tolerance on barley yield in acid soils. Eur. J. Agron. 2026, 173, 127907. DOI:10.1016/j.eja.2025.127907 [Google Scholar]
-
Hammer G, Messina C, van Oosterom E, Chapman S, Singh V, Borrell A, et al. Molecular Breeding for Complex Adaptive Traits: How Integrating Crop Ecophysiology and Modelling Can Enhance Efficiency. In Crop Systems Biology: Narrowing the Gaps Between Crop Modelling and Genetics; Yin X, Struik PC, Eds.; Springer International Publishing: Cham, Switzerland, 2016; pp. 147–162. [Google Scholar]
-
Krasensky J, Jonak C. Drought, salt, and temperature stress-induced metabolic rearrangements and regulatory networks. J. Exp. Bot. 2012, 63, 1593–1608. DOI:10.1093/jxb/err460 [Google Scholar]
-
Gupta S, Kaur R, Upadhyay A, Chauhan A, Tripathi V. Unveiling the secrets of abiotic stress tolerance in plants through molecular and hormonal insights. 3 Biotech 2024, 14, 252. DOI:10.1007/s13205-024-04083-7 [Google Scholar]
-
Ghosh UK, Islam MN, Siddiqui MN, Khan MAR. Understanding the roles of osmolytes for acclimatizing plants to changing environment: A review of potential mechanism. Plant Signal. Behav. 2021, 16, 1913306. DOI:10.1080/15592324.2021.1913306 [Google Scholar]
-
Shabala S, Bose J, Hedrich R. Salt bladders: Do they matter? Trends Plant Sci. 2014, 19, 687–691. DOI:10.1016/j.tplants.2014.09.001 [Google Scholar]
-
Hasan MM, Gong L, Nie Z-F, Li F-P, Ahammed GJ, Fang X-W. ABA-induced stomatal movements in vascular plants during dehydration and rehydration. Environ. Exp. Bot. 2021, 186, 104436. DOI:10.1016/j.envexpbot.2021.104436 [Google Scholar]
-
Chen G, Qin Y, Wang J, Li S, Zeng F, Deng F, et al. Stomatal evolution and plant adaptation to future climate. Plant Cell Environ. 2024, 47, 3299–3315. DOI:10.1111/pce.14953 [Google Scholar]
-
Teng Z, Lyu J, Chen Y, Zhang J, Ye N. Effects of stress-induced ABA on root architecture development: Positive and negative actions. Crop J. 2023, 11, 1072–1079. DOI:10.1016/j.cj.2023.06.007 [Google Scholar]
-
Cutler SR, Rodriguez PL, Finkelstein RR, Abrams SR. Abscisic Acid: Emergence of a Core Signaling Network. Annu. Rev. Plant Biol. 2010, 61, 651–679. DOI:10.1146/annurev-arplant-042809-112122 [Google Scholar]
-
Van Der Sman AJM, Voesenek LACJ, Blom CWPM, Harren FJM, Reuss J. The Role of Ethylene in Shoot Elongation with Respect to Survival and Seed Output of Flooded Rumex maritimus L. Plants. Funct. Ecol. 1991, 5, 304–313. DOI:10.2307/2389269 [Google Scholar]
-
Sharmita O, Siddique AB, Liu K, Shabala S, Harrison MT, Zhou M, et al. Boosting crop resilience to waterlogging through hormone-regulated root traits. Eur. J. Agron. 2026, 174, 127948. DOI:10.1016/j.eja.2025.127948 [Google Scholar]
-
Visser EJW, Bögemann GM. Aerenchyma formation in the wetland plant Juncus effusus is independent of ethylene. New Phytol. 2006, 171, 305–314. DOI:10.1111/j.1469-8137.2006.01764.x [Google Scholar]
-
Hartman S, Sasidharan R, Voesenek LACJ. The role of ethylene in metabolic acclimations to low oxygen. New Phytol. 2021, 229, 64–70. DOI:10.1111/nph.16378 [Google Scholar]
-
Singhal RK, Fahad S, Kumar P, Choyal P, Javed T, Jinger D, et al. Beneficial elements: New Players in improving nutrient use efficiency and abiotic stress tolerance. Plant Growth Regul. 2023, 100, 237–265. DOI:10.1007/s10725-022-00843-8 [Google Scholar]
-
Demidchik V, Shabala S, Isayenkov S, Cuin TA, Pottosin I. Calcium transport across plant membranes: Mechanisms and functions. New Phytol. 2018, 220, 49–69. DOI:10.1111/nph.15266 [Google Scholar]
