Advances in the Application of CRISPR/Cas Systems in Molecular Diagnostics
Haozhe Ren 1,2,3 Shiyuan Luo 4 Yiqing Yu 5 Pengfei Chen 6,7,* Ke Wang 1,*,†
Received: 30 October 2025 Revised: 06 January 2026 Accepted: 26 February 2026 Published: 04 March 2026
© 2026 The authors. This is an open access article under the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/).
1. Introduction
1.1. Mechanisms and Taxonomy of CRISPR/Cas Systems
CRISPR/Cas systems mediate an adaptive immune response in bacteria and archaea by the programmable, sequence-specific recognition and endonucleolytic cleavage of foreign nucleic acids. Phylogenomic analyses led Makarova et al. [1] to propose a widely adopted classification that divides these systems into Class I (multi-subunit effector complexes) and Class II (single-protein effectors). Owing to distinct effector-architectures, each class is further subdivided into six major types (I–VI), whose key molecular features are summarized in Table 1.
Table 1. The relation of CRISPR/Cas system effector proteins.
|
Cas Protein |
Class |
Type |
Target Nucleic Acid |
Trans-Cleavage Activity/Substrate |
PAM/PFS Sequence |
References |
|---|---|---|---|---|---|---|
|
Cas9 |
Class II systems |
Type II |
dsDNA |
No |
3′ GC-rich PAM |
|
|
Cas7–11 (Multi-subunit effector) |
Class I systems |
Type III |
ssRNA |
No |
--- |
|
|
Cas12a (Cpf1) |
Class II systems |
Type V |
dsDNA/ssDNA |
Yes/ssDNA |
5′ AT-rich PAM |
[7] |
|
Cas12b |
Class II systems |
Type V |
dsDNA/ssDNA |
Yes/ssDNA |
5′ AT-rich PAM |
[8] |
|
Cas12f (Cas14) |
Class II systems |
Type V |
dsDNA/ssDNA |
Yes/ssDNA |
5′ T-rich PAM |
|
|
CasΦ (Cas12j2) |
Class II systems |
Type V |
dsDNA/ssDNA |
Yes/ssDNA |
5′ TBN |
[14] |
|
Cas13 |
Class II systems |
Type VI |
ssRNA |
Yes/ssRNA |
3′ non-G-PFS |
[15] |
|
Cas3 protein (Multi-subunit effector) |
Class I systems |
Type I |
dsDNA |
Yes/ssDNA |
AAG |
[16] |
Note: Cas9 exhibits trans-cleavage activity, but this activity strictly requires a tracrRNA–crRNA duplex and cannot be elicited by a single-guide RNA (sgRNA) alone [17].
Class I effectors are multi-subunit ribonucleoprotein complexes represented by types I, III, and IV. The signature Cascade complex assembles Cas5, Cas6, Cas7, Cas8, and Cas11 to process pre-crRNA, recognize target sequences, and catalyze nucleic-acid cleavage. Pronounced differences in subunit composition between subtypes substantially enhance system adaptability [1]. However, the inherent macromolecular complexity, large molecular mass, and intricate regulation have largely excluded these systems from the diagnostic arena. Cas7–11 is a streamlined chimeric effector comprising four Cas7 domains fused to a single Cas11 domain. It overcomes these constraints by executing crRNA-guided RNA cleavage as an autonomous entity [6]. This self-contained architecture obviates auxiliary factors and collapses multistep workflows into a single reaction. Rational reprogramming of its RNA-recognition surface or seamless integration with microfluidic multiplexers now positions Cas7–11 as a complementary platform to Class 2 effectors, enabling simultaneous pathogen screening and tumor-marker quantification, thereby broadening the CRISPR diagnostic ecosystem.
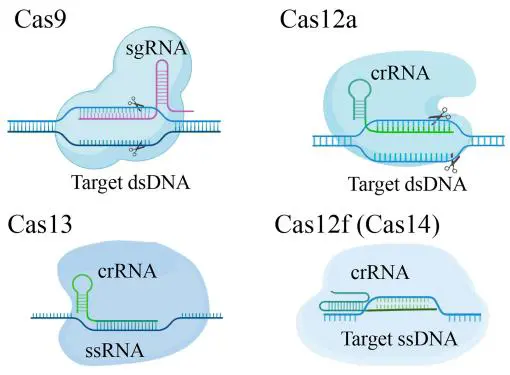
Class II immunity is mediated by single, monolithic effector proteins that have transformed genome engineering. Encompassing types II, V, and VI, this class deploys four canonical enzymes, Cas9, Cas12, Cas13, and Cas14, whose distinct substrate preferences span dsDNA, ssDNA, and RNA, thereby enabling versatile, multiplexed applications in gene editing and molecular diagnostics (Figure 1). The archetypal type II module centers on Cas9, which interrogates dsDNA via a crRNA–tracrRNA duplex (or single-guide RNA surrogate) and obligately engages a protospacer-adjacent motif (PAM). Conformational activation positions the RuvC and HNH nuclease domains for synchronized incision, yielding a staggered double-strand break with single-base precision [18,19]. Within the type II machinery, the complementary DNA strands are simultaneously docked, with the target strand engaging the HNH pocket and the non-target strand occupying the RuvC groove. Product entrapment within these catalytic clefts renders Cas9 catalytically quiescent, precluding subsequent turnovers. The type V repertoire employs the monomeric Cas12 architecture. In the paradigm V-A effector Cas12a (formerly Cpf1), a solitary RuvC domain is directed by a single crRNA to protospacer-adjacent motif (PAM)-flanked dsDNA. Upon complete complementarity, Cas12a initiates a sequential cis-cleavage cascade process in which the non-target strand is cleaved first, followed by incision of the target strand, thereby producing a precise double-strand break while maintaining enzymatic efficiency. After cis-cleavage of target dsDNA, Cas12a undergoes a conformational switch that unleashes indiscriminate, trans-acting DNase activity against non-target ssDNA. This trans-cleavage activity is converted into a fluorescence burst via rapid turnover of quenched ssDNA reporters, affording an ultrasensitive signal amplification module for diagnostic assays. Notably, crRNA-complementary single-stranded DNA can also activate Cas12a in a PAM-independent manner, eliciting efficient endonucleolytic cleavage. Cas12b exhibits exceptional thermostability within the type V-B clade, extending its utility to field-deployable point-of-care testing [8]. Cas12f (type V-F, previously Cas14) is the most compact CRISPR nuclease yet identified, spanning only 400–700 amino acids. Guided by a single crRNA, it binds cognate ssDNA and catalyzes indiscriminate trans-cleavage of bystander ssDNA, providing a minimal, high-fidelity platform for attomolar-level nucleic-acid detection without pre-amplification [9]. Compared with Cas9, Cas12 exhibits slower kinetics but substantially higher fidelity, reducing off-target activity to near-background levels. In contrast, type VI systems operate as RNA-specific ribonucleases. Upon crRNA-guided binding of a cognate transcript, Cas13a (type VI-A) undergoes allosteric activation that aligns its dual HEPN nuclease domains and triggers trans-RNase activity against proximal ssRNA. By encoding orthogonal crRNAs for each target and pairing them with spectrally distinct fluorescent reporters, single-reaction multiplexed detection of diverse analytes is achieved [15,20,21,22]. The interchangeable modularity and multiplexing capability inherent to Class 2 systems provide the foundation for advanced CRISPR-based diagnostics that achieve attomolar sensitivity while remaining directly deployable in both clinical laboratories and point-of-care settings.
1.2. Innovation and Optimization of CRISPR-Based Diagnostic Platforms
The emergence of CRISPR in nucleic-acid diagnostics reflects the combination of molecular precision and structural versatility. Through canonical Watson–Crick base pairing, a single rationally designed crRNA provides single-nucleotide specificity, enabling unambiguous discrimination of pathogen sequences or host mutations. Cas12 and Cas13 exploit orthogonal collateral pathways. dsDNA-bound Cas12 elicits trans-DNase activity that indiscriminately degrades ssDNA reporters, while RNA-bound Cas13 activates trans-RNase activity that non-specifically cleaves proximal ssRNA. Both cascades convert the primary recognition event into an amplified signal via rapid reporter turnover. Leveraging this intrinsic enzymatic amplification, the crRNA scaffold can be iteratively reprogrammed to detect DNA, RNA, or non-nucleic-acid analytes while remaining compatible with fluorescence, electrochemical, or colorimetric readouts. These features establish a modular engineering framework for portable, high-throughput diagnostic devices.
1.2.1. CRISPR-Based Diagnostic Technologies Employing Nucleic Acid Pre-Amplification
CRISPR/Cas systems achieve single-base specificity. However, their intrinsic detection limit is typically in the pM–fM range and remains insufficient for clinical trace analytes such as attomolar viral nucleic acids. To address this limitation, researchers have developed pre-amplification/CRISPR synergistic strategies that combine nucleic acid amplification with CRISPR trans-cleavage activity, achieving a 3–6 orders of magnitude improvement in analytical sensitivity. DETECTR integrates recombinase polymerase amplification (RPA) with Cas12a trans-cleavage, and HPV16/18 DNA recognized by the crRNA triggers reporter cleavage to give a detection limit of 10 copies µL−1 as shown in Figure 2A [7]. HOLMES v2 couples loop-mediated isothermal amplification (LAMP) with a thermostable Cas12b module for rapid single-tube DNA and RNA co-detection (Figure 2B) [23]. SHERLOCK combines RPA and T7 transcription with the RNA-guided trans-RNase activity of Cas13a, achieving attomolar detection of viral genomes under optimized pre-amplification conditions (Figure 2C) [15]. A NASBA–CRISPR hybrid uses toehold-mediated switches to convert Cas9 cleavage into a colorimetric readout for bedside Zika virus detection (Figure 2D) [24]. Cas14, activated by phosphorothioate-primed T7 exonuclease processing, selectively interrogates ssDNA targets and initiates downstream signal amplification with minimal components (Figure 2E) [9].
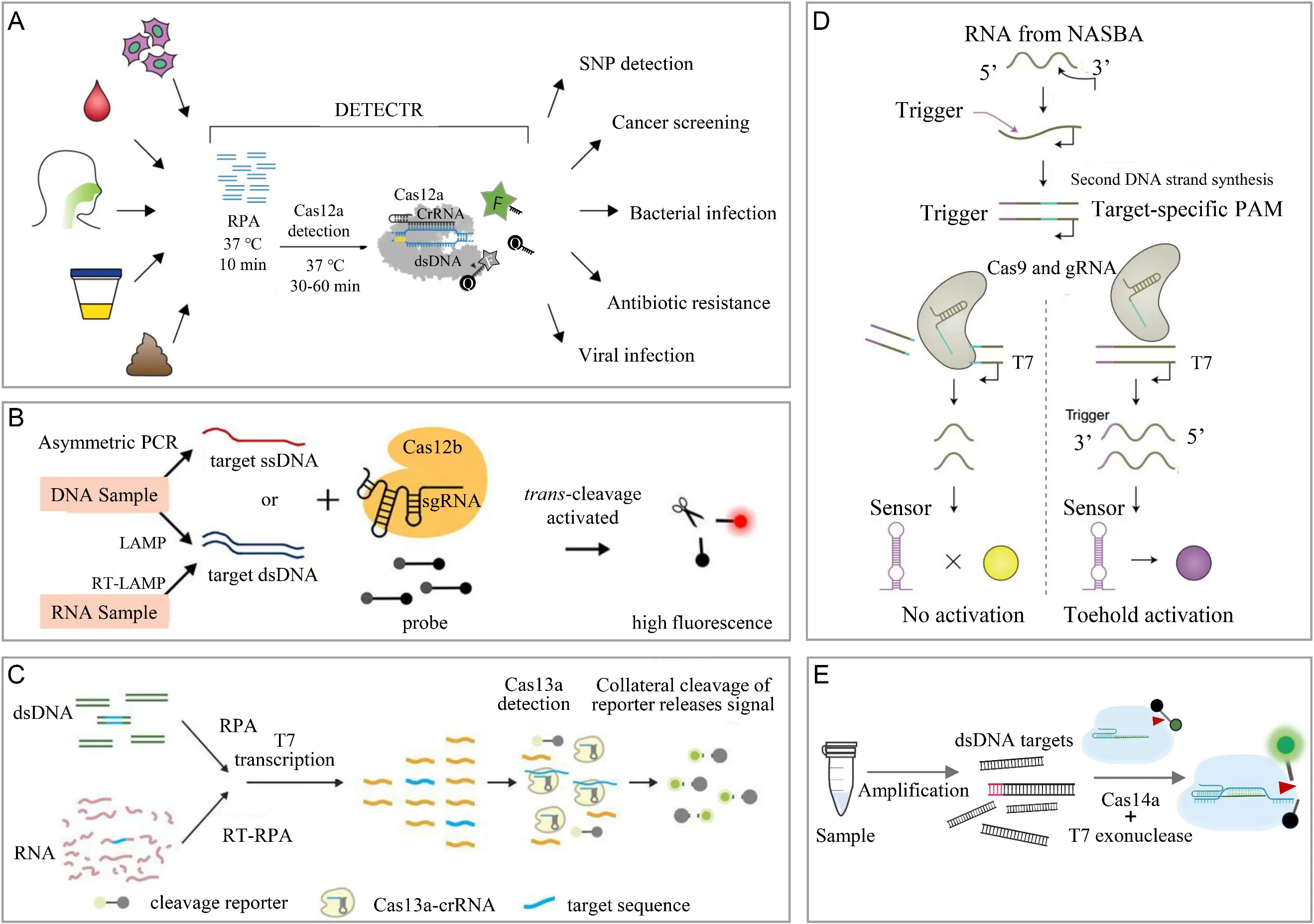
Figure 2. Schematic of CRISPR/Cas molecular diagnostic technology based on nucleic acid pre-amplification. (A) Schematic of the CRISPR/Cas12a-based DETECTR methodology for detecting clinical samples in various diagnostic application. (B) Schematic diagram of HOLMESv2 detection method. (C) Schematic of the SHERLOCK system combined with CRISPR/Cas13. (D) Schematic of the NASBACC detection method combining CRISPR/Cas9. (E) Schematic of CRISPR/Cas14a for detecting ssDNA.
