Bacteriophages in Human Gastrointestinal Health Applications
Received: 30 October 2025 Revised: 29 December 2025 Accepted: 04 February 2026 Published: 24 February 2026
© 2026 The authors. This is an open access article under the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/).
1. Introduction
Bacteriophages (also referred to as phages) are viruses that infect bacteria. It is hypothesized that most bacterial species are parasitized by specific bacteriophages [1,2]. Though phages are not considered living entities, they are the most numerous biological structures on Earth [3,4]. They populate every habitat on the planet where their respective bacteria can live: soil, water, plants, and the bodies of animals [5]. All animals are naturally and continuously exposed to phages through interactions with their environment. Phages are normal constituents of the mammalian gastrointestinal tract, where they create tightly knit communities with their corresponding bacterial hosts [3,6].
The discovery of phages in the early 1900s by Twort and d’Herelle and their application in treating bacterial disease has been covered in multiple reviews [2,7,8,9,10]. Despite early promise as an antimicrobial treatment, the therapeutic application of phages lost momentum after the discovery of antibiotics [8,11]. Subsequently, only a few institutions in France, Poland, and the former Soviet Union (currently Russia and Georgia) continued researching their development, and their records were not widely accessible at the time. However, the recent rise of multidrug-resistant bacterial strains has rekindled interest in the therapeutic potential of phages.
This renewed attention has led to extensive efforts to uncover and re-evaluate existing data on phage therapies. Several reviews have been compiled from English-language sources, online information, articles in the native languages of the aforementioned countries, and personal communication with investigators and clinical personnel [12,13,14,15,16]. Most of these reviews conclude that very few historical phage studies were conducted according to the current gold standards for clinical research, i.e., randomized, blinded, and placebo-controlled. Most previous phage applications are characterized as prophylactic or remedial “standard of care” rather than controlled trials [7]. Nevertheless, the wealth of accumulated knowledge—including semi-scientific reports, medical case studies, and anecdotal evidence—provides a valuable foundation for contemporary research.
Knowledge of bacteriophages is expanding rapidly, partially due to new sequencing techniques and methods of metagenomic analysis [17,18]. A fundamental difference among phages is whether their genetic content consists of DNA or RNA. This distinction shapes their replication mechanisms, evolution, and interaction with their hosts. DNA phages are by far the most prevalent type and represent most sequenced phage genomes. They consist of double-stranded DNA housed in a complex protein coat. In contrast, RNA phages contain RNA as their genetic material. The first known RNA phage, f2, which infects Escherichia coli, was described several decades after the discovery of DNA phages [19]. Phage RNA can be either single-stranded (ss) or double-stranded (ds). Single-stranded RNA genomes can exist in two orientations: negative sense (−ve) and positive sense (+ve). Positive sense phages have their genetic sequence oriented in the same direction as the host cell’s mRNA, allowing them to be recognized by the host cell’s translational machinery, resulting in rapid viral replication. In contrast, negative sense ssRNA phages must carry their own RNA-dependent RNA polymerase (RdRp) to transcribe the negative-sense genome into multiple strands of positive-sense RNA before translation can occur [20]. The host range of RNA phages is much narrower than that of DNA phages and originally was thought to be limited to members of the Pseudomonadota (formerly Proteobacteria) phylum but was later expanded to include Streptomyces (Actinomycetota) and potentially other bacterial groups [21].
Bacteriophage infection begins with a highly specific binding event between the phage and its bacterial host, which determines both the host range and the efficiency of subsequent infection. This process, commonly termed adsorption, is mediated by receptor-binding proteins (RBPs) located on the phage surface-typically on tail fibers, tail spikes, or baseplate structures, which recognize and bind to distinct surface receptors on the target bacterium, such as outer membrane proteins, lipopolysaccharides, teichoic acids, capsules, pili, or flagella [22,23,24]. When an RBP encounters a compatible receptor, a reversible binding can occur, allowing the phage to remain near the cell surface while searching for the productive receptor site. This step increases the probability of successful host recognition without irreversible commitment [25]. Once the correct receptor is encountered, high-affinity interactions between the phage RBP and the bacterial receptor stabilize the contact. These interactions often involve conformational changes in the phage tail structure that anchor the virion to the cell surface and trigger downstream infection processes. For example, in many-tailed phages (such as T4), the tail fibers bind to bacterial outer membrane components, leading to irreversible adsorption and structural rearrangements that prepare the phage for genome delivery [24]. The molecular specificity of these interactions underlie a phage’s host range; subtle differences in receptor structure or RBP binding domains can prevent adsorption and thereby confer phage resistance to specific bacterial strains. In some cases, binding may involve a two-step mechanism, a primary reversible interaction followed by irreversible binding to a secondary receptor, ensuring both specificity and the correct orientation for genome ejection [25]. At the structural level, high-resolution analyses have revealed how RBP architecture is tuned to recognize specific host receptors, providing insight into how phage tail complexes can trigger conformational transformations upon binding that ultimately facilitate DNA or RNA injection into the host cytoplasm [26].
Some phages can infect multiple bacterial species, with the multiple host associations stemming from the structural adaptability of tail fibers/receptor-binding proteins, enabling recognition of different receptors across species [27]. Such phages may influence bacterial ecology and horizontal gene transfer by packaging and moving genetic material between hosts [28]. Many broad-host-range phages have mosaic genomes created by recombination, enabling them to acquire genes that expand receptor specificity or overcome host resistance [29]. Metagenomic analyses show that many phage genomes are associated with multiple bacterial species in natural environments, including the human gut [27].
In parallel with gaining general knowledge of phages and their interactions with the environment, new studies have been conducted to investigate therapeutic phage applications in vitro [28,30,31,32] and in vivo in animal models, specifically murine [33,34,35,36,37,38], and in human subjects [39,40,41,42]. Since bacteria can populate or invade nearly every part of the human body, the scope of the current research is very broad. Major areas of focus include skin wounds, burns, infections, and ulcers [43,44,45]; respiratory, urinary, and digestive system diseases [41,46,47,48,49]; and septicemia [50]. The volume of reports on successful real-world cases of personalized bacteriophage therapy (BT) is growing and has instigated the most recent retrospective observational analysis of the first 100 consecutive bacteriophage therapy cases of difficult-to-treat infections, enabled by a Belgian consortium [51]. As stated by the authors, the inclusion of all BT cases submitted to the Belgum consortium from 2008–2022, rather than focusing only on successful or challenging ones, made it possible to evaluate the general efficacy of BT and examine functional relationships common across cases. The conclusion states that most cases resulted in clinical improvement, with or without eradication of the targeted bacteria, and that pathogen eradication was more likely with concomitant antibiotic use. Therefore, BT can be most effective when combined with antibiotics and can inform the design of future controlled clinical trials. In the above study, the most common indications were lower respiratory tract infections, skin infections, soft-tissue infections, and bone infections [51]. Another area of interest is the application of phages to gastrointestinal health, with a recent review of this topic primarily highlighting their use in livestock [52]. Here, we review recent information on the application of phages for gastrointestinal health in humans by summarizing the findings of clinical trials conducted according to current clinical research standards.
2. Bacteriophages as Pharmacologic Agents in Gastrointestinal Applications
Phages are a natural component of the healthy human gut, with the phageome comprising both virulent (lytic lifecycle) and temperate (lysogenic lifecycle) phages, in which temperate phages are presumed to be more abundant and stable in gut microbial communities [53,54,55,56,57,58]. Because of their natural presence in the gut and their host specificity, phages are being explored as both therapeutic agents for the treatment of bacterial GI infections as well as dietary supplements for prophylaxis and GI health maintenance. As antibacterial agents, phages are unique among pharmaceuticals because they replicate at the site of infection. However, only obligatory lytic phages have been approved for human applications as they do not avoid clearance by the host, while introducing temperate phages may present additional implications, like transfer of virulence or antibiotic resistance genes, or development of resistance in lysogens [59].
Traditionally and currently, two categories of bacteriophage preparation have been utilized: bacteriophage stocks and adapted bacteriophages [7]. Bacteriophage stocks are cocktails of multiple phages that exhibit broad-spectrum activity and target multiple pathogenic bacterial strains or species [59,60,61,62]. This approach does not require the identification of a pathogen. Adapted bacteriophages are individual phages that are active against specific bacterial strains. This preparation, though more precisely targeted, is also more time-consuming and labor-intensive as it requires infecting pathogenic bacteria to be isolated and identified, then tested against a large collection of previously isolated phages [7]. Both preparations have advantages and disadvantages, and the choice is based on the GI conditions that need to be addressed and the availability of a particular preparation. Phage cocktails are usually pre-made, ready-to-use stocks.