-
Chimungu JG, Brown KM, Lynch JP. Large Root Cortical Cell Size Improves Drought Tolerance in Maize. Plant Physiol. 2014, 166, 2166–2178. DOI:10.1104/pp.114.250449 [Google Scholar]
-
Lynch JP, Brown KM. Topsoil foraging—An architectural adaptation of plants to low phosphorus availability. Plant Soil 2001, 237, 225–237. DOI:10.1023/A:1013324727040 [Google Scholar]
-
Sabadin PK, Malosetti M, Boer MP, Tardin FD, Santos FG, Guimarães CT, et al. Studying the genetic basis of drought tolerance in sorghum by managed stress trials and adjustments for phenological and plant height differences. Theor. Appl. Genet. 2012, 124, 1389–1402. DOI:10.1007/s00122-012-1795-9 [Google Scholar]
-
Yamauchi T, Colmer TD, Pedersen O, Nakazono M. Regulation of Root Traits for Internal Aeration and Tolerance to Soil Waterlogging-Flooding Stress. Plant Physiol. 2018, 176, 1118–1130. DOI:10.1104/pp.17.01157 [Google Scholar]
-
Mackay TFC, Stone EA, Ayroles JF. The genetics of quantitative traits: Challenges and prospects. Nat. Rev. Genet. 2009, 10, 565–577. DOI:10.1038/nrg2612 [Google Scholar]
-
Bhat JA, Yu D, Bohra A, Ganie SA, Varshney RK. Features and applications of haplotypes in crop breeding. Commun. Biol. 2021, 4, 1266. DOI:10.1038/s42003-021-02782-y [Google Scholar]
-
Tardivel A, Torkamaneh D, Lemay M-A, Belzile F, O’Donoughue LS. A Systematic Gene-Centric Approach to Define Haplotypes and Identify Alleles on the Basis of Dense Single Nucleotide Polymorphism Datasets. Plant Genome 2019, 12, 180061. DOI:10.3835/plantgenome2018.08.0061 [Google Scholar]
-
Tibbs Cortes L, Zhang Z, Yu J. Status and prospects of genome-wide association studies in plants. Plant Genome 2021, 14, e20077. DOI:10.1002/tpg2.20077 [Google Scholar]
-
Hyten DL, Song Q, Zhu Y, Choi I-Y, Nelson RL, Costa JM, et al. Impacts of genetic bottlenecks on soybean genome diversity. Proc. Natl. Acad. Sci. USA 2006, 103, 16666–16671. DOI:10.1073/pnas.0604379103 [Google Scholar]
-
Mayer M, Hölker AC, González-Segovia E, Bauer E, Presterl T, Ouzunova M, et al. Discovery of beneficial haplotypes for complex traits in maize landraces. Nat. Commun. 2020, 11, 4954. DOI:10.1038/s41467-020-18683-3 [Google Scholar]
-
Liu J, Rasheed A, He Z, Imtiaz M, Arif A, Mahmood T, et al. Genome-wide variation patterns between landraces and cultivars uncover divergent selection during modern wheat breeding. Theor. Appl. Genet. 2019, 132, 2509–2523. DOI:10.1007/s00122-019-03367-4 [Google Scholar]
-
Abed A, Belzile F. Comparing Single-SNP, Multi-SNP, and Haplotype-Based Approaches in Association Studies for Major Traits in Barley. Plant Genome 2019, 12, 190036. DOI:10.3835/plantgenome2019.05.0036 [Google Scholar]
-
Hayes BJ, Chamberlain AJ, McPartlan H, Macleod I, Sethuraman L, Goddard ME. Accuracy of marker-assisted selection with single markers and marker haplotypes in cattle. Genet. Res. 2007, 89, 215–220. DOI:10.1017/S0016672307008865 [Google Scholar]
-
Alemu A, Batista L, Singh PK, Ceplitis A, Chawade A. Haplotype-tagged SNPs improve genomic prediction accuracy for Fusarium head blight resistance and yield-related traits in wheat. Theor. Appl. Genet. 2023, 136, 92. DOI:10.1007/s00122-023-04352-8 [Google Scholar]
-
Núñez-Ramírez R, Sánchez-Barrena MJ, Villalta I, Vega JF, Pardo JM, Quintero FJ, et al. Structural Insights on the Plant Salt-Overly-Sensitive 1 (SOS1) Na+/H+ Antiporter. J. Mol. Biol. 2012, 424, 283–294. DOI:10.1016/j.jmb.2012.09.015 [Google Scholar]
-