1.2.2. CRISPR/Cas-Based Molecular Diagnostic Technologies without Nucleic Acid Amplification
Conventional nucleic acid amplification enhances analytical sensitivity but confines diagnostics to sophisticated thermal cyclers, multi-step workflows, and a carryover contamination risk, severely limiting point-of-care and resource-limited applications. Amplification-free CRISPR assays circumvent these constraints by employing trans-cleavage as an intrinsic signal amplification mechanism [25]. Upon perfect crRNA–target pairing, a single nucleic acid molecule activates the collateral nuclease activity of Cas12 or Cas13, which processively cleaves fluorogenic or electrochemical reporter substrates [26]. Coupling Cas trans-cleavage activity with downstream enzymatic recycling yields exponential signal amplification and routinely achieves sub-femtomolar sensitivity [27]. An archetypal strategy integrates Cas12a with exonuclease III-mediated target recycling, in which target binding initiates iterative cleavage and reduces the detection limit to below 1 fM. Fidelity is further enhanced by rational crRNA engineering, including chemical modifications, locked nucleic acids (LNAs), and optimized quencher spacing [28,29,30,31,32]. Recent transduction schemes further lower the limit; graphene field-effect transistors (FETs) convert Cas13a-directed RNA recognition into conductance changes, enabling label-free RNA quantification at 0.1 pM [33]. Droplet digital microfluidics has eliminated the requirement for pre-amplification altogether. Zhou and colleagues encapsulated Cas13a in picoliter droplets to achieve absolute quantification of single bacterial 16S rRNA copies [34]. Subsequent work leveraged Cas12a trans-cleavage for absolute counting of circulating tumor DNA at 1 copy µL−1 in a one-step workflow [35]. Watanabe’s SATORI platform integrates Cas13a with micro-well arrays to confine SARS-CoV-2 RNA at 0.1 copies µL−1 without enzymatic amplification [36].
Beyond nucleic acids, Collins and colleagues developed CrisprZyme, which combines Cas13 with nanozyme-immunoassays to generate peroxidase-mimicking metal nanoparticle reporters that function at room temperature and are compatible with colorimetric or lateral-flow readouts [37]. The platform supports multiplexed Cas effectors and has been clinically validated for acute myocardial infarction triage and prostate cancer detection, serving as a universal signal amplification module that substantially broadens the target landscape of CRISPR-based diagnostics.
A persistent limitation is the incompatibility of isothermal amplification templates with CRISPR nucleases in single-tube reactions, as trans-cleavage indiscriminately degrades the amplicon. Split-step protocols mitigate this loss but sacrifice simplicity and heighten aerosol contamination. Photocaged guide RNAs resolve this dilemma by enabling light-triggered activation that temporally separates amplification from CRISPR cleavage within a sealed vessel, eliminating contamination while maintaining a single-tube workflow [30,32,38,39].
To approach the ultimate sensitivity limit, CRISPR-based autocatalytic circuits have been engineered to convert a single binding event into an autocatalytic cascade. CALSA (CRISPR/Cas autocatalytic amplification) is a strategy in which LNA-modified split activators initiate a positive-feedback loop in which each Cas12a trans-cleavage event exposes a new activator, achieving a limit of detection of 10 aM for both ssDNA and genomic DNA [40]. Our laboratory further advanced this approach with CONAN (CRISPR/Cas-Only Amplification Network). Here, target dsDNA first activates a canonical Cas12a module whose trans-cleavage displaces an unprotected gRNA; this gRNA then primes a second Cas12a module carrying an auxiliary probe, triggering iterative activator generation and exponential fluorescence increase [41]. Although powerful, such autocatalytic circuits may amplify the background signal if off-target cleavage erroneously initiates the cascade. Rashid Bashir mitigated this risk with a CRISPR/Cas system that embeds two orthogonal Cas12a layers separated by a temperature-gated blocker-linker; kinetic optimization enables attomolar DNA detection in 10 min without pre-amplification [42]. The assay accurately discriminates methicillin-resistant staphylococcus aureus, methicillin-sensitive staphylococcus aureus, Escherichia coli, and HBV directly from blood cultures and implements a multi-input OR gate for syndromic diagnosis. Beyond fluorescence, plasmonic and nanopore transducers are now integrated; nanopore sensors convert Cas13a trans-cleavage activity into ion-current signatures that enable single-molecule RNA quantification. Continued optimization of reaction parameters, digital microfluidic readouts, and hybrid nanomaterial interfaces is establishing a new performance ceiling for amplification-free CRISPR diagnostics, offering instrument-independent pathogen identification, tumor-mutation tracking, and resistance-gene surveillance at the point of care.
While these platforms share the common principle of CRISPR-mediated signal amplification, their clinical trajectories have diverged considerably (Table 2). Platforms integrating isothermal amplification with lateral-flow detection (e.g., DETECTR, SHERLOCK) have achieved broader adoption, whereas amplification-free approaches, despite eliminating pre-amplification bias, are constrained by sophisticated instrumentation needs.
Table 2. Comparative overview of major CRISPR-based diagnostic platforms.
|
Platform |
Cas Effector |
Target |
Readout |
Key Advantages |
Major Limitations |
Clinical Status |
References |
|---|---|---|---|---|---|---|---|
|
SHERLOCK |
Cas13a |
RNA |
Fluorescence |
Attomolar sensitivity; high SNP resolution |
Multi-step workflow; initially qualitative |
Research and limited clinical use |
|
|
SHERLOCKv2 |
Cas13 |
RNA |
Multiplex fluorescence |
Quantitative and multiplexed detection |
Increased system complexity |
Early translational stage |
[20] |
|
DETECTR |
Cas12a |
DNA |
Fluorescence/ |
Simplified workflow; strong clinical compatibility |
Requires pre-amplification |
Advanced toward clinical use |
|
|
HOLMES |
Cas12a/Cas12b |
DNA/RNA |
Fluorescence |
High sensitivity and specificity |
Amplification-associated bias |
Validated in clinical samples |
|
|
Cas12b-based HPV detection |
Cas12b |
DNA |
Fluorescence |
Direct detection in human plasma |
Pathogen-specific design |
Under clinical validation |
[46] |
|
Amplification-free CRISPR-SNP chip |
Cas–gRNA |
DNA |
Electrical signal |
No amplification; real-time SNP discrimination |
High cost; complex instrumentation |
Preclinical stage |
[47] |
2. Precision Diagnostics and Clinical Deployment Scenarios of CRISPR/Cas Molecular Technologies
2.1. Precision Diagnosis of Infectious Diseases
Infectious diseases remain a persistent and emerging threat to global health. Conventional PCR-based pathogen detection provides high analytical sensitivity but is limited in multiplex capacity, making it unsuitable for complex polymicrobial infections or undifferentiated outbreaks. High-throughput sequencing offers comprehensive genomic coverage; however, its high cost, extended turnaround times, and substantial infrastructure requirements hinder rapid population-level deployment.
2.1.1. Rapid Pathogen Screening
The COVID-19 pandemic accelerated the transition of CRISPR-based diagnostics from bench to bedside. In May 2020, Sherlock Biosciences received the first U.S. FDA Emergency Use Authorization for a CRISPR-based SARS-CoV-2 assay built on Zhang Feng’s SHERLOCKv2 platform, which multiplexes Cas13, Cas12a, and Csm6 for parallel nucleic-acid detection [48,49,50]. The original workflow required sequential pipetting and off-cartridge sample processing, increasing contamination risk and turnaround time. Researchers subsequently integrated isothermal amplification and CRISPR-mediated cleavage into a single-tube reaction (STOPCovid.v1) using a thermostable Cas12b variant and a proprietary buffer, reducing hands-on time to less than 1 h [51]. STOPCovid.v2 further compressed the sample-to-answer time to 15 min by incorporating magnetic-bead enrichment into a self-contained microfluidic cartridge, enhancing both analytical sensitivity and clinical utility [50]. This progression from SHERLOCK to STOPCovid illustrates how iterative engineering can simultaneously improve portability, sensitivity, and operational simplicity, making CRISPR-based diagnostics suitable for resource-limited settings.
To overcome the throughput bottleneck that has long constrained multiplexed pathogen detection, the Sabeti laboratory fused the single-base precision of Cas13 with microfluidic miniaturization to create CARMEN (Combinatorial Arrayed Reactions for Multiplexed Evaluation of Nucleic acids) [52]. The platform interrogates 169 viral sequences within a single sample at femto-molar sensitivity, can be rapidly adapted to emerging pathogens, and decreases reagent costs >100-fold. However, CARMEN’s clinical translation was hindered by a large instrument footprint, limited sample capacity, and labor-intensive liquid handling. The team therefore unveiled mCARMEN, a Fluidigm microfluidic cartridge that processes 192 specimens (24-plex) or 96 specimens (96-plex) at less than $13 per test, while automated assay scripting reduces hands-on time to below five hours. Kinetic optimization lifts clinical sensitivity to 98.4% at 10 copies µL−1 and compresses turnaround to one hour. By integrating Cas13 and Cas12 modules, mCARMEN delivers absolute quantification of SARS-CoV-2 and influenza viral loads, providing surveillance data seven days ahead of sequencing during the Omicron surge. These iterative advances transform CARMEN from a research prototype into a clinical-grade, high-sensitivity, high-throughput platform that closes the technological gap for multiplex screening, variant tracking, and real-time epidemic response.
2.1.2. Drug-Resistance and Mutation Surveillance
Precision diagnostics enable rapid pathogen identification, yet qualitative or quantitative data alone offer limited guidance in the era of personalized antiviral therapy. Resistance profiling redirects the diagnostic focus from microbial detection to therapeutic prediction by characterizing genotypic resistance mutations, such as HIV integrase inhibitor resistance variants or phenotypic drug susceptibility, as exemplified by Mycobacterium tuberculosis drug-susceptibility testing (DST). These approaches guide dynamic and individualized treatment regimens, and CRISPR-based platforms are ideally positioned to close this therapeutic decision loop [53]. A case in point is the Treponema pallidum assay devised by Zheng and co-workers at Southern Medical University, which merges PCR with CRISPR/Cas13a to deliver a ten-fold lower limit of detection than conventional RT-PCR and achieves true single-molecule precision [54]. The test achieved 93.3% sensitivity and 100% specificity in a clinical cohort. It has since been extended to strain typing and surveillance of macrolide-resistance mutations, positioning it for post-therapeutic efficacy monitoring.
Similarly, Sabeti’s CARMEN architecture has been adapted for HIV drug-resistance surveillance [55]. A microarray of Cas13 effectors, each guided by crRNAs tiled across reverse-transcriptase and integrase genes, interrogates key resistance alleles such as K103N and M184V at single-nucleotide resolution. Using synthetic templates, all six RT and 21 integrase resistance mutations were correctly called with clinical-grade sensitivity and specificity.
When benchmarked against patient samples, CARMEN showed 90% concordance with Sanger sequencing and 86% with next-generation sequencing; an integrated mismatch filter maintains accuracy despite HIV hyper-variability. These features establish CARMEN as a low-cost, high-throughput platform for rapid resistance mapping, providing clinicians with actionable results to tailor antiviral therapy within hours of sample collection.
2.2. Molecular Subtyping of Human Germline Variants and Tumors
Circulating tumor DNA (ctDNA), consisting of fragmented tumor-derived DNA released into peripheral blood, has emerged as a pivotal liquid-biopsy biomarker for non-invasive cancer diagnosis, treatment guidance, and prognostic monitoring [56,57,58]. Although tissue biopsy remains the histopathological gold standard, its invasiveness, sampling bias, and potential complications preclude serial sampling and dynamic disease tracking. Liquid biopsy offers a powerful non-invasive approach to profiling tumour genomes; however, the extremely low abundance of ctDNA, typically present at femtomolar concentrations and vastly outnumbered by wild-type DNA, demands analytical platforms that combine attomolar sensitivity with single-base resolution.
To address this challenge, Huo and colleagues at Chongqing University developed an ultrasensitive, label-free CRISPR biosensor that uses Cas12a trans-cleavage to initiate an RNA–DNA chimeric exponential amplification reaction [59]. The resulting G-rich amplicons self-assemble into a dimeric G-quadruplex that binds thioflavin T, yielding more than 100-fold fluorescence enhancement and a limit of detection of 57 aM. The assay is applicable across multiple cancer types and requires only one primer pair per hotspot mutation. Chen and co-workers at Sichuan University proposed an orthogonal, amplification-free strategy by engineering a competitive dual-signal fluorescence sensor [60]. Target ctDNA activates Cas12a, which trans-cleaves a pre-formed G-quadruplex–N-methyl mesoporphyrin IX complex while catalyzing dopamine oxidation to polydopamine, thereby quenching residual NMM fluorescence. The fluorescence decrease scales linearly with ctDNA concentration, enabling marker-free, separation-free quantification with a 4 aM limit of detection within a 40 min workflow for breast-cancer-specific mutations. Both strategies integrate the single-base specificity of CRISPR/Cas12a with the peroxidase-mimicking signal amplification of G-quadruplex structures, eliminating the need for fluorophore–quencher labeling and maintaining robust performance in undiluted serum. The EXPAR-mediated route achieves attomolar sensitivity through multi-turnover catalysis, whereas the competitive-quenching format provides instrument-free operation suitable for low-resource settings. These complementary workflows advance ctDNA analysis to clinical-level sensitivity and portability, offering synergistic options for early cancer detection and real-time therapeutic monitoring.