Several preparations and delivery mechanisms are currently used in phage therapies. For GI applications, phage preparations can be formulated as liquid solutions, solid tablets, or capsules. Another administration route is intrarectal, by enema or suppositories. However, this approach has not been used in clinical trials of phages for GI health; rather, it has been reported in case studies and personalized phage treatments [63,64,65]. Although not a strictly GI-focused condition, one case report mentioned intravenous and peritoneal application of phages to treat pancreatitis [66]. The most common mode of administration is through oral consumption [39,40,67,68,69]; however, when oral doses are administered, the phages can be destroyed by gastric acid. Since phages naturally colonize the human gut, it is obvious that they can escape acid destruction, at least to some extent. Many phages enter the gastrointestinal tract along with their bacterial hosts rather than as naked viral particles that overcome gastric acid barriers [70]. A large fraction of gut phages are temperate phages integrated into bacterial genomes as prophages; they travel through gastric acid within gut bacterial populations [71]. Virulent phages can also travel with their hosts before getting released by lysis. Also, some phage particles inherently withstand moderate acidic conditions better than expected [56]. However, when a therapeutic dose is administered, maximal survival of phages in acid transit is of utmost importance. Different techniques have been implemented to overcome this limitation. Traditionally, neutralizing stomach acid prior to administering phages has been considered an important factor in bacteriophage viability [2]. However, lowering gastric acidity could also permit the passage of pathogens and should be used with caution [38]. Thus, for treating acute bacterial infection (e.g., diarrhea), it may be better to increase the application dose rather than use acid-neutralizing agents [28]. The other promising method of protecting phages from stomach acidity is polymer encapsulation. The method was successfully tested in vitro and on farm animals [72,73,74].
Regarding dosing, there are two common units: multiplicity of infection (MOI) and plaque-forming units (PFU). In vitro research or dose calculations for the treatment of specific infections are based on bacterial counts and use MOI, which represents the ratio of bacteriophages to bacterial cells [65]. The dose of oral phage application is always given in plaque-forming units (PFU). The most widely used experimental method to quantify phage dose is the plaque assay, which directly measures infectious phage particles as PFU/mL [75]. In brief, a phage sample is serially diluted in a buffer, reducing the concentration of phage particles by known factors. Each phage dilution is mixed with a susceptible bacterial host and then embedded in a soft agar overlay on a nutrient agar plate. The number of plaques is counted on plates with an optimal countable range. The PFU titer is calculated by correcting for the dilution factor and the volume plated [75]. If needed, this PFU concentration can be used to determine multiplicity of infection (MOI) or actual dose administered in some studies [76]. Not all phage studies use plate counts. Some methods estimate infectivity via endpoint dilution assays or optical density changes [77]. Once PFU/mL is known, the therapeutic dose can be calculated to deliver the total infectious units (e.g., 1 × 108 PFU per dose). This ensures consistent biological dosing across treatment arms [78]. Theoretical calculations of effective phage titers vs. bacterial concentrations are presented by Danis-Wlodarczyk et al. [79]. A generic titer of 108 PFU/mL was proposed as a potentially “inundative” (as defined by the author) density that could result in the effective in situ elimination of bacteria [79]. However, this dose was developed based on “theoretical calculations” and cannot serve as a valid titer recommendation for practical use in human applications. In addition, the authors described their application, targeted at extraintestinal bacterial infections, in which phages presumably cross the epithelial barrier and reach the site of infection via the bloodstream. The range of phage titers used for the treatment of GI infections or in clinical study protocols is somewhere between 1 × 103 PFU and 3 × 109 PFU per single dose, depending on the age of the patients, the severity of the condition, and the phage formulation [39,40,41,68,69]. Finally, phage administration can be either acute, lasting 2–4 days [39,40,41], or prolonged, lasting a few weeks, and given as a single dose per day or multiple doses. Prolonged oral administration of phages was used for the therapeutic treatment of multidrug-resistant pathogens (3 weeks) [64], and in clinical trials evaluating the safety and tolerability of phage consumption and effects of phages on inflammation (4 weeks) [67,68].
Pharmacokinetics and pharmacodynamics, two key aspects of therapeutic applications, are difficult to measure and interpret for phages. These aspects are typically evaluated by three components: absorption, distribution, and elimination, and were recently reviewed by Międzybrodzki et al. for phage-based interventions [65]. That review elaborated on the challenges of pharmacokinetic evaluation of phages and the most important obstacles associated with phage therapy. The authors highlighted that intravenous and topical application of phages are relatively unproblematic. However, for oral application, there are a number of factors to consider, including phage pH sensitivity, phage penetration through epithelial barriers, inactivation of phages by body fluids, their immunogenicity, interaction with antibiotics, etc. Moreover, the components (absorption, distribution, and elimination) primarily apply to oral administration for extra-intestinal infections, where phages are absorbed into the bloodstream and travel to the site of infection. Phage pharmacokinetics for direct intra-intestinal applications requires further exploration [65].
The pharmacodynamics of GI phage therapy can be estimated in several ways. First, the number of excreted phages in stool can be compared to the administered dose, although the interpretation of data from this approach is speculative. For example, lower excretion may indicate that phages are destroyed by stomach acid, although it could also indicate temporary colonization and prolonged gut transit. There is evidence that many lytic phages can adhere to the mucus layer lining the intestinal epithelium and remain there until they contact susceptible bacteria [80,81]. If the excreted dose remains unchanged from the administered dose, it can indicate inactivity of the phages. However, it could also be interpreted as destruction of some phages in the stomach, followed by intestinal replication of surviving phages on target bacteria in the intestines [39]. Finally, a higher excreted dose unequivocally indicates phage replication on target bacteria. A second method to evaluate phage pharmacodynamics is to compare counts of target bacteria from pre-treatment and post-treatment stool samples. However, the interpretation of these data is also ambiguous since no change in bacterial counts can indicate either ineffective phages or increased replication of target bacteria that escaped phage attack. An additional challenge in gut phage therapy is the ability of phages to pass through epithelial cells into the system, which could be a sink that reduces the active dose at the target location [82,83]. Therefore, many clinical studies rely on measuring potentially phage-mediated effects on defined outcomes following phage application [7].
3. Effects and Safety of Gastrointestinal Phage Applications in Human Clinical Trials
According to the FDA, registered controlled human clinical trials (randomized, double-blinded, and placebo-controlled) are the gold standard for evaluating the effects and safety of phage applications. Multiple trials are required to comprehensively evaluate the potential of phage therapy, and conclusions should not be made based on a single trial outcome [59,84]. However, human gastrointestinal phage therapy trials conducted in accordance with current clinical research standards are limited. This section summarizes the results of several of those studies in terms of gastrointestinal effects, safety, and tolerability.
Introducing specific lytic phages into the human gastrointestinal tract at doses above naturally occurring levels can have several effects. The first to consider, of course, is their impact on target bacteria. However, this direct effect can also trigger a cascade of secondary or indirect effects. Therefore, another consideration is the effect of phages on non-target microbiota and possible modification or even disruption of the homeostatic microbial composition of the gut. Finally, there is also the potential for phages to influence the human host’s immune system and stimulate immunological reactions, either by interacting with human epithelial and immune cells or by triggering responses to bacterial metabolites produced in response to viral predation (Figure 1). These effects can be reflected in measurable changes in GI-related conditions but may also cause adverse events when used for treatment, prophylaxis, or maintenance of GI health. Several clinical studies evaluated the above-listed outcomes following oral phage administration (Table 1).
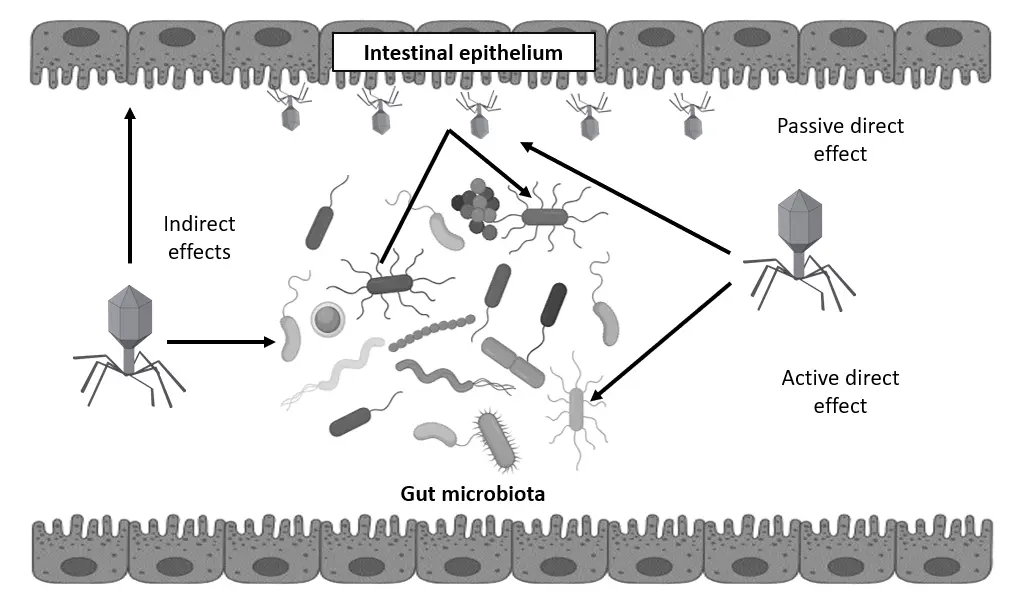
Figure 1. Possible mechanistic targets of bacteriophages in the human gastrointestinal tract: Active direct effect—attack on pathogens, Passive direct effect—prevention of pathogen adherence to intestinal mucosal epithelium, Indirect effects—modulation of gut microbiota and host immunity.