Shabala S, Chen G, Chen Z-H, Pottosin I. The energy cost of the tonoplast futile sodium leak. New Phytol. 2020, 225, 1105–1110. DOI:10.1111/nph.15758 [Google Scholar]
-
Pan T, Liu M, Kreslavski VD, Zharmukhamedov SK, Nie C, Yu M, et al. Non-stomatal limitation of photosynthesis by soil salinity. Crit. Rev. Environ. Sci. Technol. 2021, 51, 791–825. DOI:10.1080/10643389.2020.1735231 [Google Scholar]
-
Schilling RK, Tester M, Marschner P, Plett DC, Roy SJ. AVP1: One Protein, Many Roles. Trends Plant Sci. 2017, 22, 154–162. DOI:10.1016/j.tplants.2016.11.012 [Google Scholar]
-
Gaxiola R, Hirschi KD. Regulated to respond: Dual localization and dynamic control of AVP1 in plant carbon partitioning. Front. Plant Sci. 2025, 16, 1605111. DOI:10.3389/fpls.2025.1605111 [Google Scholar]
-
Wang M, Cheng J, Wu J, Chen J, Liu D, Wang C, et al. Variation in TaSPL6-D confers salinity tolerance in bread wheat by activating TaHKT1;5-D while preserving yield-related traits. Nat. Genet. 2024, 56, 1257–1269. DOI:10.1038/s41588-024-01762-2 [Google Scholar]
-
Venkataraman G, Shabala S, Véry A-A, Hariharan GN, Somasundaram S, Pulipati S, et al. To exclude or to accumulate? Revealing the role of the sodium HKT1;5 transporter in plant adaptive responses to varying soil salinity. Plant Physiol. Biochem. 2021, 169, 333–342. DOI:10.1016/j.plaphy.2021.11.030 [Google Scholar]
-
Yang Y, Ti J, Zou J, Wu Y, Rees RM, Harrison MT, et al. Optimizing crop rotation increases soil carbon and reduces GHG emissions without sacrificing yields. Agric. Ecosyst. Environ. 2023, 342, 108220. DOI:10.1016/j.agee.2022.108220 [Google Scholar]
-
Muehlbauer GJ, Specht JE, Thomas-Compton MA, Staswick PE, Bernard RL. Near-Isogenic Lines—A Potential Resource in the Integration of Conventional and Molecular Marker Linkage Maps. Crop Sci. 1988, 28, 729–735. DOI:10.2135/cropsci1988.0011183X002800050002x [Google Scholar]
-
Wu Y, Yu L, Pan C, Dai Z, Li Y, Xiao N, et al. Development of near-isogenic lines with different alleles of Piz locus and analysis of their breeding effect under Yangdao 6 background. Mol. Breed. 2016, 36, 12. DOI:10.1007/s11032-016-0433-7 [Google Scholar]
-
Sujariya S, Jongdee B, Fukai S. Estimation of flowering time and its effect on grain yield of photoperiod sensitive varieties in rainfed lowland rice in Northeast Thailand. Field Crops Res. 2023, 302, 109075. DOI:10.1016/j.fcr.2023.109075 [Google Scholar]
-
Century K, Reuber TL, Ratcliffe OJ. Regulating the Regulators: The Future Prospects for Transcription-Factor-Based Agricultural Biotechnology Products. Plant Physiol. 2008, 147, 20–29. DOI:10.1104/pp.108.117887 [Google Scholar]
-
Kummari D, Palakolanu SR, Kishor PBK, Bhatnagar-Mathur P, Singam P, Vadez V, et al. An update and perspectives on the use of promoters in plant genetic engineering. J. Biosci. 2020, 45, 119. DOI:10.1007/s12038-020-00087-6 [Google Scholar]
-
Singh S, Mackill DJ, Ismail AM. Physiological basis of tolerance to complete submergence in rice involves genetic factors in addition to the SUB1 gene. AoB PLANTS 2014, 6, plu060. DOI:10.1093/aobpla/plu060 [Google Scholar]
-
Dar MH, Zaidi NW, Waza SA, Verulkar SB, Ahmed T, Singh PK, et al. No yield penalty under favorable conditions paving the way for successful adoption of flood tolerant rice. Sci. Rep. 2018, 8, 9245. DOI:10.1038/s41598-018-27648-y [Google Scholar]
-
Henry A, Swamy BPM, Dixit S, Torres RD, Batoto TC, Manalili M, et al. Physiological mechanisms contributing to the QTL-combination effects on improved performance of IR64 rice NILs under drought. J. Exp. Bot. 2015, 66, 1787–1799. DOI:10.1093/jxb/eru506 [Google Scholar]