MicroRNAs, post-transcriptional regulators of gene expression and mediators of intercellular communication, have emerged as critical biomarkers for human disease. Encapsulated within exosomal lipid bilayers, these approximately 22-nucleotide oligomers exhibit exceptional stability and disease-specific enrichment, making exosomal miRNAs prime targets for non-invasive liquid biopsy [61,62,63]. The gold-standard quantitative RT-PCR workflow, involving RNA extraction, reverse transcription, and pre-amplification, is labor-intensive, prone to sample loss, and poorly suited for point-of-care settings. CRISPR/Cas systems, which combine programmable specificity with built-in signal amplification, now provide an efficient alternative to these constraints [64,65,66]. Prof. Nie Zhou’s group at Hunan University integrated Cas12a into a catalytic hairpin assembly (CHA) circuit to create a dual-amplification engine [67]. Hybridization of the target miRNA to the CHA initiator generates a dsDNA activator that triggers Cas12a trans-cleavage of a fluorogenic reporter, driving the limit of detection into the sub-femtomolar range and enabling multiplexed profiling in a single cuvette. K. Chang and co-workers developed a “Logic-Measurer” cascade that couples a primer-exchange reaction with exonuclease III recycling through an OR gate to quantify miR-21 and miR-375 with a 2.1 fM limit of detection [68]. Evaluation of 315 breast-cancer biopsies achieved 87.3% diagnostic accuracy, underscoring its translational potential. Beyond bulk RNA, Zhang Kaixiang’s group delivered CRISPR/Cas13a directly into exosomes via lipid-mediated membrane fusion, enabling in situ quantification of exosomal miR-21 in undiluted plasma within 30 min [69]. The assay clearly distinguished breast cancer patients from healthy donors in a pilot cohort, highlighting its potential for real-time therapy monitoring. Collectively, these studies integrate single-base CRISPR precision with diverse enzyme-coupled amplification strategies, surpassing the sensitivity limit of conventional miRNA assays and accelerating the clinical translation of CRISPR-based liquid biopsies [70,71]. While Nie Zhou’s CHA circuit achieves superior sensitivity (sub-fM) and multiplexing in a single cuvette, it requires longer incubation times (typically >60 min) compared to Zhang’s exosome-targeted delivery (30 min), and lacks the extensive clinical validation (n = 315) demonstrated by Chang’s PER-Exo III cascade.
2.3. Multifunctional Sensing of Non-Nucleic-Acid Biomarkers
2.3.1. Protein Biomarker Detection
Accurate quantification of protein biomarkers underpins early diagnosis, treatment stratification, and mechanistic studies. However, conventional enzyme-linked immunosorbent assays (ELISA) are limited by restricted sensitivity, lengthy incubations, and inherent dependence on antibody stability, shortcomings that are amplified when analytes are scarce or matrices are complex. By introducing programmable nucleic-acid recognition and exponential signal amplification into protein detection, CRISPR/Cas systems are reshaping the analytical landscape, enabling label-free immunoassay workflows in which aptamers, nanobodies, or DNAzymes convert protein binding into CRISPR-activating nucleic-acid outputs with attomolar sensitivity.
Early diagnosis of Alzheimer’s disease hinges on the precise quantification of Aβ40 and Aβ42 in cerebrospinal fluid. Liu et al. addressed this proteomic challenge with a CRISPR/Cas12a-driven aptasensor, in which Aβ-aptamer binding induces a conformational change that releases a DNA activator, triggering Cas12a trans-cleavage and yielding an ultrasensitive readout of the Aβ42/Aβ40 ratio within minutes. The modular design provides a universal scaffold for any biomarker linked to an aptamer switch [72].
In cardiovascular emergencies, rapid triage of acute myocardial infarction (AMI) demands femtomolar quantification of cardiac troponin I. Zhang and co-workers at Nanjing University developed DATAS-Cas13d (Dual-Aptamer Transcription Amplification Strategy), in which simultaneous binding of two orthogonal aptamers to cTnI nucleates an in-vitro transcription module whose RNA output is sensed by Cas13 trans-cleavage activity, achieving 9.3 pM sensitivity in serum at half the cost of conventional ELISA [73]. The assay maintains a LOD of 0.67 ng·mL−1 in 10% human serum and discriminates cTnI from interferents proteins such as myoglobin, underscoring its utility in busy emergency departments.
Beyond aptamer-based sensors, CRISPR has been harnessed to monitor enzymatic activity directly. We recently introduced PR-Cas (Protease-Responsive CRISPR), a cascade that couples cancer-associated matrix metalloproteinase-2 (MMP-2) to Cas12a trans-cleavage [74]. An MMP-2-cleavable peptide tethers an RNA polymerase; a single proteolytic event releases the enzyme, triggering multi-copy gRNA transcription that fuels robust Cas12a collateral activity. The platform attains femtomolar sensitivity for MMP-2 in undiluted clinical serum and stratifies patients by metastatic burden, offering a new paradigm for real-time cancer-progression monitoring.
To confront the formidable challenge of quantifying trace proteins in viscous, matrix-rich specimens such as whole blood or sweat, Goldys and colleagues developed CAFI, a fiber-optic immunosensor augmented with CRISPR–Cas12a [75]. In this platform, an antibody–aptamer sandwich captures interferon-γ (IFN-γ), releasing a DNA activator that triggers Cas12a trans-cleavage of a fluorescent reporter. Using only 100 µL of sample, CAFI achieves a limit of detection of 1 fg·mL−1 (58.8 aM), representing over 1000-fold improvement compared with conventional ELISA while maintaining picomolar accuracy across undiluted serum, whole blood, and saliva. Swappable capture reagents endow the platform with broad protein versatility, underscoring the modular power of CRISPR-based protein sensing. By integrating aptamer-based sensors, protease-activity reporters, and fiber-optic transduction, CRISPR technology overcomes the traditional limitations of protein detection. Future marriage with hand-held readers, multiplexed biomarker panels, and real-time in-vivo probes will catalyze the clinical translation of CRISPR protein sensors, forging a foundational toolset for precision medicine at the point of care.
2.3.2. Antibody/Metabolite Detection
Accurate quantification of anti-SARS-CoV-2 antibodies is essential for evaluating post-infection immunity, vaccine effectiveness, and protective humoral responses in immunocompromised individuals. Conventional immunoassays, including ELISA and chemiluminescence platforms, suffer from limited sensitivity and labor-intensive workflows, making them ill-suited for point-of-care deployment [76,77,78,79,80,81]. More critically, these platforms frequently fail to detect the low-titre antibodies that characterize early infection or the blunted seroconversion observed in immunosuppressed patients after either natural infection or vaccination [79,82,83,84,85,86].
To overcome this analytical gap, Li and colleagues at Sichuan University developed UCAD (Ultrasensitive CRISPR-based Antibody Detection). This system converts antibody binding into a CRISPR/Cas12a-recognizable DNA barcode, achieving attomolar-level sensitivity [87]. The assay harnesses antibody-induced proximity hybridization to form a dsDNA template; after RPA, the amplicon triggers Cas12a trans-cleavage of a fluorogenic or lateral-flow readout. Testing 197 clinical sera (65 from vaccinated donors), UCAD demonstrated 100% sensitivity and 98.5% specificity. Remarkably, in 85 vaccinated kidney-transplant recipients, 85.9% classified as antibody-negative by standard chemiluminescence, UCAD readily quantified low-level antibodies and documented a ≥84.8% seroconversion rate following a third vaccine dose. This capability provides a critical tool for immune monitoring in immunocompromised populations and, as such, constitutes a technical cornerstone that enables both early-phase infection diagnosis and large-scale epidemiological investigations. The Francesco Ricci group has developed diverse DNA-engineered strategies for activating CRISPR/Cas12a signaling, including PAM-engineered toehold switch-mediated strand displacement reactions, synthetic antibody–DNA conjugates for direct activation, and the MAIGRET (Molecular Assay based on antibody-Induced Guide-RNA Enzymatic Transcription) system, which collectively enable the transduction of antibody–antigen recognition events into Cas enzyme collateral cleavage signals. These approaches significantly expand the scope of CRISPR-based detection beyond nucleic acids, enabling femtomolar-to-picomolar sensitivity for various targets, including antibodies (e.g., anti-digoxigenin, anti-HA, and Cetuximab), protein antigens (e.g., SARS-CoV-2 spike protein, EGFR, and MUC1), and small molecules (e.g., digoxigenin) in complex biological matrices [88,89,90].
Accurate quantification of small molecule metabolites such as uric acid and glucose is central to the management of gout, diabetes, and related chronic disorders. Conventional approaches, such as high performance liquid chromatography or invasive blood sampling followed by benchtop analysis, limit the feasibility of point-of-care testing. Zhang and co-workers addressed this limitation with CaT SMelor, an allosteric transcription factor (aTF) gated CRISPR sensor that converts small molecule binding into an amplifiable DNA output [91]. Uric-acid-mediated conformational switching of the aTF licenses RPA amplification of a Cas12a activator; subsequent trans-cleavage of a fluorogenic reporter yields a 10 nM limit of detection that correlates seamlessly with HPLC and clinical chemistry analyser readouts across 32 patient sera. The reagent cost (less than fifty cents per test) and ambient-temperature operation position CaT-SMelor as a field-forward alternative for resource-limited settings.
Building on this concept, a non-invasive CRISPR biosensor has been designed for painless glucose surveillance in tears and saliva [92]. Glucose oxidase (GOx) oxidizes glucose to gluconolactone with stoichiometric H2O2 release; the peroxide triggers self-cleavage of a pistol-like DNAzyme (PLDz), liberating a DNA fragment that activates Cas12a trans-cleavage of a quenched reporter. This cascade achieves 0.1 μM sensitivity, significantly outperforming conventional electrochemical glucose sensors while eliminating the need for finger-stick blood sampling. Collectively, these metabolite-to-DNA transducers exemplify how CRISPR/Cas systems can make chronic-disease monitoring accessible through amplification free handheld workflows.
CRISPR-based biomarker detection follows a universal design principle in which a molecular recognition element, such as an antibody, an allosteric transcription factor, a DNAzyme, or its synthetic analogue, converts a non-nucleic acid target into a programmable DNA cue that triggers the trans-cleavage cascade of CRISPR/Cas. UCAD initiates DNA probe hybridisation through antibody–antigen proximity, whereas CaT-SMelor uses ligand-induced conformational changes in an aTF to release a DNA template, and the glucose sensor links enzymatic H2O2 production to DNAzyme self-cleavage to furnish the required activator. This modular architecture retains attomolar sensitivity and single-base specificity across analyte classes and remains independent of the upstream recognition modality.
Developing next-generation affinity reagents such as aptamers, de novo protein binders, and artificial receptors will expand CRISPR diagnostics to tumour antigens, environmental toxins, and other emerging biomarkers. Concurrent integration with lateral-flow strips, microfluidic cartridges, and handheld readers should translate these molecular advances into household-ready devices suitable for resource-limited settings, establishing a versatile technological platform for precision medicine and global health surveillance.
In addition to clinical applications, CRISPR/Cas diagnostics are increasingly deployed in agriculture and environmental monitoring. Multiplex RT–RPA–Cas12 panels detect potato virus X, potato virus Y, wheat stripe rust, and Magnaporthe oryzae in a single sealed tube [93,94,95,96,97,98]. Attomolar sensitivity and single-base specificity are exploited to scan GMO inserts in food and feed, discriminate SNPs linked to herbicide resistance, and trace yield-determining loci in crops [99]. Environmental deployments extend the same molecular logic to quantify viral and bacterial load, heavy-metal ions, antibiotic residues, and pesticide contaminants in water, soil, and aerosol samples. These cross-sector successes illuminate a versatile future for CRISPR/Cas molecular diagnostics stretching from clinic to crop and riverbank to factory floor.
3. Future Perspectives on CRISPR-Based Diagnostics
3.1. Innovations in Miniaturization and Integration Technologies
The future development of CRISPR-based diagnostic technologies is expected to focus on miniaturization, high-throughput detection, and interdisciplinary integration, aiming to overcome current technological limitations and expand application scenarios. Miniaturization and portability represent key directions in the evolution of CRISPR/Cas diagnostic systems [100]. As an integrated platform, microfluidic chips adopt an integrated design concept that consolidates sample pretreatment, nucleic acid amplification, and CRISPR/Cas-based detection within a single device, thereby achieving complete automation and miniaturization. Fingernail-sized chips, for example, extract and detect HIV nucleic acids directly from whole blood [101,102]. Concurrently, smartphone-integrated readers and wearable form factors propel the field toward an “instrument-free” paradigm. The use of low-energy Bluetooth or Wi-Fi for streaming signals from various detection modalities (colorimetric, fluorometric, amperometric) to a mobile processor enables real-time, multi-modal image deconvolution via convolutional neural networks. This system minimizes hardware overhead and facilitates diagnostics in resource-limited environments [103].