Table 1. Gastrointestinal phage applications in clinical trials.
|
References |
Study Design |
Study Population |
Dose of Phage Administration |
Major Study Outcomes |
|---|---|---|---|---|
|
Bruttin and Brussow [39], 2005 |
Single-center, randomized, double-blind, placebo-controlled trial |
Healthy adults (n = 15) |
1 × 103 PFU, 1 × 105 PFU |
Safety of phage administration. Effect on target bacteria. |
|
Sarker et al. [40], 2012 |
Single-center, randomized, double-blind, placebo-controlled trial |
Healthy adults (n = 15) |
3 × 107 PFU, 3 × 109 PFU |
Safety of phage administration. Effect on gut microbiota. |
|
McCallin et al. [69], 2013 |
Single-center, randomized, double-blind, placebo-controlled trial |
Healthy adults (n = 5), healthy children (n = 10) |
7 × 106 PFU |
Safety of phage administration. Effect on gut microbiota. |
|
Sarker et al. [41], 2016 |
Randomized, double-blind, placebo-controlled, parallel-group trial |
Male children with acute bacterial diarrhea (n = 120) |
3.6 × 108 PFU 1.4 × 109 PFU |
Safety of phage administration in children with acute diarrhea. Assessment of lytic phage activity. Effect on diarrhea parameters. |
|
Gindin et al. [68], 2018. Febvre et al. [67], 2019 |
Single-center, randomized, double-blind, placebo-controlled trial |
Healthy adults with self-reported gastrointestinal distress (n = 43) |
1 × 106 PFU |
Safety of phage administration. Effect on gut microbiota. Effect on markers of local and systemic inflammation. |
The first four studies presented in Table 1 were in continuity with each other, sharing similar methods [39,40,41,69]. Each subsequent study was based on the results of the previous study, extending the findings to a different study population and/or modified phage preparation. The following subsections describe the details of the above-listed studies, highlight the differences in their methods, and summarize findings related to particular outcomes.
3.1. Clinical Trial Designs and Outcomes
Bruttin and Brussow administered T4 phage that targeted unspecified strains of E. coli to 15 healthy adult Caucasian volunteers between 23–54 years old in a single-center, randomized, double-blind, and placebo-controlled trial [39]. It was a three-period crossover comparison of two oral T4 phage doses, with each subject receiving a higher dose (105 PFU/mL), a lower dose (103 PFU/mL), and a placebo in drinking water. After baseline data collection, each subject received a treatment dose for two consecutive days, followed by a five-day washout. This procedure was repeated for three consecutive weeks. The primary outcome was the safety of phage administration; the secondary outcome was its effect on target bacteria.
Sarker et al. conducted the next two clinical trials in Bangladesh [40,41]. The primary goal was to treat pediatric diarrhea in hospitalized children. The main objective of the first study was to create a safe and effective anti-diarrheal phage cocktail and to test it on healthy adult volunteers [40]. The genomic diversity of 99 T4-like coliphages was investigated by sequencing an equimolar mixture and screening them against different databases for horizontal gene transfer and undesired genes. The resulting cocktail consisted of nine independent isolates of T4-like E. coli phages, which were tested on 15 healthy adults in a trial following the design of the previous Bruttin and Brussow study [39]. This single-center, randomized, double-blind, placebo-controlled trial consisted of a three-period crossover design comparing two doses of the oral T4-like phage cocktail. The timing was similar to the previous study (two-day intervention followed by er to test for possible adverse events before applying the cocktail to children. Each subject received a higher phage dose (3 × 109 PFU/mL), a lower phage dose (3 × 107 PFU/mL), and a placebo in drinking water. The primary outcome was the safety of phage administration; the secondary outcome was its effect on the gut microbiota.
The follow-up study by McCallin et al. in Bangladesh investigated the composition of a phage cocktail (Microgen, Moscow, Russia) targeting Escherichia coli/Proteus infections [69]. Two methods of characterization were used: electron microscopy and metagenome analysis. Electron microscopy identified six phage types, with numerically T7-like phages dominating over T4-like phages. A metagenomic approach using taxonomic classification, reference mapping, and de novo assembly identified 18 distinct phage types. The complexity of the phage cocktail, as stated by the authors, raises questions about its safety, as it is difficult to maintain comparable quality and reproducibility in complex cocktail production compared with simpler formulations. The product was evaluated for safety by running phage sequences against a database of undesired genes, where no matches of concern were identified. Then, a small safety trial was set up with healthy human volunteers. The study design mimicked the preceding study by Sarker et al. [40], except for the study population and the phage doses used. Fifteen healthy subjects (five adults, five children aged 5–10, and five children under the age of 5) received a high dose of approximately 7 × 106 PFU/mL, a low dose (ten times lower), and a placebo. For the human trial, the primary outcome was the safety of phage administration; the secondary outcome was its effect on gut microbiota.
A second Sarker et al. clinical study took place in a hospital setting in Bangladesh and was based on the results of the previous two trials [41]. It utilized a T4-like coliphage cocktail [40] and a commercial coliphage product (Microgen ColiProteus, Moscow, Russia) [69] to treat acute bacterial diarrhea in children. In a randomized, placebo-controlled, parallel-group clinical trial, 120 male children were enrolled and allocated into 3 groups: 40 received orally 1.4 × 109 PFU/mL of the Microgen ColiProteus cocktail, 39 children received 3.6 × 108 PFU/mL of the T4-like coliphage cocktail, and 41 received a placebo with standard treatment for diarrhea patients. All treatments were given for 4 consecutive days. The primary outcome was the safety of oral coliphage in children infected with E. coli, the secondary outcome was the titration of fecal coliphage and E. coli pathogen to assess in vivo lytic phage activity, and the tertiary outcome was the impact of oral phage administration on quantitative diarrhea parameters.
The “Bacteriophage for Gastrointestinal Health (PHAGE)” clinical trial was the most comprehensive study of the oral application of bacteriophages in adults, and the results were presented in two publications. Gindin et al. evaluated the effects of bacteriophage intake on gastrointestinal health and the overall safety and tolerability of the treatment [68]. Febvre et al. assessed the effects of bacteriophage intake on inflammation and gut microbiota [67]. The PHAGE study was a randomized, double-blind, placebo-controlled crossover intervention. Forty-three healthy adults with self-reported gastrointestinal distress were enrolled in the study. Of these participants, 36 completed at least one arm of the trial, and 32 completed the entire study. Enrolled participants were randomly assigned to either the placebo or treatment starting groups and were asked to consume one 15-mg capsule daily for 28 days. This intervention was followed by a 2-week washout period prior to starting an additional 28-day intervention with the opposite treatment. The treatment was an encapsulated cocktail of four supplemental bacteriophage strains (LH01-Myoviridae, LL5-Siphoviridae, T4D-Myoviridae, and LL12-Myoviridae) at a titer of 106 PFU per dose included in the PreforPro commercial preparation (Deerland Enzymes, Kennesaw, GA, USA). These phages are known to infect a range of Escherichia coli strains, including E. coli K12, 16 enterotoxigenic E. coli strains, and 2 enterohemorrhagic strains. The primary outcome was the safety of phage administration; the secondary outcomes were its effects on gut microbiota and markers of local and systemic inflammation.
3.2. Safety of Phage Applications and the Effects on Gastrointestinal and Metabolic Health
The safety of the phage intervention was the major outcome in the above-described clinical trials. The Bruttin and Brussow study reported only five mild adverse events in the trial, four of which were related to GI distress (stomach pain, nausea, increased peristalsis) [39]. Adverse events were not reported more often with the higher phage dose than with the lower phage dose or placebo. All treatments were well tolerated. Serum levels of alanine aminotransferase and aspartate aminotransferase were chosen to evaluate phage application on liver toxicity. No significant increases in the levels of these two liver enzymes were observed after the intervention. In addition, neither T4 phage nor T4-specific antibodies were observed in the serum of the subjects at the end of the study. The authors concluded that phage treatments at both doses were safe and well-tolerated in the test population.
In the Sarker et al. study with healthy adult subjects, some shifts in white blood cell proportions and levels of aminotransferase were recorded, but there was no consistent pattern suggestive of an adverse effect of phage therapy. The authors concluded that no adverse events were observed by self-report, clinical examination, or laboratory tests for liver, kidney, and hematology function [40]. McCallin et al. assessed the safety of phage applications using the same parameters as Sarker et al. [69]: self-reports of adverse events, clinical examination, and laboratory tests of liver, kidney, and hematologic function. As to the authors’ conclusions, a few recorded adverse events were not associated with oral phage exposure. Also, no phages were detected in the blood, and no phage antibodies were found in the serum. In the follow-up study, which featured treatment of children with acute diarrhea, no adverse events attributable to oral phage application were observed [41]. However, it was noted that the release of endotoxin could possibly place a limit on the phage treatment of Gram-negative infections [7]. As discussed by the authors of the Sarker et al. trial, in acute cases of E. coli diarrhea, the barrier function of the gut mucosa in the small intestine is compromised: inflammation of the tissue affects the tight junctions between epithelial cells, creating a passageway for paracellular transit of substances. When the pathogen is lysed by phages, in vivo released endotoxins and enterotoxins can potentially cross into the circulation. However, no systemic reaction to endotoxin was observed in that trial. While the phage application in children with acute diarrhea was demonstrated to be both safe and well tolerated, it failed to improve gastrointestinal symptoms. Neither the T4-like nor the Russian coliphage cocktail showed a clinical benefit over standard care in terms of stool output, stool frequency, or rehydration [41].