-
Dwivedi P, Ramawat N, Dhawan G, Gopala Krishnan S, Vinod KK, Singh MP, et al. Drought Tolerant near Isogenic Lines (NILs) of Pusa 44 Developed through Marker Assisted Introgression of qDTY2.1 and qDTY3.1 Enhances Yield under Reproductive Stage Drought Stress. Agriculture 2021, 11, 64. DOI:10.3390/agriculture11010064 [Google Scholar]
-
Munns R, James RA, Xu B, Athman A, Conn SJ, Jordans C, et al. Wheat grain yield on saline soils is improved by an ancestral Na+ transporter gene. Nat. Biotechnol. 2012, 30, 360–364. DOI:10.1038/nbt.2120 [Google Scholar]
-
Schnurbusch T, Hayes J, Hrmova M, Baumann U, Ramesh SA, Tyerman SD, et al. Boron Toxicity Tolerance in Barley through Reduced Expression of the Multifunctional Aquaporin HvNIP2;1. Plant Physiol. 2010, 153, 1706–1715. DOI:10.1104/pp.110.158832 [Google Scholar]
-
Yadav S, Sandhu N, Majumder RR, Dixit S, Kumar S, Singh SP, et al. Epistatic interactions of major effect drought QTLs with genetic background loci determine grain yield of rice under drought stress. Sci. Rep. 2019, 9, 2616. DOI:10.1038/s41598-019-39084-7 [Google Scholar]
-
Delhaize E, James RA, Ryan PR. Aluminium tolerance of root hairs underlies genotypic differences in rhizosheath size of wheat (Triticum aestivum) grown on acid soil. New Phytol. 2012, 195, 609–619. DOI:10.1111/j.1469-8137.2012.04183.x [Google Scholar]
-
Khabaz-Saberi H, Setter TL, Waters I. Waterlogging Induces High to Toxic Concentrations of Iron, Aluminum, and Manganese in Wheat Varieties on Acidic Soil. J. Plant Nutr. 2006, 29, 899–911. DOI:10.1080/01904160600649161 [Google Scholar]
-
Chen J, Zhao C, Harrison MT, Zhou M. Nitrogen Fertilization Alleviates Barley (Hordeum vulgare L.) Waterlogging. Agronomy 2024, 14, 1712. DOI:10.3390/agronomy14081712 [Google Scholar]
-
Bernardo R. Reinventing quantitative genetics for plant breeding: Something old, something new, something borrowed, something BLUE. Heredity 2020, 125, 375–385. DOI:10.1038/s41437-020-0312-1 [Google Scholar]
-
Guerrero RF, Muir CD, Josway S, Moyle LC. Pervasive antagonistic interactions among hybrid incompatibility loci. PLOS Genet. 2017, 13, e1006817. DOI:10.1371/journal.pgen.1006817 [Google Scholar]
-
Heffner EL, Lorenz AJ, Jannink J-L, Sorrells ME. Plant Breeding with Genomic Selection: Gain per Unit Time and Cost. Crop Sci. 2010, 50, 1681–1690. DOI:10.2135/cropsci2009.11.0662 [Google Scholar]
-
Hickey LT, Hafeez AN, Robinson H, Jackson SA, Leal-Bertioli SCM, Tester M, et al. Breeding crops to feed 10 billion. Nat. Biotechnol. 2019, 37, 744–754. DOI:10.1038/s41587-019-0152-9 [Google Scholar]
-
Crossa J, Pérez-Rodríguez P, Cuevas J, Montesinos-López O, Jarquín D, de los Campos G, et al. Genomic Selection in Plant Breeding: Methods, Models, and Perspectives. Trends Plant Sci. 2017, 22, 961–975. DOI:10.1016/j.tplants.2017.08.011 [Google Scholar]
-
Cheng C-Y, Li Y, Varala K, Bubert J, Huang J, Kim GJ, et al. Evolutionarily informed machine learning enhances the power of predictive gene-to-phenotype relationships. Nat. Commun. 2021, 12, 5627. DOI:10.1038/s41467-021-25893-w [Google Scholar]
-
Varshney RK, Gaur PM, Chamarthi SK, Krishnamurthy L, Tripathi S, Kashiwagi J, et al. Fast-Track Introgression of “QTL-hotspot” for Root Traits and Other Drought Tolerance Traits in JG 11, an Elite and Leading Variety of Chickpea. Plant Genome 2013, 6, plantgenome2013-07. DOI:10.3835/plantgenome2013.07.0022 [Google Scholar]