Li and colleagues developed CLIPON (CRISPR and Large DNA assembly Induced Pregnancy strip for signal-ON detection), a portable platform that repurposes commercial pregnancy dipsticks for wash free, visual detection of both single and double stranded DNA in homogeneous solution [104]. A dedicated smartphone application and disposable microfluidic chip convert the visual readout into a quantitative result, delivering a fully portable assay. To maximize simplicity, Wang and collaborators developed SCOPE (Streamlined CRISPR On Pod Evaluation), a single-step platform that compresses sample-to-answer time to 12 min [105]. Rapid chemical lysis liberates viral nucleic acids from rash exudate, oral swabs, saliva, or urine within 2 min, followed by a one-pot RPA-Cas13a reaction that achieves monkeypox virus detection in 10 min. All steps, lysis, amplification, CRISPR cleavage, signal acquisition, and automated interpretation, which are executed on the palm-sized CPod device, offering a proper sample-in/answer-out solution for field diagnostics.
Wearable and real-time pathogen surveillance is transitioning from laboratory prototypes to a daily-life necessity. Collins and colleagues have seamlessly interwoven CRISPR chemistry into a 3-g, freeze-dried, cell-free synthetic-biology sensor that retains laboratory-grade sensitivity and specificity while embedded in everyday textiles [106]. Laminated within the fabric of an N95 respirator, the device passively captures SARS-CoV-2 aerosols, rehydrates on exhaled moisture, and completes a colorimetric Cas12 readout within 90 min without requiring an external power source. Encapsulated in soft silicone elastomers or woven fibres, the sensor continuously logs exposure events and transmits time-stamped data via low-energy Bluetooth to a smartphone gateway. By fusing micro-scale CRISPR reactors with ultra-low-power circuitry and cloud analytics, the platform compresses sample-to-answer time to under 30 min. It establishes a three-tier surveillance network spanning individual, community, and clinical levels, delivering instantaneous, high-resolution intelligence for outbreak containment, chronic-disease management, and personalized health tracking.
Amid the global COVID-19 pandemic, microfluidic technology has significantly advanced the performance of point-of-care testing (POCT) of SARS-CoV-2 assay through synergistic integration with nucleic acid analysis, immunoassays, and electrochemical biosensing platforms [107,108,109,110,111,112]. As a versatile multimodal platform, microfluidics accommodates nucleic-acid amplification, antigen-antibody recognition, and optoelectronic signal transduction within a miniaturized, automated framework. Notably, CRISPR-based microfluidic platforms, by precisely coupling molecular recognition with robust signal amplification, have substantially enhanced detection sensitivity and broadened the diagnostic scope [113,114].
Compared with conventional diagnostic workflows, microfluidic systems offer several advantages, including reduced reagent consumption, streamlined automation, and minimized risk of cross-contamination. Customized microfluidic designs have also addressed critical technical challenges associated with CRISPR-based assays [115]. For instance, paper-based microfluidic devices enable simple, colorimetric visual readouts, significantly simplifying assay procedures [116]. Enclosed microfluidic cartridges effectively prevent aerosol leakage, enhancing biosafety during testing [117,118,119,120]. Centrifugal or electrochemically integrated modules can consolidate nucleic acid extraction, purification, and detection into a single, standardized workflow [121,122]. Furthermore, by precisely manipulating discrete droplets, digital microfluidics enables single-molecule-level absolute quantification, pushing the limits of analytical sensitivity [123,124].
Current research efforts are increasingly focused on identifying and optimizing microfluidic architectures that align with the specific methodological requirements of different detection modalities, aiming to maximize automation, reproducibility, and diagnostic performance. This endeavor requires in-depth innovation, including microchannel topology design, fluidic control systems, and system-level engineering integration. In the future, CRISPR/Cas-powered microfluidic diagnostics, through interdisciplinary convergence, are poised to establish transformative technological paradigms in pathogen detection at the point of care, antimicrobial resistance gene screening, and precision medicine. These advances are expected to drive the evolution of in vitro diagnostics toward greater portability, intelligence, and networked connectivity, ultimately reshaping the landscape of global health diagnostics.
3.2. High-Throughput and Intelligent Diagnostics
As CRISPR diagnostic technology gradually advances toward clinical-grade testing, achieving high-throughput, multi-target analysis, and intelligent result interpretation has become a central direction for future development. Regarding multiplex detection, the CARMEN platform integrates Cas12/Cas13 hybrid systems with fluorescence-barcoded microfluidic chips to simultaneously identify 169 pathogens in a single reaction. Droplet microfluidic sorting technologies, such as Drop-CRISPR, partition samples into millions of microdroplets and, combined with the trans-cleavage activity of Cas proteins, enable single-molecule counting and multi-target genotyping, thereby markedly enhancing throughput and sensitivity. With the continuous development of artificial intelligence, especially breakthroughs in machine learning, CRISPR/Cas systems are shifting from basic research tools to precision, intelligent molecular diagnostic platforms, opening new avenues for innovation. Zhang Feng’s team developed the machine-learning model CRISPRscan, which predicts crRNA–target binding efficiency and off-target risk, guiding the design of high-affinity crRNAs and improving detection sensitivity by more than tenfold. Youchun Xu’s group at Tsinghua University proposed “mutaSCAN”, an extraction-free mutation-detection strategy [125] that combines sensitive RT-LAMP amplification with specific CRISPR/Cas12a detection in a single microfluidic chip unit. This approach uses an innovative lyophilization strategy to resolve temperature compatibility between the two steps and successfully integrates variable-throughput microfluidic chips. In addition, the workflow incorporates AI algorithms and user-friendly imaging devices to achieve high-throughput and automated result reading. mutaSCAN can detect 96 samples within 30 min at a sensitivity of 250 copies mL−1, enabling direct detection of SARS-CoV-2 and its variants from patient samples. The method demonstrates an ultrahigh-throughput analysis system using microfluidic multiwell plates, achieving rapid and accurate detection of SARS-CoV-2 and its variants with no false positives in negative samples. The research team also improved detection versatility and field applicability by developing a lysis buffer and optimizing microfluidic chip design. It achieved on-site automated result interpretation through AI-based image recognition algorithms. Clinical sample testing showed that mutaSCAN provides high positive and negative predictive concordance. Furthermore, synergistic innovation between bioinformatics and CRISPR technology offers a new approach for pathogen tracing. The deep integration of high-throughput and intelligent features significantly enhances the throughput, sensitivity, and accuracy of CRISPR diagnostics. It lays a solid foundation for its broad application in clinical medicine, disease surveillance, and public health [126,127,128,129,130].
Compared with other detection methods, CRISPR systems exhibit distinct advantages. First, owing to their high specificity and sensitivity, CRISPR systems achieve single-base resolution through precise crRNA–target nucleic acid pairing, enabling accurate discrimination between conserved pathogen sequences and host gene mutations. Second, CRISPR systems possess strong programmability and multifunctionality, allowing customized detection of diverse pathogens and biomarkers to meet diversified and personalized diagnostic needs. Their integrated detection capability also makes “sample-in, answer-out” automated workflows feasible, reducing cumbersome operational steps and lowering contamination risks. In addition, integrating CRISPR systems with artificial intelligence endows them with intelligent data-processing capacity, enabling rapid and accurate test results analysis. These advantages in specificity, sensitivity, programmability, integrated detection, and AI compatibility position CRISPR systems as powerful tools for high-throughput and intelligent diagnostic applications. As cross-disciplinary technologies advance, CRISPR diagnostic platforms are expected to evolve into intelligent molecular diagnostic systems that integrate detection, analysis, and decision-making.
3.3. Core Challenges in Clinical Translation
CRISPR-based diagnostics have demonstrated exceptional performance in research settings, yet their transition to the clinic remains constrained by a series of upstream bottlenecks, foremost among them the absence of a standardized, high-fidelity sample-preparation pipeline [25]. The extraction efficiency and purity of cell-free nucleic acids, such as circulating tumor DNA and viral RNA, directly influence detection sensitivity. However, extraction approaches such as magnetic bead-based and column-based methods show limited capacity to enrich low-abundance targets, resulting in suboptimal recovery rates. Moreover, heterogeneous pre-processing requirements for diverse specimens, including sputum, feces, and tissue sections, necessitate the development of universal lysis reagents and automated platforms. Nonetheless, most current CRISPR-based assays still rely on manual operations, thereby introducing variability in reproducibility.
Beyond sample-to-answer integration, the credibility of absolute quantification remains an equally formidable bottleneck. Digital CRISPR formats, exemplified by droplet-microfluidic single-molecule counting, achieve a limit of detection of 1 copy µL−1, yet they demand high-precision syringe pumps, high-numerical-aperture optics, and defect-free microfluidic masters, pushing instrument costs beyond 50,000 dollars, a price point incompatible with district hospitals or low-resource settings. The Quake-group dPCR–CRISPR hybrid reduces the hardware bill of materials to approximately 5000 dollars, but the long-term drift of the mandatory standard curve is still unresolved. Inter-batch variation in droplet volume, surface fluorophore adsorption, and thresholding algorithms routinely produces more than 25% relative standard deviation (RSD) in quantitative calls.
The core components of CRISPR-based assays include Cas proteins and crRNA, which must be stored at low temperatures to maintain activity and ensure the reliability of test results. Although lyophilization can extend their shelf life, Cas protein activity typically declines after rehydration, reducing detection accuracy. Because Cas proteins are susceptible to temperature, future studies could explore optimized storage conditions and reaction buffers, or the addition of protein stabilizers to prevent denaturation and degradation, thereby enhancing the stability and reliability of diagnostic systems [131]. In addition, the large-scale synthesis cost of crRNA far exceeds that of traditional PCR. Delays in regulatory approval and clinical validation further exacerbate the challenges of commercialization. Confronted by intertwined technical and economic bottlenecks, the field is turning to convergent, cross-disciplinary innovation as the most credible escape route. Systematic delays in navigating regulatory approval (averaging 18–36 months for novel biochemistry) and the absence of standardized clinical validation protocols further exacerbate commercialization challenges. A flagship example is mutaSCAN: a Cas12a-gated, high-throughput microfluidic platform that merges one-pot RT-LAMP amplification with convolutional-neural-network (CNN)-driven image analytics to deliver 96-plex results in 30 min. The system retains an analytical limit of 250 copies mL−1 across diverse clinical matrices while simultaneously profiling emergent mutations in the same workflow.
Equally instructive is the generative-design strategy introduced by Hayden C. Metsky and co-workers [132]. By retrofitting the ADAPT pipeline with a generative adversarial network coupled with an evolutionary algorithm, they established an in silico design environment that enables unconstrained exploration of crRNA sequence space, transcending the finite repertoire of naturally occurring viral genomes. The resulting software suite outputs synthetic guide sequences that intentionally harbour engineered mismatches yet preserve on-target activity. In head-to-head comparisons, these artificial crRNAs tripled the limit of detection relative to wild-type counterparts. They discriminated single-nucleotide variants of SARS-CoV-2 with 98% concordance against next-generation sequencing, thereby setting a new benchmark for precision CRISPR diagnostics.
CRISPR/Cas-driven molecular diagnostic technologies are rapidly moving from laboratory research to clinical application, with their core breakthrough being the realization of accurate “sample-in, answer-out” testing that meets the universal demand for equipment-free and rapid results [133,134]. Integrating nucleic acid isothermal amplification modules with CRISPR/Cas molecular recognition elements allows these platforms to screen multiple pathogens in complex biological samples without sophisticated instruments. Synergistic innovations in synthetic biology and artificial intelligence further enhance assay performance. Deep-learning-based crRNA design algorithms markedly improve the target specificity of Cas proteins, while Cas variants produced through directed evolution can elevate detection sensitivity to the single-copy level. Because of their inherently visual signal output, CRISPR biosensors show unique advantages in primary healthcare and at-home self-testing. Platforms built on lateral-flow strips, using colloidal gold nanoparticles or fluorescent probes, provide dual-mode colorimetric and fluorescence readouts, thereby overcoming the traditional dependence of molecular diagnostics on specialized instruments [135,136,137,138]. Clinical validation data show that home self-test strips for influenza A/B and Helicobacter pylori achieved more than 90% concordance in multicenter trials, and participants following standardized procedures could determine infection status within 15 min. This combination of operational simplicity and result reliability drives precision diagnostics toward a distributed model of community-based prescreening and at-home testing. With their visual and portable advantages, CRISPR test strips are well-suited for primary healthcare scenarios, and their complementary integration may create a complete POCT system covering community screening and precise pathogen subtyping.
The cross-integration of CRISPR/Cas biosensors with cutting-edge technologies such as nucleic acid design, nanomaterials, biochips, and artificial intelligence is opening new possibilities for molecular diagnostics and enabling the development of more efficient, intelligent, and versatile POCT devices. This convergence drives the evolution of next-generation diagnostic tools and lays the foundation for rapid molecular testing within POCT applications. To realize the full potential of CRISPR diagnostics in clinical practice and global precision medicine, it is essential to overcome persistent technical, economic, and regulatory bottlenecks. Otherwise, these constraints will significantly hinder the widespread application of CRISPR-based diagnostic technologies.