The “PHAGE” study reported no adverse events during the trial [68]. Self-assessment of gastrointestinal symptoms suggested improvements in most parameters measured. In addition, there were bidirectional crossover effects, suggesting that participants perceived relief of gastrointestinal distress just from participating in the study. The majority of individuals maintained or reduced baseline levels of gastrointestinal distress during treatment, suggesting that it was tolerable and did not exacerbate symptoms. Comprehensive metabolic panels were used to monitor blood chemistry and liver function and to assess the safety of bacteriophage consumption. Although there were statistically significant changes in a few parameters, overall, the analyte averages were within clinical reference ranges for all measurements. The authors concluded that consumption of therapeutic doses of a mixture of four bacteriophages was both safe and tolerable in healthy individuals with mild to moderate gastrointestinal distress.
In summary, the results of available human clinical trials indicate that oral phage treatments were safe and well-tolerated in the studied doses and timeframes. Also, the results of prolonged consumption of phages (28 days) suggest that bacteriophage supplementation did not exacerbate symptoms in healthy individuals with mild to moderate gastrointestinal distress [68]. However, the acute therapeutic application of phages to treat diarrhea did not achieve a clinical advantage over traditional methods. As stated by the authors, these observations should be interpreted with caution because the trial was not powered to detect a treatment effect, but only to assess safety issues and gain mechanistic insight into phage therapy [41].
3.3. Effects of Phages on Target Bacteria
Lytic bacteriophages infect and destroy representatives of specific strains of bacteria. In the process, they multiply and, after the lysis of the host cell, are ready to infect additional susceptible bacterial cells. The cycles of infection, multiplication, and lysis are supposed to be continued until susceptible cells are present within the reach of the phages. After predation cycles stop, the remaining phages are evacuated from the gut with bowel movements. Thus, the direct effect of phages is the destruction of target strains. However, in the highly interconnected microbial communities of the GI tract, the destruction of some of its members almost inevitably impacts the rest of the microbiome to some degree. Therefore, in GI phage applications, both target bacteria and the overall microbiota composition can be affected, and both effects should be evaluated in clinical trials.
The effect of phages on target bacteria is most commonly measured by comparing fecal counts of bacteria in pre- and post-treatment stool samples. The results of different studies seem to be somewhat conflicting, reporting either a reduction of bacterial counts or no effect on bacterial counts after phage application. In all of the above-described clinical trials, the target bacteria were different strains of E. coli.
Bruttin et al. reported no change in fecal E. coli counts after a 2-day intervention with phage T4 at both lower and higher doses [39]. The same result was observed in the Sarker et al. study with healthy adults, as expected, since no E. coli was detected in the initial stool samples [40]. In the study of children with diarrhea, viable E. coli colony counts represented only 5% of all fecal bacteria in diarrhea patients and were at most 10-fold higher as compared to non-E. coli diarrhea patients. Children with non-E. coli diarrhea and healthy controls mainly showed less than 1% Escherichia abundance. No significant decrease in E. coli (ETEC) titers was observed after phage treatment. According to the authors’ interpretation, the pathogenic E. coli titer, even in confirmed cases of E. coli diarrhea, might not be high enough to sustain phage replication in vivo [41].
Why might target bacteria be unaffected when exposed to their respective lytic phages? Although the “kill the winner” model was not based on intestinal microbial ecosystems, we can speculate that it is potentially applicable to the gut microbiota. The model implies that phages and other predators will mainly attack targets that are dominant in an ecosystem, so only when the overgrowth of target bacteria occurs do the phages reduce their number, balancing the ecosystem. Another potentially applicable model proposes that the “spatial organization” of bacteria may allow some members of a target host to escape predation by phages [85]. It suggests that the spatial separation of bacteria prevents phages from reaching and attacking all available hosts. Both models denote that the composition and structure of the host microbiome can possibly determine the response to treatment. The idea of response vs. non-response to the GI treatments being rooted in the host gut environment is gaining popularity in nutritional supplementation studies [86,87] and is worth further investigation in phage therapy. However, in phage formulations, the dose and duration of treatment are major considerations. It is important to note that all these studies measuring effects on host bacterial populations used phages for only 2–4 days. Acute and prolonged applications have quite different pharmacodynamics, and acute application may not be effective enough to significantly reduce the number of target bacteria, particularly if there is no pronounced overgrowth of the bacterial host.
In contrast, Febvre et al. observed a significant reduction in the relative abundance of E. coli in the “PHAGE” study following 4 weeks of phage consumption [67]. However, this reduction was observed only in a subpopulation of the study that had detectable baseline levels of E. coli. Only 21 participants (out of 43) had detectable levels of E. coli before the treatment period began. Baseline levels varied significantly among participants, ranging from 0.01–3.2% of total operational taxonomic units (OTUs). The response rate was ~71%, with 15 of the 21 participants who had detectable E. coli levels prior to the treatment period showing reduced or undetectable levels after the treatment. Overall, the number of E. coli reads was reduced by ~40% after treatment and only by 14% after the placebo, although there was high variability between participants. Though the reason for the E. coli reduction after the placebo was not discussed by the authors, we can speculate that in a randomized crossover design, some carry-over effect is possible (in participants who received the treatment in the first period of the study, its effects may last through the washout period and affect the placebo phase). Finally, it should be noted that these data were based on 16S rRNA sequencing reads, which cannot reliably distinguish E. coli from Shigella and other similar taxa and only account for relative, not absolute, changes in a bacterial population.
3.4. Effects of Phages on Gastrointestinal Microbiota
The effect of phage consumption on microbiota was evaluated in several clinical trials. None of the studies reported disruption or global modification of microbiota. However, some decreases or increases in the counts of certain taxa may occur after phage application.
In Sarker et al.’s study, some subjects showed changes in the relative abundance of two phyla, Actinomycetota and Bacteroidota, over the observation period; the authors did not discuss the possible effects of this change [40]. There were no consistent trends and no consistent differences between placebo and phage applications. The relative abundance before and after the intervention was evaluated for the major fecal bacterial families using 16S rRNA analysis. Data points from phage recipients overlapped those from placebo recipients. When the fecal microbiota data were compared for overall bacterial diversity and richness at the strain level (defined as 99% identity), phage-treated subjects displayed the same range as placebo-treated. The authors concluded that there were no impacts of oral phage consumption on the fecal microbiota composition.
McCallin et al. based their microbiota analysis on bacterial 16S rRNA sequencing of a very limited number of samples, only four adults in the study [69]. When feces were sampled three times over a one-week observation period, no consistent change in microbiota composition was seen in adult volunteers receiving the high phage dose.
The “PHAGE” study also presents the results of the 16S rRNA sequencing of gut microbiota [67]. There were no significant differences in bacterial taxa between treatment groups and time points. Richness estimates (CHAO1) and α-diversity, calculated as Shannon and Simpson indices, did not differ across groups, and PCoA of Bray–Curtis distances did not reveal significant clustering between treatment groups or time points. However, the counts of some taxa were affected by the treatment. One read that mapped to Bifidobacterium bifidum increased significantly in relative abundance after phage consumption in the responder subpopulation (defined as participants with detectable baseline E. coli who showed reductions after treatment), reaching ~5.5-fold above baseline. In the total study population, Clostridium perfringens counts decreased, and two OTUs representing Eubacterium were relatively increased during treatment compared with placebo. The authors concluded that phage consumption did not disrupt the gut microbiota but did elicit minor changes that may be considered beneficial.
3.5. Effects on the Immune System of the Human Host
Immunological interactions between mammalian hosts and bacteriophages are very complex and multifaceted. These interactions were mostly studied in vitro or in vivo using animal models, and multiple reviews have covered different aspects of these interactions [34,88,89,90]. To date, there is somewhat limited information on the effects of lytic phage consumption on the human immune system in registered clinical trials. Of the five clinical studies covered in this review, only one briefly addressed immunity in its results report, noting that no T4-specific antibodies were detected in the subjects’ serum at the end of the intervention [39]. One of the points of emphasis here is that the neutralization or inactivation of bacteriophages by the immune system could be a complicating factor in phage therapy [3,52] and should be considered in terms of phage dosage and treatment duration. The effects of phage application on the human immune system are beyond the scope of this article.
4. The Potential of Supplemental Phage Applications in Gastrointestinal Health
Supplemental bacteriophage applications hold significant promise for enhancing gastrointestinal health. The U.S. Food and Drug Administration lists many bacteriophages as Generally Recognized as Safe (GRAS) for human consumption because they are abundant in nature, naturally occur in the human gastrointestinal tract, and are inadvertently consumed daily by humans [68]. Unlike broad-spectrum antibiotics, which indiscriminately eliminate both pathogenic and beneficial bacteria, current research suggests that bacteriophages offer a precision-based approach that preserves the integrity of the gut ecosystem. As discussed, this selective targeting has been associated with beneficial shifts in microbial populations, including increases in butyrate-producing bacteria such as Eubacterium, which are known to support gut barrier function and reduce inflammation. Moreover, supplemental phage intake has shown potential in lowering markers of intestinal and systemic inflammation, suggesting a role in managing gastrointestinal distress and promoting overall gut health when used alone or in combination with probiotics. Grubb et al. conducted a “PHAGE 2” study using a combination of bacteriophages and a probiotic in one capsule [91]. The phage component was the same E. coli-targeting cocktail that was used in the above-described “PHAGE” study [67,68]. The probiotic component in the study was Bifidobacterium animalis subspecies lactis strain BL04 (B. lactis BL04). Probiotics are defined as live microorganisms that, when administered in adequate amounts, confer benefits to the host [92]. Bifidobacterium and Lactobacillus are the most common and widely used traditional probiotics.
PHAGE 2 was a randomized, parallel-arm, double-blind, placebo-controlled clinical intervention trial designed to test whether combining supplemental bacteriophages with a probiotic would provide additional benefits for GI health and the gut microbiota compared with consuming the probiotic alone. Participants, healthy adults some of whom had mild to moderate GI distress, were randomly assigned to one of 3 treatment groups: (1) 1 × 109 Colony Forming Units (CFU) B. lactis BL04; (2) 1 × 109 CFU B. lactis BL04 + 1 × 106 PFU of the phage cocktail; (3) placebo. One 15 mg capsule per day was consumed for four weeks. The primary outcome was GI health assessment, and the secondary outcome was the effect on gut microbiota. Participants who consumed B. lactis BL04 + phages showed improvements in digestive symptoms related to GI inflammation and colon pain and had the highest percentage of individuals reporting overall reduced symptom severity. The combination of B. lactis BL04 + phages did not significantly impact gut microbiota; no shifts were recorded in either alpha- or beta-diversity parameters or large phyla-level changes in taxa. However, there were small but statistically significant changes in the number of lower-level taxa after four weeks of B. lactis BL04 + phage consumption, including an increase in Lactobacillus and a decrease in Citrobacter and Desulfovibrio. In summary, the combination of a phage cocktail and a probiotic positively affected some GI symptoms without disrupting the microbiota.
Phage-containing dietary supplements have been designed for prophylaxis and maintenance of GI health [7]. A recent study investigated the potential of prophylactic bacteriophage administration to reduce the burden of Salmonella in vivo in mice colonized with a synthetic microbiota [93]. The bacteriophage preparation, FOP (foodborne outbreak pill), contained lytic phages targeting Salmonella (SalmoFresh phage cocktail), Shiga toxin-producing Escherichia coli (STEC), and Listeria monocytogenes. The authors reported that the administration of bacteriophages was safe and efficient in lowering the Salmonella burden and in reducing perturbations of the microbiota caused by Salmonella infection, suggesting the use of phages as a promising prophylactic intervention to reduce the spread of foodborne epidemics.
Probiotics introduce beneficial bacteria that interfere with pathogenic bacteria’s ability to colonize the GI tract by competing for attachment sites and nutritional resources. Phages, besides targeting specific pathogenic bacteria, could also have a preventive function due to their ability to adhere to the mucus layer. The effect is described as “Bacteriophage Adherence to Mucin” (BAM): it has been demonstrated that T4 phage proteins exposed on the capsid can bind mucin glycoproteins and protect mucus-producing tissue culture cells in vitro [80,81]. Thus, phages can aid GI health by protecting enterocytes from adherence and attacks of pathogenic bacteria.
Another advantage of phages is their ability to mutate and evolve to counter the development of phage-resistant bacterial strains [15,94]. The development of bacterial resistance to phages is a major consideration in long-term phage supplementation strategies [95]. Bacteria evolve resistance to phages through several well-characterized mechanisms. These mechanisms arise from long-term antagonistic co-evolution between bacterial hosts and phage predators and can be grouped into surface, genetic, and intracellular defense strategies. The first one is the modification or loss of phage receptors (surface exclusion), which prevents phage adsorption and thereby blocks infection at the first step [95]. The second way is bacterial adaptive immunity via CRISPR-Cas systems. During a phage attack, bacterial cells can incorporate short fragments of phage DNA (spacers) into their CRISPR arrays. These spacers are transcribed and processed into guide RNAs that direct Cas nucleases to recognize and cleave matching sequences in invading phage genomes upon subsequent encounters. This mechanism provides sequence-specific immunity, effectively destroying phage DNA before it can replicate [96]. Restriction-modification (R-M) systems are widespread bacterial defenses that distinguish self from non-self DNA. A host methyltransferase modifies specific nucleotide sequences in the bacterial genome. An associated restriction endonuclease then cleaves foreign DNA (such as unmodified phage DNA) at those same sequences. Phage DNA lacking the host’s methylation pattern is cut and degraded, preventing replication. Bacteria can combine R-M and CRISPR mechanisms to enhance defense [96]. Some bacteria possess abortive infection (Abi) systems that sacrifice infected cells to block phage proliferation. Abi systems trigger cellular suicide or aggressive responses when phage replication begins, terminating the infection at the cost of the individual cell’s life. Toxin-antitoxin modules and other less-specific intracellular systems also halt phage replication, serving as a last line of defense [97]. Despite all these elaborate mechanisms of bacterial defense, phages are still abundant and successful in all bacterial environments due to their ability to co-evolve in parallel with their respective hosts [98]. In addition, therapeutic application of phages can employ mechanisms beyond the natural phage adaptations to overcome host defenses. A recent review by Bleriot et al. summarized therapeutic approaches that are being evaluated to overcome the bacterial defense systems, including circumvention of phage receptor mutations, modification of prophages, targeting of CRISPR-Cas systems and the biofilm matrix, engineering of safer and more efficacious phages, and inhibition of the anti-persister strategies used by bacteria [99]. Therefore, even prolonged supplementary application of phages does not impose the same problem in terms of bacterial resistance as using drugs and antibiotics. Thus, as antibiotic resistance continues to rise, bacteriophages may emerge as a compelling alternative, offering a safe, tolerable, and microbiome-friendly strategy for maintaining and restoring gastrointestinal homeostasis. For example, nosocomial intestinal infections and “traveler’s diarrhea” may be good targets for phage-based supplements. These conditions are commonly caused by members of the family Enterobacteriaceae [64], and gut colonization by multidrug-resistant Enterobacteriaceae has been observed in healthy travelers returning from endemic areas of antimicrobial resistance [100,101,102]. A phage cocktail designed to lyse selected species of Enterobacteriaceae could be consumed during international travel as a low-risk preventive measure against prospective pathogens.
5. Conclusions
In a healthy gut, there is a natural balance between bacteria and phages. Currently, bacteriophages are underutilized as therapeutic, prophylactic, and maintenance agents. Introduced at the right time and in the right doses, phages have the potential to treat and/or prevent GI infections and aid GI health. The dynamics, effects, and mechanisms of action of bacteriophages in the human gastrointestinal tract are prospects for further elucidation.
Author Contributions
N.B.W. wrote the review and created figures and T.L.W. edited and provided revisions.
Ethics Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Funding
N.B.W. is supported by an assistantship funded by ADM Deerland Enzymes and Probiotics.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
-
Cowan MK. Microbiology: A Systems Approach, 6th ed.; Mc-Graw Hill: New York, NY, USA, 2020. [Google Scholar]
-
Keen EC. Felix d’Herelle and our microbial future. Future Microbiol. 2012, 7, 1337–1339. DOI:10.2217/fmb.12.115 [Google Scholar]
-