4. Conclusions
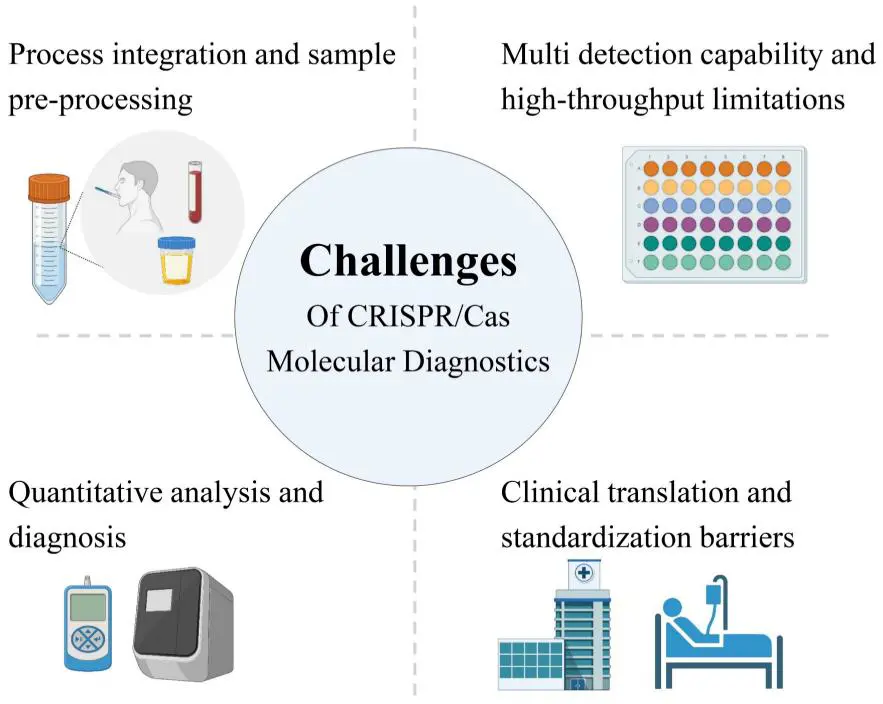
CRISPR–Cas systems have catalysed a paradigm shift in molecular biology and now stand at the forefront of next-generation diagnostics, poised to transform pathogen detection, genetic-variant profiling, and patient-tailored medicine [139,140]. Yet for all their promise, current assays still fail to converge on the quartet of clinical imperatives—high throughput, single-molecule sensitivity, pocket-sized hardware and dollar-scale unit costs. Therefore, a dispassionate audit of these persistent bottlenecks (Figure 3) and a roadmap for their future dissolution are indispensable to accelerate the bench-to-bedside journey of CRISPR-based diagnostics and unlock its full theoretical and practical potential [141,142,143].
Foremost among the hurdles to clinical translation is the imperfect specificity of CRISPR diagnostics. However, Cas effectors can be programmed through crRNA–target complementarity, complex biological matrices still yield measurable trans-cleavage [144,145]. The problem is exacerbated when discriminating near-identical pathogens or interrogating single-nucleotide variants, where a trans-cleavage base mismatch can tilt the balance between a true positive and a false positive call [146,147]. Dual gating recognition circuits, seed trimmed crRNAs, and high fidelity Cas variants have all shown promise in proof of concept studies, yet none have advanced beyond retrospective cohorts. Therefore, a rigorously validated, design rule based toolkit for suppressing mispriming and background noise remains an unmet and rate limiting clinical need.
Secondly, streamlining detection workflows and standardizing sample preprocessing are pivotal for translating CRISPR/Cas systems into practical point-of-care diagnostics. While the ideal molecular diagnostic platform would enable a “one-step” workflow, most current CRISPR-based assays still depend on a multi-step process involving nucleic acid extraction, amplification, and signal readout. These systems are commonly coupled with isothermal amplification techniques such as RPA or LAMP to achieve clinically relevant sensitivity. However, this multistage approach increases operational complexity and raises the risk of cross-contamination, limiting the feasibility of rapid deployment in resource-limited or field settings [148,149]. To overcome these challenges, future research should prioritize the development of highly integrated platforms such as microfluidic-based, fully enclosed detection chips or in situ reaction systems that eliminate the need for nucleic acid extraction, thereby advancing the realization of a true “sample-to-result” diagnostic paradigm.
Quantitative inadequacy remains a cardinal bottleneck constraining the clinical translation of CRISPR–Cas molecular diagnostics. Predominant end-point formats offer only binary or semi-quantitative readouts. They are blind to reaction kinetics, precluding precise viral-load monitoring, treatment-response surveillance, or any application requiring copy-number resolution. Coupling Cas-based assays with real-time fluorescence tracking, electrochemical transduction, or digital-droplet partitioning is an urgent priority; such integrations promise next-generation platforms that are simultaneously quantitative, high-throughput, and automation-compatible.
Multiplexing and massively parallel processing must evolve in parallel. Divergent cleavage kinetics among Cas orthologs, and the narrow reaction windows demanded by orthogonal targets, frustrate simultaneous multi-analyte detection. Although proof-of-concept strategies such as SHERLOCKv2, CARMEN, and their microfluidic-array, bar-coded, or spatially segregated progeny have demonstrated impressive scalability, they remain dependent on complex instrumentation and labor-intensive workflows, which are incompatible with decentralized testing. A convergent engineering agenda that unifies reaction chemistry, normalizes signal output, and standardizes readout modalities is thus essential if multiplex CRISPR diagnostics are to graduate from specialist laboratories to routine clinical use.
Notwithstanding the hurdles described above, the long-range outlook for CRISPR/Cas molecular diagnostics remains unequivocally promising. Evolutionary momentum is already evident, with an expanding arsenal of newly annotated Cas effectors differentiated by mass, cleavage mechanism, PAM plasticity, and thermal tolerance. This diversity has broadened the engineering design space, providing unprecedented freedom to develop lighter, faster, and more robust assays. Concurrently, machine-learning algorithms trained on million-sequence datasets are permeating every stratum of the development pipeline, accelerating guide-RNA optimisation, in-silico off-target filtering, kinetic modelling, and automated signal triage, while simultaneously shrinking user intervention and cycle time.
Beyond analytical performance, the strategic value of CRISPR diagnostics is becoming indelible. Attomolar sensitivity, minute footprint, and minimal power requirements converge to enable outbreak interception, personalised therapy titration, and decentralised testing in low-resource theatres. The COVID-19 pandemic provided an unplanned but decisive validation: field-forward CRISPR prototypes repeatedly delivered laboratory-grade sensitivity and specificity within 30 min, surpassing gold-standard RT-PCR in logistical agility and time-to-action, with speed translating directly into outbreak containment. Prospectively, more profound symbiosis with nanomaterials science, micro/nanofluidics, flexible electronics, and cloud-connected analytics is poised to engender a continuum of sensing ecosystems. These next-generation systems will evolve beyond episodic testing to function as continuous health monitors. By integrating real-time molecular surveillance, AI-powered early-warning algorithms for pathogen threats, and closed-loop therapeutic feedback, they will reposition CRISPR diagnostics from a reactive tool to a foundational component of proactive, precision public health.
In conclusion, CRISPR/Cas diagnostics have vaulted to the forefront of molecular medicine by uniting attomolar sensitivity, single-nucleotide specificity, and field-deployable simplicity in one cohesive platform. Future assays should balance sensitivity with cost and speed, as many diagnostic decisions do not benefit from attomolar limits. Ongoing engineering advances, driven by cross-disciplinary efforts, are expected to overcome existing challenges and enable these assays to transition from research tools to clinical standards. This will ultimately establish them as key components in global health security and precision therapeutics.
Statement of the Use of Generative AI and AI-Assisted Technologies in the Writing Process
During the preparation of this manuscript, the author(s) used Kimi and DeepSeek in order to assist with language editing and manuscript polishing. After using these tools, the author(s) reviewed and edited the content as needed and take(s) full responsibility for the content of the published article.
Acknowledgments
The authors thank all colleagues who provided technical assistance and helpful discussions during the preparation of this manuscript.
Author Contributions
Conceptualization, K.W.; Writing—Original Draft Preparation, H.R. and K.W.; Writing—Review & Editing, S.L., Y.Y. and P.C.
Ethics Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
No data was used for the research described in the article.
Funding
This research received no external funding.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- Makarova KS, Wolf YI, Iranzo J, Shmakov SA, Alkhnbashi OS, Brouns SJ, et al. Evolutionary classification of CRISPR–Cas systems: A burst of class 2 and derived variants. Nat. Rev. Microbiol. 2020, 18, 67–83. DOI:10.1038/s41579-019-0299-x [Google Scholar]
- Abudayyeh OO, Gootenberg JS, Essletzbichler P, Han S, Joung J, Belanto JJ, et al. RNA targeting with CRISPR–Cas13. Nature 2017, 550, 280–284. DOI:10.1038/nature24049 [Google Scholar]
- Zhou W, Hu L, Ying L, Zhao Z, Chu PK, Yu X-F. A CRISPR–Cas9-triggered strand displacement amplification method for ultrasensitive DNA detection. Nat. Commun. 2018, 9, 5012. DOI:10.1038/s41467-018-07324-5 [Google Scholar]
- Hu C, van Beljouw SPB, Nam KH, Schuler G, Ding F, Cui Y, et al. Craspase is a CRISPR RNA-guided, RNA-activated protease. Science 2022, 377, 1278–1285. DOI:10.1126/science.add5064 [Google Scholar]
- Strecker J, Demircioglu FE, Li D, Faure G, Wilkinson ME, Gootenberg JS, et al. RNA-activated protein cleavage with a CRISPR-associated endopeptidase. Science 2022, 378, 874–881. DOI:10.1126/science.add7450 [Google Scholar]
- Yu G, Wang X, Zhang Y, An Q, Wen Y, Li X, et al. Structure and function of a bacterial type III-E CRISPR–Cas7-11 complex. Nat. Microbiol. 2022, 7, 2078–2088. DOI:10.1038/s41564-022-01256-z [Google Scholar]
- Chen JS, Ma E, Harrington LB, Da Costa M, Tian X, Palefsky JM, et al. CRISPR-Cas12a target binding unleashes indiscriminate single-stranded DNase activity. Science 2018, 360, 436–439. DOI:10.1126/science.aar6245 [Google Scholar]
- Tong X, Zhang K, Han Y, Li T, Duan M, Ji R, et al. Fast and sensitive CRISPR detection by minimized interference of target amplification. Nat. Chem. Biol. 2024, 20, 885–893. DOI:10.1038/s41589-023-01534-9 [Google Scholar]
- Harrington LB, Burstein D, Chen JS, Paez-Espino D, Ma E, Witte IP, et al. Programmed DNA destruction by miniature CRISPR-Cas14 enzymes. Science 2018, 362, 839–842. DOI:10.1126/science.aav4294 [Google Scholar]
- Karvelis T, Bigelyte G, Young JK, Hou Z, Zedaveinyte R, Budre K, et al. PAM recognition by miniature CRISPR–Cas12f nucleases triggers programmable double-stranded DNA target cleavage. Nucleic Acids Res. 2020, 48, 5016–5023. DOI:10.1093/nar/gkaa208 [Google Scholar]
- Wu Z, Zhang Y, Yu H, Pan D, Wang Y, Wang Y, et al. Programmed genome editing by a miniature CRISPR-Cas12f nuclease. Nat. Chem. Biol. 2021, 17, 1132–1138. DOI:10.1038/s41589-021-00868-6 [Google Scholar]
- Wang Y, Wang Y, Pan D, Yu H, Zhang Y, Chen W, et al. Guide RNA engineering enables efficient CRISPR editing with a miniature Syntrophomonas palmitatica Cas12f1 nuclease. Cell Rep. 2022, 40, 111418. DOI:10.1016/j.celrep.2022.111418 [Google Scholar]
- Wang Y, Tang N, Ji Q. Systematic trans-Activity Comparison of Several Reported Cas12f Nucleases. Chin. J. Chem. 2025, 43, 1339–1347. DOI:10.1002/cjoc.202401325 [Google Scholar]
- Pausch P, Al-Shayeb B, Bisom-Rapp E, Tsuchida CA, Li Z, Cress BF, et al. CRISPR-CasΦ from huge phages is a hypercompact genome editor. Science 2020, 369, 333–337. DOI:10.1126/science.abb1400 [Google Scholar]