Dabrowska K, Switala-Jelen K, Opolski A, Weber-Dabrowska B, Gorski A. Bacteriophage penetration in vertebrates. J. Appl. Microbiol. 2005, 98, 7–13. DOI:10.1111/j.1365-2672.2004.02422.x [Google Scholar]
-
Willey JM, Prescott LM, Sandman KM, Wood DH. Prescott’s Microbiology, 12th ed.; McGraw-Hill: New York, NY, USA, 2023. [Google Scholar]
-
Comeau AM, Hatfull GF, Krisch HM, Lindell D, Mann NH, Prangishvili D. Exploring the prokaryotic virosphere. Res. Microbiol. 2008, 159, 306–313. DOI:10.1016/j.resmic.2008.05.001 [Google Scholar]
-
Gorski A, Dabrowska K, Switala-Jelen K, Nowaczyk M, Weber-Dabrowska B, Boratynski J, et al. New insights into the possible role of bacteriophages in host defense and disease. Med. Immunol. 2003, 2, 2. DOI:10.1186/1476-9433-2-2 [Google Scholar]
-
Abedon ST, Kuhl SJ, Blasdel BG, Kutter EM. Phage treatment of human infections. Bacteriophage 2011, 1, 66–85. DOI:10.4161/bact.1.2.15845 [Google Scholar]
-
Golkar Z, Bagasra O, Pace DG. Bacteriophage therapy: A potential solution for the antibiotic resistance crisis. J. Infect. Dev. Ctries. 2014, 8, 129–136. DOI:10.3855/jidc.3573 [Google Scholar]
-
Keen EC. A century of phage research: Bacteriophages and the shaping of modern biology. Bioessays 2015, 37, 6–9. DOI:10.1002/bies.201400152 [Google Scholar]
-
Summers WC. In the beginning. Bacteriophage 2011, 1, 50–51. DOI:10.4161/bact.1.1.14070 [Google Scholar]
-
Wittebole X, De Roock S, Opal SM. A historical overview of bacteriophage therapy as an alternative to antibiotics for the treatment of bacterial pathogens. Virulence 2014, 5, 226–235. DOI:10.4161/viru.25991 [Google Scholar]
-
Chanishvili N. Phage therapy—History from Twort and d’Herelle through Soviet experience to current approaches. Adv. Virus Res. 2012, 83, 3–40. DOI:10.1016/B978-0-12-394438-2.00001-3 [Google Scholar]
-
Gorski A, Miedzybrodzki R, Borysowski J, Weber-Dabrowska B, Lobocka M, Fortuna W, et al. Bacteriophage therapy for the treatment of infections. Curr. Opin. Investig. Drugs 2009, 10, 766–774. [Google Scholar]
-
Guo M, Li M, Zhang C, Zhang X, Wu Y. Dietary Administration of the Bacillus subtilis Enhances Immune Responses and Disease Resistance in Chickens. Front. Microbiol. 2020, 11, 1768. DOI:10.3389/fmicb.2020.01768 [Google Scholar]
-
Ho K. Bacteriophage therapy for bacterial infections. Rekindling a memory from the pre-antibiotics era. Perspect. Biol. Med. 2001, 44, 1–16. DOI:10.1353/pbm.2001.0006 [Google Scholar]
-
Miedzybrodzki R, Borysowski J, Weber-Dabrowska B, Fortuna W, Letkiewicz S, Szufnarowski K, et al. Clinical aspects of phage therapy. Adv. Virus Res. 2012, 83, 73–121. DOI:10.1016/B978-0-12-394438-2.00003-7 [Google Scholar]
-
Breitbart M, Hewson I, Felts B, Mahaffy JM, Nulton J, Salamon P, et al. Metagenomic analyses of an uncultured viral community from human feces. J. Bacteriol. 2003, 185, 6220–6223. DOI:10.1128/JB.185.20.6220-6223.2003 [Google Scholar]
-
Minot S, Sinha R, Chen J, Li H, Keilbaugh SA, Wu GD, et al. The human gut virome: Inter-individual variation and dynamic response to diet. Genome Res. 2011, 21, 1616–1625. DOI:10.1101/gr.122705.111 [Google Scholar]
-
Loeb T, Zinder ND. A bacteriophage containing RNA. Proc. Natl. Acad. Sci. USA 1961, 47, 282–289. DOI:10.1073/pnas.47.3.282 [Google Scholar]
-
Callanan J, Stockdale SR, Shkoporov A, Draper LA, Ross RP, Hill C. RNA Phage Biology in a Metagenomic Era. Viruses 2018, 10, 386. DOI:10.3390/v10070386 [Google Scholar]
-
Krishnamurthy SR, Janowski AB, Zhao G, Barouch D, Wang D. Hyperexpansion of RNA Bacteriophage Diversity. PLoS Biol. 2016, 14, e1002409. DOI:10.1371/journal.pbio.1002409 [Google Scholar]
-
Bertozzi Silva J, Storms Z, Sauvageau D. Host receptors for bacteriophage adsorption. FEMS Microbiol. Lett. 2016, 363, fnw002. DOI:10.1093/femsle/fnw002 [Google Scholar]
-
Degroux S, Effantin G, Linares R, Schoehn G, Breyton C. Deciphering Bacteriophage T5 Host Recognition Mechanism and Infection Trigger. J. Virol. 2023, 97, e0158422. DOI:10.1128/jvi.01584-22 [Google Scholar]
-
Kortright KE, Chan BK, Turner PE. High-throughput discovery of phage receptors using transposon insertion sequencing of bacteria. Proc. Natl. Acad. Sci. USA 2020, 117, 18670–18679. DOI:10.1073/pnas.2001888117 [Google Scholar]
-
Zeng X, Wang W, Zhu D, Liu M, Wang M, Jia R, et al. Two receptor-targeting mechanisms of lambda-like siphophage Gifsy-1 of Salmonella Typhimurium. PLoS Pathog. 2025, 21, e1013352. DOI:10.1371/journal.ppat.1013352 [Google Scholar]
-
Ge X, Wang J. Structural mechanism of bacteriophage lambda tail’s interaction with the bacterial receptor. Nat. Commun. 2024, 15, 4185. DOI:10.1038/s41467-024-48686-3 [Google Scholar]
-
Bignaud A, Conti DE, Thierry A, Serizay J, Labadie K, Poulain J, et al. Phages with a broad host range are common across ecosystems. Nat. Microbiol. 2025, 10, 2537–2549. DOI:10.1038/s41564-025-02108-2 [Google Scholar]
-
Göller PC, Elsener T, Lorgé D, Radulovic N, Bernardi V, Naumann A, et al. Multi-species host range of staphylococcal phages isolated from wastewater. Nat. Commun. 2021, 12, 6965. DOI:10.1038/s41467-021-27037-6 [Google Scholar]
-
Fong K, Wong CWY, Wang S, Delaquis P. How Broad Is Enough: The Host Range of Bacteriophages and Its Impact on the Agri-Food Sector. Ther. Appl. Res. 2021, 2, 83–91. DOI:10.1089/phage.2020.0036 [Google Scholar]
-
Meader E, Mayer MJ, Gasson MJ, Steverding D, Carding SR, Narbad A. Bacteriophage treatment significantly reduces viable Clostridium difficile and prevents toxin production in an in vitro model system. Anaerobe 2010, 16, 549–554. DOI:10.1016/j.anaerobe.2010.08.006 [Google Scholar]
-
Moye ZD, Woolston J, Abbeele PVD, Duysburgh C, Verstrepen L, Das CR, et al. A Bacteriophage Cocktail Eliminates Salmonella Typhimurium from the Human Colonic Microbiome while Preserving Cytokine Signaling and Preventing Attachment to and Invasion of Human Cells by Salmonella In Vitro. J. Food Prot. 2019, 82, 1336–1349. DOI:10.4315/0362-028X.JFP-18-587 [Google Scholar]
-
Yu L, Wang S, Guo Z, Liu H, Sun D, Yan G, et al. A guard-killer phage cocktail effectively lyses the host and inhibits the development of phage-resistant strains of Escherichia coli. Appl. Microbiol. Biotechnol. 2018, 102, 971–983. DOI:10.1007/s00253-017-8591-z [Google Scholar]
-
Cepko LCS, Garling EE, Dinsdale MJ, Scott WP, Bandy L, Nice T, et al. Myoviridae phage PDX kills enteroaggregative Escherichia coli without human microbiome dysbiosis. J. Med. Microbiol. 2020, 69, 309–323. DOI:10.1099/jmm.0.001162 [Google Scholar]
-
Federici S, Kredo-Russo S, Valdes-Mas R, Kviatcovsky D, Weinstock E, Matiuhin Y, et al. Targeted suppression of human IBD-associated gut microbiota commensals by phage consortia for treatment of intestinal inflammation. Cell 2022, 185, 2879–2898.e24. DOI:10.1016/j.cell.2022.07.003 [Google Scholar]
-
Gu J, Liu X, Li Y, Han W, Lei L, Yang Y, et al. A method for generation phage cocktail with great therapeutic potential. PLoS ONE 2012, 7, e31698. DOI:10.1371/journal.pone.0031698 [Google Scholar]
-
Hsu BB, Gibson TE, Yeliseyev V, Liu Q, Lyon L, Bry L, et al. Dynamic Modulation of the Gut Microbiota and Metabolome by Bacteriophages in a Mouse Model. Cell Host Microbe 2019, 25, 803–814.e5. DOI:10.1016/j.chom.2019.05.001 [Google Scholar]
-
Maura D, Morello E, du Merle L, Bomme P, Le Bouguenec C, Debarbieux L. Intestinal colonization by enteroaggregative Escherichia coli supports long-term bacteriophage replication in mice. Environ. Microbiol. 2012, 14, 1844–1854. DOI:10.1111/j.1462-2920.2011.02644.x [Google Scholar]
-
Vahedi A, Soltan Dallal MM, Douraghi M, Nikkhahi F, Rajabi Z, Yousefi M, et al. Isolation and identification of specific bacteriophage against enteropathogenic Escherichia coli (EPEC) and in vitro and in vivo characterization of bacteriophage. FEMS Microbiol. Lett. 2018, 365, fny136. DOI:10.1093/femsle/fny136 [Google Scholar]
-