- Gootenberg JS, Abudayyeh OO, Lee JW, Essletzbichler P, Dy AJ, Joung J, et al. Nucleic acid detection with CRISPR-Cas13a/C2c2. Science 2017, 356, 438–442. DOI:10.1126/science.aam9321 [Google Scholar]
- Yoshimi K, Takeshita K, Yamayoshi S, Shibumura S, Yamauchi Y, Yamamoto M, et al. CRISPR-Cas3-based diagnostics for SARS-CoV-2 and influenza virus. iScience 2022, 25, 103830. DOI:10.1016/j.isci.2022.103830 [Google Scholar]
- Chen J, Chen Y, Huang L, Lin X, Chen H, Xiang W, et al. Trans-nuclease activity of Cas9 activated by DNA or RNA target binding. Nat. Biotechnol. 2025, 43, 558–568. DOI:10.1038/s41587-024-02255-7 [Google Scholar]
- Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. A programmable dual-RNA–guided DNA endonuclease in adaptive bacterial immunity. Science 2012, 337, 816–821. DOI:10.1126/science.1225829 [Google Scholar]
- Zhan T, Rindtorff N, Betge J, Ebert MP, Boutros M. CRISPR/Cas9 for cancer research and therapy. Semin. Cancer Biol. 2019, 55, 106–119. DOI:10.1016/j.semcancer.2018.04.001 [Google Scholar]
- Gootenberg JS, Abudayyeh OO, Kellner MJ, Joung J, Collins JJ, Zhang F. Multiplexed and portable nucleic acid detection platform with Cas13, Cas12a, and Csm6. Science 2018, 360, 439–444. DOI:10.1126/science.aaq0179 [Google Scholar]
- Knott GJ, East-Seletsky A, Cofsky JC, Holton JM, Charles E, O’Connell MR, et al. Guide-bound structures of an RNA-targeting A-cleaving CRISPR–Cas13a enzyme. Nat. Struct. Mol. Biol. 2017, 24, 825–833. DOI:10.1038/nsmb.3466 [Google Scholar]
- Wang B, Zhang T, Yin J, Yu Y, Xu W, Ding J, et al. Structural basis for self-cleavage prevention by tag: Anti-tag pairing complementarity in type VI Cas13 CRISPR systems. Mol. Cell 2021, 81, 1100–1115.e5. DOI:10.1016/j.molcel.2020.12.033 [Google Scholar]
- Li L, Li S, Wu N, Wu J, Wang G, Zhao G, et al. HOLMESv2: A CRISPR-Cas12b-assisted platform for nucleic acid detection and DNA methylation quantitation. ACS Synth. Biol. 2019, 8, 2228–2237. DOI:10.1021/acssynbio.9b00209 [Google Scholar]
- Pardee K, Green AA, Takahashi MK, Braff D, Lambert G, Lee JW, et al. Rapid, low-cost detection of Zika virus using programmable biomolecular components. Cell 2016, 165, 1255–1266. DOI: 10.1016/j.cell.2016.04.059 [Google Scholar]
- Li H, Xie Y, Chen F, Bai H, Xiu L, Zhou X, et al. Amplification-free CRISPR/Cas detection technology: Challenges, strategies, and perspectives. Chem. Soc. Rev. 2023, 52, 361–382. DOI:10.1039/D2CS00594H [Google Scholar]
- Myhrvold C, Freije CA, Gootenberg JS, Abudayyeh OO, Metsky HC, Durbin AF, et al. Field-deployable viral diagnostics using CRISPR-Cas13. Science 2018, 360, 444–448. DOI:10.1126/science.aas8836 [Google Scholar]
- Li SY, Cheng QX, Wang JM, Li XY, Zhang ZL, Gao S, et al. CRISPR-Cas12a-assisted nucleic acid detection. Cell Discov. 2018, 4, 20. DOI:10.1038/s41421-018-0028-z [Google Scholar]
- Wan Y, Li S, Xu W, Wang K, Guo W, Yang C, et al. Terminal chemical modifications of crRNAs enable improvement in the performance of CRISPR-Cas for point-of-care nucleic acid detection. Anal. Chem. 2024, 96, 16346–16354. DOI:10.1021/acs.analchem.4c03698 [Google Scholar]
- Yang H, Eremeeva E, Abramov M, Jacquemyn M, Groaz E, Daelemans D, et al. CRISPR-Cas9 recognition of enzymatically synthesized base-modified nucleic acids. Nucleic Acids Res. 2023, 51, 1501–1511. DOI:10.1093/nar/gkac1147 [Google Scholar]
- Hu M, Qiu Z, Bi Z, Tian T, Jiang Y, Zhou X. Photocontrolled crRNA activation enables robust CRISPR-Cas12a diagnostics. Proc. Natl. Acad. Sci. USA 2022, 119, e2202034119. DOI:10.1073/pnas.2202034119 [Google Scholar]
- Rossetti M, Merlo R, Bagheri N, Moscone D, Valenti A, Saha A, et al. Enhancement of CRISPR/Cas12a trans-cleavage activity using hairpin DNA reporters. Nucleic Acids Res. 2022, 50, 8377–8391. DOI:10.1093/nar/gkac578 [Google Scholar]
- Hu M, Liu R, Qiu Z, Cao F, Tian T, Lu Y, et al. Light-start CRISPR-Cas12a reaction with caged crRNA enables rapid and sensitive nucleic acid detection. Angew Chem. Int. Ed. 2023, 62, e202300663. DOI:10.1002/ange.202300663 [Google Scholar]
- Chen D, Huang W, Zhang Y, Chen B, Tan J, Yuan Q, et al. CRISPR-mediated profiling of viral RNA at single-nucleotide resolution. Angew Chem. Int. Ed. 2023, 62, e202304298. DOI:10.1002/anie.202304298 [Google Scholar]
- Tian T, Shu B, Jiang Y, Ye M, Liu L, Guo Z, et al. An ultralocalized Cas13a assay enables universal and nucleic acid amplification-free single-molecule RNA diagnostics. ACS Nano 2020, 15, 1167–1178. DOI:10.1021/acsnano.0c08165 [Google Scholar]
- Yue H, Shu B, Tian T, Xiong E, Huang M, Zhu D, et al. Droplet Cas12a assay enables DNA quantification from unamplified samples at the single-molecule level. Nano Lett. 2021, 21, 4643–4653. DOI:10.1021/acs.nanolett.1c00715 [Google Scholar]
- Shinoda H, Iida T, Makino A, Yoshimura M, Ishikawa J, Ando J, et al. Automated amplification-free digital RNA detection platform for rapid and sensitive SARS-CoV-2 diagnosis. Commun. Biol. 2022, 5, 473. DOI:10.1038/s42003-022-03433-6 [Google Scholar]
- Broto M, Kaminski MM, Adrianus C, Kim N, Greensmith R, Dissanayake-Perera S, et al. Nanozyme-catalysed CRISPR assay for preamplification-free detection of non-coding RNAs. Nat. Nanotechnol. 2022, 17, 1120–1126. DOI:10.1038/s41565-022-01179-0 [Google Scholar]
- Liu P, Lin Y, Zhuo X, Zeng J, Chen B, Zou Z, et al. Universal crRNA Acylation Strategy for Robust Photo-Initiated One-Pot CRISPR–Cas12a Nucleic Acid Diagnostics. Angew Chem. Int. Ed. 2024, 63, e202401486. DOI:10.1002/anie.202401486 [Google Scholar]
- Chen Y, Xu X, Wang J, Zhang Y, Zeng W, Liu Y, et al. Photoactivatable CRISPR/Cas12a Strategy for One-Pot DETECTR Molecular Diagnosis. Anal. Chem. 2022, 94, 9724–9731. DOI:10.1021/acs.analchem.2c01193 [Google Scholar]
- Sun K, Pu L, Chen C, Chen M, Li K, Li X, et al. An autocatalytic CRISPR-Cas amplification effect propelled by the LNA-modified split activators for DNA sensing. Nucleic Acids Res. 2024, 52, e39. DOI:10.1093/nar/gkae176 [Google Scholar]
- Shi K, Xie S, Tian R, Wang S, Lu Q, Gao D, et al. A CRISPR-Cas autocatalysis-driven feedback amplification network for supersensitive DNA diagnostics. Sci. Adv. 2021, 7, eabc7802. DOI:10.1126/sciadv.abc7802 [Google Scholar]
- Lim J, Van AB, Koprowski K, Wester M, Valera E, Bashir R. Amplification-free, OR-gated CRISPR-Cascade reaction for pathogen detection in blood samples. Proc. Natl. Acad. Sci. USA 2025, 122, e2420166122. DOI:10.1073/pnas.2420166122 [Google Scholar]
- Kellner MJ, Koob JG, Gootenberg JS, Abudayyeh OO, Zhang F. SHERLOCK: Nucleic acid detection with CRISPR nucleases. Nat. Protoc. 2019, 14, 2986–3012. DOI:10.1038/s41596-019-0210-2 [Google Scholar]
- Zhou X, Ye C, Xie M, Wei Y, Zhao Y, Liu X, et al. Advances in the application of CRISPR technology in pathogen detection: amplification-based and amplification-free strategies. Front Cell Infect Microbiol. 2025, 15, 1645699. DOI:10.3389/fcimb.2025.1645699 [Google Scholar]
- Xiao G, He X, Zhang S, Liu Y, Liang Z, Liu H, et al. Cas12a/guide RNA-based platform for rapid and accurate identification of major Mycobacterium species. J. Clin. Microbiol. 2020, 58, 10-1128. DOI:10.1128/jcm.01368-19 [Google Scholar]
- Dai Y, Somoza RA, Wang L, Welter JF, Li Y, Caplan AI, et al. Exploring the trans-cleavage activity of CRISPR-Cas12a (cpf1) for the development of a universal electrochemical biosensor. Angew. Chem. Int. Ed. Engl. 2019, 58, 17399–17405. DOI:10.1002/anie.201910772 [Google Scholar]
- Balderston S, Taulbee JJ, Celaya E, Fung K, Jiao A, Smith K, et al. Discrimination of single-point mutations in unamplified genomic DNA via Cas9 immobilized on a graphene field effect transistor. Nat. Biomed. Eng. 2021, 5, 713–725. DOI:10.1038/s41551-021-00706-z [Google Scholar]
- Tang Y, Gao L, Feng W, Guo C, Yang Q, Li F, et al. The CRISPR-Cas toolbox for analytical and diagnostic assay development. Chem. Soc. Rev. 2021, 50, 11844–11869. DOI:10.1039/d1cs00098e [Google Scholar]
- Broughton JP, Deng X, Yu G, Fasching CL, Servellita V, Singh J, et al. CRISPR–Cas12-based detection of SARS-CoV-2. Nat. Biotechnol. 2020, 38, 870–874. DOI:10.1038/s41587-020-0513-4 [Google Scholar]
- Joung J, Ladha A, Saito M, Kim N-G, Woolley AE, Segel M, et al. Detection of SARS-CoV-2 with SHERLOCK one-pot testing. N. Engl. J. Med. 2020, 383, 1492–1494. DOI:10.1056/NEJMc2026172 [Google Scholar]
- Joung J, Ladha A, Saito M, Segel M, Bruneau R, Huang M-LW, et al. Point-of-care testing for COVID-19 using SHERLOCK diagnostics. MedRxiv 2020. DOI:10.1101/2020.05.04.20091231 [Google Scholar]
- Welch NL, Zhu M, Hua C, Weller J, Mirhashemi ME, Nguyen TG, et al. Multiplexed CRISPR-based microfluidic platform for clinical testing of respiratory viruses and identification of SARS-CoV-2 variants. Nat. Med. 2022, 28, 1083–1094. DOI:10.1038/s41591-022-01734-1 [Google Scholar]
- Yan M-Y, Zheng D, Li S-S, Ding X-Y, Wang C-L, Guo X-P, et al. Application of combined CRISPR screening for genetic and chemical-genetic interaction profiling in Mycobacterium tuberculosis. Sci. Adv. 2022, 8, eadd5907. DOI:10.1126/sciadv.add5907 [Google Scholar]
- Chen W, Luo H, Zeng L, Pan Y, Parr JB, Jiang Y, et al. A suite of PCR-LwCas13a assays for detection and genotyping of Treponema pallidum in clinical samples. Nat. Commun. 2022, 13, 4671. DOI:10.1038/s41467-022-32250-y [Google Scholar]
- Ackerman CM, Myhrvold C, Thakku SG, Freije CA, Metsky HC, Yang DK, et al. Massively multiplexed nucleic acid detection with Cas13. Nature 2020, 582, 277–282. DOI:10.1038/s41586-020-2279-8 [Google Scholar]
- Wan JCM, Massie C, Garcia-Corbacho J, Mouliere F, Brenton JD, Caldas C, et al. Liquid biopsies come of age: towards implementation of circulating tumour DNA. Nat. Rev. Cancer 2017, 17, 223–238. DOI: 10.1038/nrc.2017.7 [Google Scholar]
- Underhill HR, Kitzman JO, Hellwig S, Welker NC, Daza R, Baker DN, et al. Fragment Length of Circulating Tumor DNA. PLoS Genet 2016, 12, e1006162. DOI:10.1371/journal.pgen.1006162 [Google Scholar]
- Merker JD, Oxnard GR, Compton C, Diehn M, Hurley P, Lazar AJ, et al. Circulating Tumor DNA Analysis in Patients With Cancer: American Society of Clinical Oncology and College of American Pathologists Joint Review. Arch. Pathol. Lab. Med. 2018, 142, 1242–1253. DOI:10.5858/arpa.2018-0901-SA [Google Scholar]
- Dong J, Li X, Deng L, Zhou S, Hou J, Hou C, et al. CRISPR/Cas12a cleavage-mediated isothermal amplification lights up the dimeric G-quadruplex signal unit for ultrasensitive and label-free detection of circulating tumor DNA. Sens. Actuators B Chem. 2024, 404, 135292. DOI:10.1016/j.snb.2024.135292 [Google Scholar]
- Zhao X, Liu W, Qu R, Jiang X, Chen J, Chen P. Target-activated CRISPR/Cas12a recognize multifunctional G-quadruplex and dual fluorescent indicators enable rapid non-extraction analysis of circulating tumor DNA in breast cancer. Sens. Actuators B Chem. 2025, 430, 137372. DOI:10.1016/j.snb.2025.137372 [Google Scholar]