Bruttin A, Brussow H. Human volunteers receiving Escherichia coli phage T4 orally: A safety test of phage therapy. Antimicrob. Agents Chemother. 2005, 49, 2874–2878. DOI:10.1128/AAC.49.7.2874-2878.2005 [Google Scholar]
-
Sarker SA, McCallin S, Barretto C, Berger B, Pittet AC, Sultana S, et al. Oral T4-like phage cocktail application to healthy adult volunteers from Bangladesh. Virology 2012, 434, 222–232. DOI:10.1016/j.virol.2012.09.002 [Google Scholar]
-
Sarker SA, Sultana S, Reuteler G, Moine D, Descombes P, Charton F, et al. Oral Phage Therapy of Acute Bacterial Diarrhea with Two Coliphage Preparations: A Randomized Trial in Children from Bangladesh. EBioMedicine 2016, 4, 124–137. DOI:10.1016/j.ebiom.2015.12.023 [Google Scholar]
-
Wright A, Hawkins CH, Anggard EE, Harper DR. A controlled clinical trial of a therapeutic bacteriophage preparation in chronic otitis due to antibiotic-resistant Pseudomonas aeruginosa; a preliminary report of efficacy. Clin. Otolaryngol. 2009, 34, 349–357. DOI:10.1111/j.1749-4486.2009.01973.x [Google Scholar]
-
Marza JA, Soothill JS, Boydell P, Collyns TA. Multiplication of therapeutically administered bacteriophages in Pseudomonas aeruginosa infected patients. Burns 2006, 32, 644–646. DOI:10.1016/j.burns.2006.02.012 [Google Scholar]
-
Merabishvili M, Pirnay JP, Verbeken G, Chanishvili N, Tediashvili M, Lashkhi N, et al. Quality-controlled small-scale production of a well-defined bacteriophage cocktail for use in human clinical trials. PLoS ONE 2009, 4, e4944. DOI:10.1371/journal.pone.0004944 [Google Scholar]
-
Rhoads DD, Wolcott RD, Kuskowski MA, Wolcott BM, Ward LS, Sulakvelidze A. Bacteriophage therapy of venous leg ulcers in humans: Results of a phase I safety trial. J. Wound Care 2009, 18, 237–243. DOI:10.12968/jowc.2009.18.6.42801 [Google Scholar]
-
Chegini Z, Khoshbayan A, Vesal S, Moradabadi A, Hashemi A, Shariati A. Bacteriophage therapy for inhibition of multi drug-resistant uropathogenic bacteria: A narrative review. Ann. Clin. Microbiol. Antimicrob. 2021, 20, 30. DOI:10.1186/s12941-021-00433-y [Google Scholar]
-
Jennes S, Merabishvili M, Soentjens P, Pang KW, Rose T, Keersebilck E, et al. Use of bacteriophages in the treatment of colistin-only-sensitive Pseudomonas aeruginosa septicaemia in a patient with acute kidney injury—A case report. Crit. Care 2017, 21, 129. DOI:10.1186/s13054-017-1709-y [Google Scholar]
-
Leitner L, Sybesma W, Chanishvili N, Goderdzishvili M, Chkhotua A, Ujmajuridze A, et al. Bacteriophages for treating urinary tract infections in patients undergoing transurethral resection of the prostate: A randomized, placebo-controlled, double-blind clinical trial. BMC Urol. 2017, 17, 90. DOI:10.1186/s12894-017-0283-6 [Google Scholar]
-
Malik S, Nehra K, Rana JS. Bacteriophage cocktail and phage antibiotic synergism as promising alternatives to conventional antibiotics for the control of multi-drug-resistant uropathogenic Escherichia coli. Virus Res. 2021, 302, 198496. DOI:10.1016/j.virusres.2021.198496 [Google Scholar]
-
Gorski A, Jonczyk-Matysiak E, Lusiak-Szelachowska M, Miedzybrodzki R, Weber-Dabrowska B, Borysowski J. The Potential of Phage Therapy in Sepsis. Front. Immunol. 2017, 8, 1783. DOI:10.3389/fimmu.2017.01783 [Google Scholar]
-
Pirnay J-P, Djebara S, Steurs G, Griselain J, Cochez C, De Soir S, et al. Personalized bacteriophage therapy outcomes for 100 consecutive cases: A multicentre, multinational, retrospective observational study. Nat. Microbiol. 2024, 9, 1434–1453. DOI:10.1038/s41564-024-01705-x [Google Scholar]
-
Mahmud MR, Tamanna SK, Akter S, Mazumder L, Akter S, Hasan MR, et al. Role of bacteriophages in shaping gut microbial community. Gut Microbes 2024, 16, 2390720. DOI:10.1080/19490976.2024.2390720 [Google Scholar]
-
Hu J, Ye H, Wang S, Wang J, Han D. Prophage Activation in the Intestine: Insights into Functions and Possible Applications. Front. Microbiol. 2021, 12, 785634. DOI:10.3389/fmicb.2021.785634 [Google Scholar]
-
Kim MS, Park EJ, Roh SW, Bae JW. Diversity and abundance of single-stranded DNA viruses in human feces. Appl. Environ. Microbiol. 2011, 77, 8062–8070. DOI:10.1128/AEM.06331-11 [Google Scholar]
-
Lourenco M, Chaffringeon L, Lamy-Besnier Q, Pedron T, Campagne P, Eberl C, et al. The Spatial Heterogeneity of the Gut Limits Predation and Fosters Coexistence of Bacteria and Bacteriophages. Cell Host Microbe 2020, 28, 390–401.e5. DOI:10.1016/j.chom.2020.06.002 [Google Scholar]
-
Manrique P, Bolduc B, Walk ST, van der Oost J, de Vos WM, Young MJ. Healthy human gut phageome. Proc. Natl. Acad. Sci. USA 2016, 113, 10400–10405. DOI:10.1073/pnas.1601060113 [Google Scholar]
-
Mathieu A, Dion M, Deng L, Tremblay D, Moncaut E, Shah SA, et al. Virulent coliphages in 1-year-old children fecal samples are fewer, but more infectious than temperate coliphages. Nat. Commun. 2020, 11, 378. DOI:10.1038/s41467-019-14042-z [Google Scholar]
-
Shkoporov AN, Clooney AG, Sutton TDS, Ryan FJ, Daly KM, Nolan JA, et al. The Human Gut Virome Is Highly Diverse, Stable, and Individual Specific. Cell Host Microbe 2019, 26, 527–541.e5. DOI:10.1016/j.chom.2019.09.009 [Google Scholar]
-
Brussow H. What is needed for phage therapy to become a reality in Western medicine? Virology 2012, 434, 138–142. DOI:10.1016/j.virol.2012.09.015 [Google Scholar]
-
Chan BK, Abedon ST, Loc-Carrillo C. Phage cocktails and the future of phage therapy. Future Microbiol. 2013, 8, 769–783. DOI:10.2217/fmb.13.47 [Google Scholar]
-
Menor-Flores M, Vega-Rodriguez MA, Molina F. Computational design of phage cocktails based on phage-bacteria infection networks. Comput. Biol. Med. 2022, 142, 105186. DOI:10.1016/j.compbiomed.2021.105186 [Google Scholar]
-
Pelfrene E, Willebrand E, Cavaleiro Sanches A, Sebris Z, Cavaleri M. Bacteriophage therapy: A regulatory perspective. J. Antimicrob. Chemother. 2016, 71, 2071–2074. DOI:10.1093/jac/dkw083 [Google Scholar]
-
Bochkareva SS, Karaulov AV, Aleshkin AV, Novikova LI, Kiseleva IA, Rubal’skii EO, et al. Analysis of the Pharmacokinetics of Suppository Forms of Bacteriophages. Bull. Exp. Biol. Med. 2020, 168, 748–752. DOI:10.1007/s10517-020-04794-w [Google Scholar]
-
Corbellino M, Kieffer N, Kutateladze M, Balarjishvili N, Leshkasheli L, Askilashvili L, et al. Eradication of a Multidrug-Resistant, Carbapenemase-Producing Klebsiella pneumoniae Isolate Following Oral and Intra-rectal Therapy with a Custom Made, Lytic Bacteriophage Preparation. Clin. Infect. Dis. 2020, 70, 1998–2001. DOI:10.1093/cid/ciz782 [Google Scholar]
-
Miedzybrodzki R, Kasprzak H, Letkiewicz S, Rogoz P, Zaczek M, Thomas J, et al. Pharmacokinetic and Pharmacodynamic Obstacles for Phage Therapy From the Perspective of Clinical Practice. Clin. Infect. Dis. 2023, 77, S395–S400. DOI:10.1093/cid/ciad516 [Google Scholar]
-
Schooley RT, Biswas B, Gill JJ, Hernandez-Morales A, Lancaster J, Lessor L, et al. Development and Use of Personalized Bacteriophage-Based Therapeutic Cocktails To Treat a Patient with a Disseminated Resistant Acinetobacter baumannii Infection. Antimicrob. Agents Chemother. 2017, 61, 10-1128. DOI:10.1128/AAC.00954-17 [Google Scholar]
-
Febvre HP, Rao S, Gindin M, Goodwin NDM, Finer E, Vivanco JS, et al. PHAGE Study: Effects of Supplemental Bacteriophage Intake on Inflammation and Gut Microbiota in Healthy Adults. Nutrients 2019, 11, 666. DOI:10.3390/nu11030666 [Google Scholar]
-
Gindin M, Febvre HP, Rao S, Wallace TC, Weir TL. Bacteriophage for Gastrointestinal Health (PHAGE) Study: Evaluating the Safety and Tolerability of Supplemental Bacteriophage Consumption. J. Am. Coll. Nutr. 2019, 38, 68–75. DOI:10.1080/07315724.2018.1483783 [Google Scholar]
-
McCallin S, Alam Sarker S, Barretto C, Sultana S, Berger B, Huq S, et al. Safety analysis of a Russian phage cocktail: From metagenomic analysis to oral application in healthy human subjects. Virology 2013, 443, 187–196. DOI:10.1016/j.virol.2013.05.022 [Google Scholar]
-