- Kim VN. MicroRNA biogenesis: Coordinated cropping and dicing. Nat. Rev. Mol. Cell Biol. 2005, 6, 376–385. DOI:10.1038/nrm1644 [Google Scholar]
- Calura E, Fruscio R, Paracchini L, Bignotti E, Ravaggi A, Martini P, et al. miRNA Landscape in Stage I Epithelial Ovarian Cancer Defines the Histotype Specificities. Clin. Cancer Res. 2013, 19, 4114–4123. DOI:10.1158/1078-0432.CCR-13-0360 [Google Scholar]
- Zheng H, Zhang L, Zhao Y, Yang D, Song F, Wen Y, et al. Plasma miRNAs as Diagnostic and Prognostic Biomarkers for Ovarian Cancer. PLoS ONE 2013, 8, e77853. DOI:10.1371/journal.pone.0077853 [Google Scholar]
- Milanez-Almeida P, Martins AJ, Germain RN, Tsang JS. Cancer prognosis with shallow tumor RNA sequencing. Nat. Med. 2020, 26, 188–192. DOI:10.1038/s41591-019-0729-3 [Google Scholar]
- Tian B, Minero Gabriel Antonio S, Fock J, Dufva M, Hansen MF. CRISPR-Cas12a based internal negative control for nonspecific products of exponential rolling circle amplification. Nucleic Acids Res. 2020, 48, e30. DOI:10.1093/nar/gkaa017 [Google Scholar]
- Xing S, Lu Z, Huang Q, Li H, Wang Y, Lai Y, et al. An ultrasensitive hybridization chain reaction-amplified CRISPR-Cas12a aptasensor for extracellular vesicle surface protein quantification. Theranostics 2020, 10, 10262–10273. DOI:10.7150/thno.49047 [Google Scholar]
- Peng S, Tan Z, Chen S, Lei C, Nie Z. Integrating CRISPR-Cas12a with a DNA circuit as a generic sensing platform for amplified detection of microRNA. Chem. Sci. 2020, 11, 7362–7368. DOI:10.1039/d0sc03084h [Google Scholar]
- Xie Z, Zhao S, Deng R, Tang X, Feng L, Xie S, et al. Logic-Measurer: A Multienzyme-Assisted Ultrasensitive Circuit for Logical Detection of Exosomal MicroRNAs. ACS Nano 2025, 19, 12222–12236. DOI:10.1021/acsnano.5c00258 [Google Scholar]
- Zhang J, Guan M, Ma C, Liu Y, Lv M, Zhang Z, et al. Highly Effective Detection of Exosomal miRNAs in Plasma Using Liposome-Mediated Transfection CRISPR/Cas13a. ACS Sens. 2023, 8, 565–575. DOI:10.1021/acssensors.2c01683 [Google Scholar]
- Yang Q, Dong M-J, Xu J, Xing Y, Wang Y, Yang J, et al. CRISPR/RNA Aptamer System Activated by an AND Logic Gate for Biomarker-Driven Theranostics. J. Am. Chem. Soc. 2025, 147, 169–180. DOI:10.1021/jacs.4c08719 [Google Scholar]
- Yan H, Wen Y, Tian Z, Hart N, Han S, Hughes SJ, et al. A one-pot isothermal Cas12-based assay for the sensitive detection of microRNAs. Nat. Biomed. Eng. 2023, 7, 1583–1601. DOI:10.1038/s41551-023-01033-1 [Google Scholar]
- Jia Z, Maghaydah Y, Zdanys K, Kuchel GA, Diniz BS, Liu C. CRISPR-Powered Aptasensor for Diagnostics of Alzheimer’s Disease. ACS Sens. 2024, 9, 398–405. DOI:10.1021/acssensors.3c02167 [Google Scholar]
- Feng Z-Y, Liu R, Li X, Zhang J. Harnessing the CRISPR-Cas13d System for Protein Detection by Dual-Aptamer-Based Transcription Amplification. Chem. Eur. J. 2023, 29, e202202693. DOI:10.1002/chem.202202693 [Google Scholar]
- Liu F, Chen R, Song W, Li L, Lei C, Nie Z. Modular Combination of Proteolysis-Responsive Transcription and Spherical Nucleic Acids for Smartphone-Based Colorimetric Detection of Protease Biomarkers. Anal. Chem. 2021, 93, 3517–3525. DOI:10.1021/acs.analchem.0c04894 [Google Scholar]
- Deng F, Li Y, Qiao L, Goldys E. A CRISPR/Cas12a-assisted on-fibre immunosensor for ultrasensitive small protein detection in complex biological samples. Anal. Chim. Acta 2022, 1192, 339351. DOI:10.1016/j.aca.2021.339351 [Google Scholar]
- Ainsworth M, Andersson M, Auckland K, Baillie JK, Barnes E, Beer S, et al. Performance characteristics of five immunoassays for SARS-CoV-2: A head-to-head benchmark comparison. Lancet Infect. Dis. 2020, 20, 1390–1400. DOI:10.1016/s1473-3099(20)30634-4 [Google Scholar]
- GeurtsvanKessel CH, Okba NMA, Igloi Z, Bogers S, Embregts CWE, Laksono BM, et al. An evaluation of COVID-19 serological assays informs future diagnostics and exposure assessment. Nat. Commun. 2020, 11, 3436. DOI:10.1038/s41467-020-17317-y [Google Scholar]
- Rhoads DD, Cherian SS, Roman K, Stempak LM, Schmotzer CL, Sadri N. Comparison of Abbott ID Now, DiaSorin Simplexa, and CDC FDA Emergency Use Authorization Methods for the Detection of SARS-CoV-2 from Nasopharyngeal and Nasal Swabs from Individuals Diagnosed with COVID-19. J. Clin. Microbiol. 2020, 58, e00760-20. DOI:10.1128/jcm.00760-20 [Google Scholar]
- Whitman JD, Hiatt J, Mowery CT, Shy BR, Yu R, Yamamoto TN, et al. Evaluation of SARS-CoV-2 serology assays reveals a range of test performance. Nat. Biotechnol. 2020, 38, 1174–1183. DOI:10.1038/s41587-020-0659-0 [Google Scholar]
- Elledge SK, Zhou XX, Byrnes JR, Martinko AJ, Lui I, Pance K, et al. Engineering luminescent biosensors for point-of-care SARS-CoV-2 antibody detection. Nat. Biotechnol. 2021, 39, 928–935. DOI:10.1038/s41587-021-00878-8 [Google Scholar]
- Yao Z, Drecun L, Aboualizadeh F, Kim SJ, Li Z, Wood H, et al. A homogeneous split-luciferase assay for rapid and sensitive detection of anti-SARS CoV-2 antibodies. Nat. Commun. 2021, 12, 1806. DOI:10.1038/s41467-021-22102-6 [Google Scholar]
- Long Q-X, Liu B-Z, Deng H-J, Wu G-C, Deng K, Chen Y-K, et al. Antibody responses to SARS-CoV-2 in patients with COVID-19. Nat. Med. 2020, 26, 845–848. DOI:10.1038/s41591-020-0897-1 [Google Scholar]
- Agnolon V, Contato A, Meneghello A, Tagliabue E, Toffoli G, Gion M, et al. ELISA assay employing epitope-specific monoclonal antibodies to quantify circulating HER2 with potential application in monitoring cancer patients undergoing therapy with trastuzumab. Sci. Rep. 2020, 10, 3016. DOI:10.1038/s41598-020-59630-y [Google Scholar]
- Zhang W, Du RH, Li B, Zheng XS, Yang XL, Hu B, et al. Molecular and serological investigation of 2019-nCoV infected patients: Implication of multiple shedding routes. Emerg. Microbes Infect. 2020, 9, 386–389. DOI:10.1080/22221751.2020.1729071 [Google Scholar]
- Acharya AP, Nafisi PM, Gardner A, MacKay JL, Kundu K, Kumar S, et al. A fluorescent peroxidase probe increases the sensitivity of commercial ELISAs by two orders of magnitude. Chem. Commun. 2013, 49, 10379–10381. DOI:10.1039/C3CC44783A [Google Scholar]
- Carter QL, Dotzlaf J, Swearingen C, Brittain I, Chambers M, Duffin K, et al. Development and characterization of a novel ELISA based assay for the quantitation of sub-nanomolar levels of neoepitope exposed NITEGE-containing aggrecan fragments. J. Immunol. Methods 2007, 328, 162–168. DOI:10.1016/j.jim.2007.08.018 [Google Scholar]
- Tang Y, Song T, Gao L, Yin S, Ma M, Tan Y, et al. A CRISPR-based ultrasensitive assay detects attomolar concentrations of SARS-CoV-2 antibodies in clinical samples. Nat. Commun. 2022, 13, 4667. DOI:10.1038/s41467-022-32371-4 [Google Scholar]
- Miceli F, Bracaglia S, Sorrentino D, Porchetta A, Ranallo S, Ricci F. MAIGRET: A CRISPR-based immunoassay that employs antibody-induced cell-free transcription of CRISPR guide RNA strands. Nucleic Acids Res. 2025, 53, gkaf238. DOI:10.1093/nar/gkaf238 [Google Scholar]
- Paialunga E, Bagheri N, Rossetti M, Fabiani L, Micheli L, Chamorro-Garcia A, et al. Leveraging Synthetic Antibody-DNA Conjugates to Expand the CRISPR-Cas12a Biosensing Toolbox. ACS Synth. Biol. 2025, 14, 171–178. DOI:10.1021/acssynbio.4c00541 [Google Scholar]
- Bagheri N, Chamorro A, Idili A, Porchetta A. PAM-Engineered Toehold Switches as Input-Responsive Activators of CRISPR-Cas12a for Sensing Applications. Angew. Chem. Int. Ed. Engl. 2024, 63, e202319677. DOI:10.1002/anie.202319677 [Google Scholar]
- Liang M, Li Z, Wang W, Liu J, Liu L, Zhu G, et al. A CRISPR-Cas12a-derived biosensing platform for the highly sensitive detection of diverse small molecules. Nat. Commun. 2019, 10, 3672. DOI:10.1038/s41467-019-11648-1 [Google Scholar]
- Zhang C, Yao H, Ma Q, Yu B. Ultrasensitive glucose detection from tears and saliva through integrating a glucose oxidase-coupled DNAzyme and CRISPR–Cas12a. Analyst 2021, 146, 6576–6581. DOI:10.1039/D1AN01385H [Google Scholar]
- Wheatley MS, Wang Q, Wei W, Bottner-Parker KD, Zhao Y, Yang Y. Cas12a-Based Diagnostics for Potato Purple Top Disease Complex Associated with Infection by ‘Candidatus Phytoplasma trifolii’-Related Strains. Plant Dis. 2022, 106, 2039–2045. DOI:10.1094/pdis-09-21-2119-re [Google Scholar]
- Aman R, Mahas A, Marsic T, Hassan N, Mahfouz MM. Efficient, rapid, and sensitive detection of plant RNA viruses with one-pot RT-RPA–CRISPR/Cas12a assay. Front. Microbiol. 2020, 11, 610872. DOI:10.3389/fmicb.2020.610872 [Google Scholar]
- Marqués M-C, Sánchez-Vicente J, Ruiz R, Montagud-Martínez R, Márquez-Costa R, Gómez G, et al. Diagnostics of Infections Produced by the Plant Viruses TMV, TEV, and PVX with CRISPR-Cas12 and CRISPR-Cas13. ACS Synth. Biol. 2022, 11, 2384–2393. DOI:10.1021/acssynbio.2c00090 [Google Scholar]
- Mahas A, Hassan N, Aman R, Marsic T, Wang Q, Ali Z, et al. LAMP-Coupled CRISPR–Cas12a Module for Rapid and Sensitive Detection of Plant DNA Viruses. Viruses 2021, 13, 466. DOI:10.3390/v13030466 [Google Scholar]
- Ramachandran V, Weiland JJ, Bolton MD. CRISPR-based isothermal next-generation diagnostic method for virus detection in sugarbeet. Front. Microbiol. 2021, 12, 679994. DOI:10.3389/fmicb.2021.679994 [Google Scholar]
- Kang H, Peng Y, Hua K, Deng Y, Bellizzi M, Gupta DR, et al. Rapid Detection of Wheat Blast Pathogen Magnaporthe oryzae Triticum Pathotype Using Genome-Specific Primers and Cas12a-mediated Technology. Engineering 2021, 7, 1326–1335. DOI:10.1016/j.eng.2020.07.016 [Google Scholar]
- Gong X-Y, Wang Z-H, Bashir M, Tang T, Gan X, Yang W-C. Recent application of CRISPR/Cas in plant disease detection. TrAC Trends Anal. Chem. 2025, 189, 118251. DOI:10.1016/j.trac.2025.118251 [Google Scholar]
- Liu G. Advancing CRISPR/Cas Biosensing with Integrated Devices. ACS Sens. 2025, 10, 575–576. DOI:10.1021/acssensors.5c00330 [Google Scholar]
- Li X, Wang T, Liu X, Jiang H, Wang X. Advances of engineered microfluidic biosensors via CRISPR/Cas in bacteria and virus monitoring. Chem. Eng. J. 2024, 491, 152038. DOI:10.1016/j.cej.2024.152038 [Google Scholar]
- Li Z, Uno N, Ding X, Avery L, Banach D, Liu C. Bioinspired CRISPR-Mediated Cascade Reaction Biosensor for Molecular Detection of HIV Using a Glucose Meter. ACS Nano 2023, 17, 3966–3975. DOI:10.1021/acsnano.2c12754 [Google Scholar]
- Ge H, Feng J, Huang L, Luo Z, Ling H, Ma L, et al. Development of a highly sensitive, high-throughput and automated CRISPR-based device for the contamination-free pathogen detection. Biosens. Bioelectron. 2025, 278, 117323. DOI:10.1016/j.bios.2025.117323 [Google Scholar]