Pargin E, Roach MJ, Skye A, Papudeshi B, Inglis LK, Mallawaarachchi V, et al. The human gut virome: Composition, colonization, interactions, and impacts on human health. Front. Microbiol. 2023, 14, 963173. DOI:10.3389/fmicb.2023.963173 [Google Scholar]
-
Townsend EM, Kelly L, Muscatt G, Box JD, Hargraves N, Lilley D, et al. The Human Gut Phageome: Origins and Roles in the Human Gut Microbiome. Front. Cell Infect. Microbiol. 2021, 11, 643214. DOI:10.3389/fcimb.2021.643214 [Google Scholar]
-
Boggione DMG, Batalha LS, Gontijo MTP, Lopez MES, Teixeira AVNC, Santos IJB, et al. Evaluation of microencapsulation of the UFV-AREG1 bacteriophage in alginate-Ca microcapsules using microfluidic devices. Colloids Surf. B Biointerfaces 2017, 158, 182–189. DOI:10.1016/j.colsurfb.2017.06.045 [Google Scholar]
-
Soto MJ, Retamales J, Palza H, Bastías R. Encapsulation of specific Salmonella Enteritidis phage f3αSE on alginate-spheres as a method for protection and dosification. Electron. J. Biotechnol. 2018, 31, 57–60. DOI:10.1016/j.ejbt.2017.11.006 [Google Scholar]
-
Stanford K, McAllister TA, Niu YD, Stephens TP, Mazzocco A, Waddell TE, et al. Oral delivery systems for encapsulated bacteriophages targeted at Escherichia coli O157:H7 in feedlot cattle. J. Food Prot. 2010, 73, 1304–1312. DOI:10.4315/0362-028x-73.7.1304 [Google Scholar]
-
Jo SJ, Lee YM, Cho K, Park SY, Kwon H, Giri SS, et al. Standardization of the Agar Plate Method for Bacteriophage Production. Antibiotics 2024, 14, 2. DOI:10.3390/antibiotics14010002 [Google Scholar]
-
Ács N, Gambino M, Brøndsted L. Bacteriophage Enumeration and Detection Methods. Front. Microbiol. 2020, 11, 594868. DOI:10.3389/fmicb.2020.594868 [Google Scholar]
-
Fridholm H, Everitt E. Rapid and reproducible infectivity end-point titration of virulent phage in a microplate system. J. Virol. Methods 2005, 128, 67–71. DOI:10.1016/j.jviromet.2005.04.004 [Google Scholar]
-
Javaudin F, Latour C, Debarbieux L, Lamy-Besnier Q. Intestinal Bacteriophage Therapy: Looking for Optimal Efficacy. Clin. Microbiol. Rev. 2021, 34, e0013621. DOI:10.1128/CMR.00136-21 [Google Scholar]
-
Danis-Wlodarczyk K, Dabrowska K, Abedon ST. Phage Therapy: The Pharmacology of Antibacterial Viruses. Curr. Issues Mol. Biol. 2021, 40, 81–164. DOI:10.21775/cimb.040.081 [Google Scholar]
-
Barr JJ, Auro R, Furlan M, Whiteson KL, Erb ML, Pogliano J, et al. Bacteriophage adhering to mucus provide a non-host-derived immunity. Proc. Natl. Acad. Sci. USA 2013, 110, 10771–10776. DOI:10.1073/pnas.1305923110 [Google Scholar]
-
Barr JJ, Auro R, Sam-Soon N, Kassegne S, Peters G, Bonilla N, et al. Subdiffusive motion of bacteriophage in mucosal surfaces increases the frequency of bacterial encounters. Proc. Natl. Acad. Sci. USA 2015, 112, 13675–13680. DOI:10.1073/pnas.1508355112 [Google Scholar]
-
Bichet MC, Chin WH, Richards W, Lin Y-W, Avellaneda-Franco L, Hernandez CA, et al. Bacteriophage uptake by mammalian cell layers represents a potential sink that may impact phage therapy. iScience 2021, 24, 102287. DOI:10.1016/j.isci.2021.102287 [Google Scholar]
-
Douadi C, Theodorou I, Lamy-Besnier Q, Schiettekatte O, Sbardella Y, Brot L, et al. Differential translocation of bacteriophages across the intestinal barrier in health and Crohn’s disease. Cell Rep. 2024, 45, 116726. DOI:10.1101/2024.09.17.613249 [Google Scholar]
-
Parracho HM, Burrowes BH, Enright MC, McConville ML, Harper DR. The role of regulated clinical trials in the development of bacteriophage therapeutics. J. Mol. Genet. Med. 2012, 6, 279–286. DOI:10.4172/1747-0862.1000050 [Google Scholar]
-
Eriksen RS, Mitarai N, Sneppen K. Sustainability of spatially distributed bacteria-phage systems. Sci. Rep. 2020, 10, 3154. DOI:10.1038/s41598-020-59635-7 [Google Scholar]
-
Armstrong HK, Bording-Jorgensen M, Santer DM, Zhang Z, Valcheva R, Rieger AM, et al. Unfermented β-fructan Fibers Fuel Inflammation in Select Inflammatory Bowel Disease Patients. Gastroenterology 2023, 164, 228–240. DOI:10.1053/j.gastro.2022.09.034 [Google Scholar]
-
Rytter H, Naimi S, Wu G, Lewis J, Duquesnoy M, Vigué L, et al. In vitro microbiota model recapitulates and predicts individualised sensitivity to dietary emulsifier. Gut 2025, 74, 761–774. DOI:10.1136/gutjnl-2024-333925 [Google Scholar]
-
Carroll-Portillo A, Lin HC. Bacteriophage and the Innate Immune System: Access and Signaling. Microorganisms 2019, 7, 625. DOI:10.3390/microorganisms7120625 [Google Scholar]
-
Podlacha M, Grabowski L, Kosznik-Kawsnicka K, Zdrojewska K, Stasilojc M, Wegrzyn G, et al. Interactions of Bacteriophages with Animal and Human Organisms-Safety Issues in the Light of Phage Therapy. Int. J. Mol. Sci. 2021, 22, 8937. DOI:10.3390/ijms22168937 [Google Scholar]
-
Van Belleghem JD, Dabrowska K, Vaneechoutte M, Barr JJ, Bollyky PL. Interactions between Bacteriophage, Bacteria, and the Mammalian Immune System. Viruses 2018, 11, 10. DOI:10.3390/v11010010 [Google Scholar]
-
Grubb DS, Wrigley SD, Freedman KE, Wei Y, Vazquez AR, Trotter RE, et al. PHAGE-2 Study: Supplemental Bacteriophages Extend Bifidobacterium animalis subsp. lactis BL04 Benefits on Gut Health and Microbiota in Healthy Adults. Nutrients 2020, 12, 2474. DOI:10.3390/nu12082474 [Google Scholar]
-
Hill C, Guarner F, Reid G, Gibson GR, Merenstein DJ, Pot B, et al. Expert consensus document. The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat. Rev. Gastroenterol. Hepatol. 2014, 11, 506–514. DOI:10.1038/nrgastro.2014.66 [Google Scholar]
-
Lamy-Besnier Q, Chaffringeon L, Lourenço M, Payne RB, Trinh JT, Schwartz JA, et al. Prophylactic Administration of a Bacteriophage Cocktail Is Safe and Effective in Reducing Salmonella enterica Serovar Typhimurium Burden In Vivo. Microbiol. Spectr. 2021, 9, e00497-21. DOI:10.1128/Spectrum.00497-21 [Google Scholar]
-
Matsuzaki S, Rashel M, Uchiyama J, Sakurai S, Ujihara T, Kuroda M, et al. Bacteriophage therapy: A revitalized therapy against bacterial infectious diseases. J. Infect. Chemother. 2005, 11, 211–219. DOI:10.1007/s10156-005-0408-9 [Google Scholar]
-
Egido JE, Costa AR, Aparicio-Maldonado C, Haas P-J, Brouns SJJ. Mechanisms and clinical importance of bacteriophage resistance. FEMS Microbiol. Rev. 2022, 46, fuab048. DOI:10.1093/femsre/fuab048 [Google Scholar]
-
Dupuis M-È, Villion M, Magadán AH, Moineau S. CRISPR-Cas and restriction-modification systems are compatible and increase phage resistance. Nat. Commun. 2013, 4, 2087. DOI:10.1038/ncomms3087 [Google Scholar]
-
Dy RL, Richter C, Salmond GPC, Fineran PC. Remarkable Mechanisms in Microbes to Resist Phage Infections. Annu. Rev. Virol. 2014, 1, 307–331. DOI:10.1146/annurev-virology-031413-085500 [Google Scholar]
-
Liu Y, Dai L, Dong J, Chen C, Zhu J, Rao VB, et al. Covalent Modifications of the Bacteriophage Genome Confer a Degree of Resistance to Bacterial CRISPR Systems. J. Virol. 2020, 94, e01630-20. DOI:10.1128/JVI.01630-20 [Google Scholar]
-
Bleriot I, Pacios O, Blasco L, Fernández-García L, López M, Ortiz-Cartagena C, et al. Improving phage therapy by evasion of phage resistance mechanisms. JAC-Antimicrob. Resist. 2023, 6, dlae017. DOI:10.1093/jacamr/dlae017 [Google Scholar]
-
Arcilla MS, van Hattem JM, Haverkate MR, Bootsma MCJ, van Genderen PJJ, Goorhuis A, et al. Import and spread of extended-spectrum beta-lactamase-producing Enterobacteriaceae by international travellers (COMBAT study): A prospective, multicentre cohort study. Lancet Infect. Dis. 2017, 17, 78–85. DOI:10.1016/S1473-3099(16)30319-X [Google Scholar]
-
Armand-Lefevre L, Andremont A, Ruppe E. Travel and acquisition of multidrug-resistant Enterobacteriaceae. Med. Mal. Infect. 2018, 48, 431–441. DOI:10.1016/j.medmal.2018.02.005 [Google Scholar]
-
Rondinaud E, Clermont O, Petitjean M, Ruppe E, Esposito-Farese M, Nazimoudine A, et al. Acquisition of Enterobacterales carrying the colistin resistance gene mcr following travel to the tropics. J. Travel. Med. 2023, 30, taac141. DOI:10.1093/jtm/taac141 [Google Scholar]