- Tang Y, Qi L, Liu Y, Guo L, Zhao R, Yang M, et al. CLIPON: A CRISPR-enabled strategy that turns commercial pregnancy test strips into general Point-of-Need test devices. Angew. Chem. Int. Ed. 2022, 61, e202115907. DOI:10.1002/anie.202115907 [Google Scholar]
- Wang Y, Chen H, Lin K, Han Y, Gu Z, Wei H, et al. Ultrasensitive single-step CRISPR detection of monkeypox virus in minutes with a vest-pocket diagnostic device. Nat. Commun. 2024, 15, 3279. DOI:10.1038/s41467-024-47518-8 [Google Scholar]
- Nguyen PQ, Soenksen LR, Donghia NM, Angenent-Mari NM, de Puig H, Huang A, et al. Wearable materials with embedded synthetic biology sensors for biomolecule detection. Nat. Biotechnol. 2021, 39, 1366–1374. DOI:10.1038/s41587-021-00950-3 [Google Scholar]
- Ji M, Xia Y, Loo J, Li L, Ho H-P, He J, et al. Automated multiplex nucleic acid tests for rapid detection of SARS-CoV-2, influenza A and B infection with direct reverse-transcription quantitative PCR (dirRT-qPCR) assay in a centrifugal microfluidic platform. RSC Adv. 2020, 10, 34088–34098. DOI:10.1039/D0RA04507A [Google Scholar]
- Lin Z, Zou Z, Pu Z, Wu M, Zhang Y. Application of microfluidic technologies on COVID-19 diagnosis and drug discovery. Acta Pharm. Sin. B 2023, 13, 2877–2896. DOI:10.1016/j.apsb.2023.02.014 [Google Scholar]
- Chen Y, Zong N, Ye F, Mei Y, Qu J, Jiang X. Dual-CRISPR/Cas12a-Assisted RT-RAA for Ultrasensitive SARS-CoV-2 Detection on Automated Centrifugal Microfluidics. Anal. Chem. 2022, 94, 9603–9609. DOI:10.1021/acs.analchem.2c00638 [Google Scholar]
- Lim J, Ahn JW, Maeng I, Lee J, Kim R, Mun B, et al. TwinDemic detection: A non-enzymatic signal amplification system for on-site detection of multiple respiratory viruses. Sens. Actuators B Chem. 2025, 424, 136933. DOI:10.1016/j.snb.2024.136933 [Google Scholar]
- Nguyen HQ, Nguyen VD, Phan VM, Seo TS. Development of a self-contained microfluidic chip and an internet-of-things-based point-of-care device for automated identification of respiratory viruses. Lab Chip 2024, 24, 2485–2496. DOI:10.1039/D3LC00933E [Google Scholar]
- Nguyen PQM, Wang M, Ann Maria N, Li AY, Tan HY, Xiong GM, et al. Modular micro-PCR system for the onsite rapid diagnosis of COVID-19. Microsyst. Nanoeng. 2022, 8, 82. DOI:10.1038/s41378-022-00400-3 [Google Scholar]
- Shan X, Gong F, Yang Y, Qian J, Tan Z, Tian S, et al. Nucleic Acid Amplification-Free Digital Detection Method for SARS-CoV-2 RNA Based on Droplet Microfluidics and CRISPR–Cas13a. Anal. Chem. 2023, 95, 16489–16495. DOI:10.1021/acs.analchem.3c02007 [Google Scholar]
- Peng R, Lu Z, Liu M, Hu F. RT-RPA-assisted CRISPR/Cas12a for rapid and multiplex detection of respiratory infectious viruses based on centrifugal microfluidics. Sens. Actuators B Chem. 2024, 399, 134838. DOI:10.1016/j.snb.2023.134838 [Google Scholar]
- Chen J, Yang D, Ji D, Guo B, Guo Y, Lin H, et al. A Fully Automated Point-of-Care Device Using Organic Electrochemical Transistor-Enhanced CRISPR/Cas12a for Amplification-Free Nucleic Acid Detection. Adv. Funct. Mater. 2025, 35, 2420701. DOI:10.1002/adfm.202420701 [Google Scholar]
- Xu J, Suo W, Goulev Y, Sun L, Kerr L, Paulsson J, et al. Handheld Microfluidic Filtration Platform Enables Rapid, Low-Cost, and Robust Self-Testing of SARS-CoV-2 Virus. Small 2021, 17, e2104009. DOI:10.1002/smll.202104009 [Google Scholar]
- Wang D, Wang X, Ye F, Zou J, Qu J, Jiang X. An Integrated Amplification-Free Digital CRISPR/Cas-Assisted Assay for Single Molecule Detection of RNA. ACS Nano 2023, 17, 7250–7256. DOI:10.1021/acsnano.2c10143 [Google Scholar]
- Yin B, Wan X, Sohan ASMMF, Lin X. Microfluidics-Based POCT for SARS-CoV-2 Diagnostics. Micromachines 2022, 13, 1238. DOI:10.3390/mi13081238 [Google Scholar]
- Li Z, Ding X, Yin K, Avery L, Ballesteros E, Liu C. Instrument-free, CRISPR-based diagnostics of SARS-CoV-2 using self-contained microfluidic system. Biosens. Bioelectron. 2022, 199, 113865. DOI:10.1016/j.bios.2021.113865 [Google Scholar]
- Fu Q, Tu Y, Cheng L, Zhang L, Qiu X. A fully-enclosed prototype ‘pen’ for rapid detection of SARS-CoV-2 based on RT-RPA with dipstick assay at point-of-care testing. Sens. Actuators B Chem. 2023, 383, 133531. DOI:10.1016/j.snb.2023.133531 [Google Scholar]
- Xiao Y, Zhou M, Liu C, Gao S, Wan C, Li S, et al. Fully integrated and automated centrifugal microfluidic chip for point-of-care multiplexed molecular diagnostics. Biosens. Bioelectron. 2024, 255, 116240. DOI:10.1016/j.bios.2024.116240 [Google Scholar]
- Li P, Xiong H, Yang B, Jiang X, Kong J, Fang X. Recent progress in CRISPR-based microfluidic assays and applications. TrAC Trends Anal. Chem. 2022, 157, 116812. DOI:10.1016/j.trac.2022.116812 [Google Scholar]
- Cui JQ, Liu FX, Park H, Chan KW, Leung T, Tang BZ, et al. Droplet digital recombinase polymerase amplification (ddRPA) reaction unlocking via picoinjection. Biosens. Bioelectron. 2022, 202, 114019. DOI:10.1016/j.bios.2022.114019 [Google Scholar]
- Liu FX, Cui JQ, Park H, Chan KW, Leung T, Tang BZ, et al. Isothermal Background-Free Nucleic Acid Quantification by a One-Pot Cas13a Assay Using Droplet Microfluidics. Anal. Chem. 2022, 94, 5883–5892. DOI:10.1021/acs.analchem.2c00067 [Google Scholar]
- Zhang L, Wang H, Yang S, Liu J, Li J, Lu Y, et al. High-Throughput and Integrated CRISPR/Cas12a-Based Molecular Diagnosis Using a Deep Learning Enabled Microfluidic System. ACS Nano 2024, 18, 24236–24251. DOI:10.1021/acsnano.4c05734 [Google Scholar]
- Li X, Liu M, Men D, Duan Y, Deng L, Zhou S, et al. Rapid, portable, and sensitive detection of CaMV35S by RPA-CRISPR/Cas12a-G4 colorimetric assays with high accuracy deep learning object recognition and classification. Talanta 2024, 278, 126441. DOI:10.1016/j.talanta.2024.126441 [Google Scholar]
- Zhao J, Kong D, Zhang G, Zhang S, Wu Y, Dai C, et al. An Efficient CRISPR/Cas Cooperative Shearing Platform for Clinical Diagnostics Applications. Angew. Chem. Int. Ed. 2024, 63, e202411705. DOI:10.1002/anie.202411705 [Google Scholar]
- Huang B, Guo L, Yin H, Wu Y, Zeng Z, Xu S, et al. Deep learning enhancing guide RNA design for CRISPR/Cas12a-based diagnostics. Imeta 2024, 3, e214. DOI:10.1002/imt2.214 [Google Scholar]
- Wessels H-H, Stirn A, Méndez-Mancilla A, Kim EJ, Hart SK, Knowles DA, et al. Prediction of on-target and off-target activity of CRISPR–Cas13d guide RNAs using deep learning. Nat. Biotechnol. 2024, 42, 628–637. DOI:10.1038/s41587-023-01830-8 [Google Scholar]
- Xu T, Zhang Y, Li S, Dai C, Wei H, Chen D, et al. Deep Learning-Enhanced Hand-Driven Microfluidic Chip for Multiplexed Nucleic Acid Detection Based on RPA/CRISPR. Adv. Sci. 2025, 12, 2414918. DOI:10.1002/advs.202414918 [Google Scholar]
- Li Z, Zhao W, Ma S, Li Z, Yao Y, Fei T. A chemical-enhanced system for CRISPR-Based nucleic acid detection. Biosens. Bioelectron. 2021, 192, 113493. DOI:10.1016/j.bios.2021.113493 [Google Scholar]
- Mantena S, Pillai PP, Petros BA, Welch NL, Myhrvold C, Sabeti PC, et al. Model-directed generation of artificial CRISPR–Cas13a guide RNA sequences improves nucleic acid detection. Nat. Biotechnol. 2025, 43, 1266–1273. DOI:10.1038/s41587-024-02422-w [Google Scholar]
- Huang Z, Lyon CJ, Wang J, Lu S, Hu TY. CRISPR Assays for Disease Diagnosis: Progress to and Barriers Remaining for Clinical Applications. Adv. Sci. 2023, 10, 2301697. DOI:10.1002/advs.202301697 [Google Scholar]
- Weng Z, You Z, Yang J, Mohammad N, Lin M, Wei Q, et al. CRISPR-Cas Biochemistry and CRISPR-Based Molecular Diagnostics. Angew. Chem. Int. Ed. 2023, 62, e202214987. DOI:10.1002/anie.202214987 [Google Scholar]
- Zhou T, Shen G, Zhong L, Chen G, Meng L, He W, et al. crRNA array-mediated CRISPR/Cas12a coupling with dual RPA for highly sensitive detection of Streptomyces aureofaciens Tü117 from hypertension with multi-signal output. Biosens. Bioelectron. 2025, 282, 117493. DOI:10.1016/j.bios.2025.117493 [Google Scholar]
- Mao Z, Chen R, Huang L, Ren S, Liu B, Gao Z. CRISPR analysis based on Pt@MOF dual-modal signal for multichannel fluorescence and visual detection of norovirus. Biosens. Bioelectron. 2025, 273, 117153. DOI:10.1016/j.bios.2025.117153 [Google Scholar]
- Hu M, Yuan C, Tian T, Wang X, Sun J, Xiong E, et al. Single-Step, Salt-Aging-Free, and Thiol-Free Freezing Construction of AuNP-Based Bioprobes for Advancing CRISPR-Based Diagnostics. J. Am. Chem. Soc. 2020, 142, 7506–7513. DOI:10.1021/jacs.0c00217 [Google Scholar]
- Xiong E, Jiang L, Tian T, Hu M, Yue H, Huang M, et al. Simultaneous Dual-Gene Diagnosis of SARS-CoV-2 Based on CRISPR/Cas9-Mediated Lateral Flow Assay. Angew. Chem. Int. Ed. 2021, 60, 5307–5315. DOI:10.1002/anie.202014506 [Google Scholar]
- Abudayyeh OO, Gootenberg JS. CRISPR diagnostics. Science 2021, 372, 914–915. DOI:10.1126/science.abi9335 [Google Scholar]
- Zuo X, Fan C, Chen H-Y. Biosensing: CRISPR-powered diagnostics. Nat. Biomed. Eng. 2017, 1, 0091. DOI:10.1038/s41551-017-0091 [Google Scholar]
- Du Y, Gao H, Liu J, Liu X, Xing Z, Zhang T, et al. Research progress of the CRISPR-Cas system in the detecting pathogen nucleic acids. Synth. Biol. J. 2024, 5, 202–216. DOI:10.12211/2096-8280.2022-068 [Google Scholar]
- Li JC, Xiao MJ, Chen YP. Progress in In Vitro CRISPR/Cas-based Nucleic Acid Detection. China Biotechnol. 2024, 44, 73–87. DOI:10.13523/j.cb.2401044 [Google Scholar]
- Pacesa M, Pelea O, Jinek M. Past, present, and future of CRISPR genome editing technologies. Cell 2024, 187, 1076–1100. DOI:10.1016/j.cell.2024.01.042 [Google Scholar]
- Teng F, Guo L, Cui T, Wang XG, Xu K, Gao Q, et al. CDetection: CRISPR-Cas12b-based DNA detection with sub-attomolar sensitivity and single-base specificity. Genome Biol. 2019, 20, 132. DOI:10.1186/s13059-019-1742-z [Google Scholar]
- Cao L, Wang Z, Lei C, Nie Z. Engineered CRISPR/Cas Ribonucleoproteins for Enhanced Biosensing and Bioimaging. Anal. Chem. 2025, 97, 5866–5879. DOI:10.1021/acs.analchem.4c06789 [Google Scholar]
- Guan X, Yang R, Zhang J, Moon J, Hou C, Guo C, et al. Programmable Multiplexed Nucleic Acid Detection by Harnessing Specificity Defect of CRISPR-Cas12a. Adv. Sci. 2025, 12, 2411021. DOI:10.1002/advs.202411021 [Google Scholar]
- Molina Vargas Adrian M, Sinha S, Osborn R, Arantes Pablo R, Patel A, Dewhurst S, et al. New design strategies for ultra-specific CRISPR-Cas13a-based RNA detection with single-nucleotide mismatch sensitivity. Nucleic Acids Res. 2023, 52, 921–939. DOI:10.1093/nar/gkad1132 [Google Scholar]
- Ye X, Wu H, Liu J, Xiang J, Feng Y, Liu Q. One-pot diagnostic methods based on CRISPR/Cas and Argonaute nucleases: Strategies and perspectives. Trends Biotechnol. 2024, 42, 1410–1426. DOI:10.1016/j.tibtech.2024.06.009 [Google Scholar]
- Ding X, Yin K, Li Z, Lalla RV, Ballesteros E, Sfeir MM, et al. Ultrasensitive and visual detection of SARS-CoV-2 using all-in-one dual CRISPR-Cas12a assay. Nat. Commun. 2020, 11, 4711. DOI:10.1038/s41467-020-18575-6 [Google Scholar]