Validation of Thymidylate Synthase as a Key Gene Inhibiting WSSV Proliferation in Procambarus clarkii
Received: 31 December 2025 Revised: 24 February 2026 Accepted: 30 March 2026 Published: 10 April 2026
© 2026 The authors. This is an open access article under the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/).
1. Introduction
The red swamp crayfish (P. clarkii) is a prominent aquaculture species in China, valued for its palatable flavor and high nutritional content [1]. However, the sustainability of its farming is increasingly threatened by infectious diseases, particularly white spot syndrome (WSS), which is caused by the white spot syndrome virus (WSSV; genus Whispovirus, family Nimaviridae) [2]. WSSV exhibits a broad host range among crustaceans and possesses an enveloped genome of approximately 300,000 base pairs, encoding 448 putative open reading frames (ORFs) [3]. The majority of these genes are implicated in critical viral processes, including host cell invasion, replication, immune evasion, and induction of host apoptosis. High-density transportation practices further exacerbate the risk of WSSV transmission and disease outbreaks by inducing physiological stress, immunosuppression, and physical injuries in crayfish populations [4].
The two conserved WSSV genes, thymidylate synthase (TS) and ribonucleotide reductase (RR), have been the focus of research for their potential application in the prevention and control of WSS [5,6]. TS catalyzes the conversion of dUMP to dTMP (a rate-limiting step for the DNA replication precursor dTTP) and is an early-expressed WSSV gene (detectable at 4 h post-infection, peaking at 24 h) [7]. This enzyme, in concert with RR and thymidine kinase (TK), hijacks host dUMP for viral replication, thereby accelerating the death process. RR (composed of RR1/RR2 subunits) is the sole ribonucleotide-deoxyribonucleotide transaminase and is crucial as a rate-limiting enzyme for WSSV genome amplification [8]. RNA interference (RNAi) mediates post-transcriptional gene silencing: Dicer enzymes cleave double-stranded RNA into siRNA, guiding RISC to degrade target mRNAs [9,10]. RNAi exhibits significant inhibitory effects against WSSV: double-stranded RNAs (dsRNA) targeting either host genes (e.g., Akirin, proPO, ferritin, Rab7) or viral genes (e.g., GDH, ASAT, VP28-associated LvAMP13.4) effectively suppress replication [11,12,13,14,15,16]. Previous studies demonstrated that homologous dsRNA interference targeting the RR2 subunit suppresses WSSV replication [17]. However, the practical application of dsRNA is limited by high costs and error rates associated with chemical synthesis. And mainly by the delivery method, being fundamental to the development of oral-delivery strategies [10,18,19]. Large-scale dsRNA production can be achieved using an enzymatic in vivo expression system based on T7 polymerase-expressing Escherichia coli strain HT115 (DE3), an RNase III-deficient strain, carrying the L4440 plasmid that contains inverted T7 promoters [20].
In this study, the TS and RR genes (specifically the RR1 subunit of the RR gene) were selected as target candidates for the prevention and control of WSSV in P. clarkii using RNAi technology. The functional efficacy of dsRNAs targeting WSSV TS (dsRNA-TS) and RR (dsRNA-RR) will first be validated. Subsequently, an engineered E. coli HT115 strain will be constructed to enable high-yield expression of target dsRNAs. Following isolation and purification of the dsRNAs, antiviral efficacy will be assessed based on the effective and more valuable gene target, paving the way for large-scale, low-cost dsRNA production. This research aims to develop a sustainable biological control strategy against WSSV in crayfish aquaculture. The findings are expected to contribute to the optimization of prokaryotic dsRNA production systems, the development of targeted delivery approaches, and the integration of RNAi with molecular breeding techniques, thereby advancing sustainable disease management practices in aquaculture.
2. Materials and Methods
2.1. Experimental Crayfish
Healthy crayfish were sourced from the aquaculture base of Huazhong Agricultural University. The crayfish were reared in tanks maintained at 24–26 °C, fed once daily with commercial pellet feed (Tongwei Co., Ltd., Chengdu, China) at a rate of 2% of body weight. Tank water was changed every 2 days to maintain water quality. Only active crayfish with intact limbs were selected for the experiment.
2.2. Preparation and Efficacy Validation of dsRNA-TS and dsRNA-RR
DsRNAs targeting WSSV TS (dsRNA-TS) and RR (dsRNA-RR) were synthesized via in vitro transcription using the T7RiboMAX™ RNAi Expression System (Promega, Madison, WI, USA). The TS gene was amplified with primers dsRNA-TS-F/R, and the RR gene with dsRNA-RR-F/R (Table 1).
Twelve healthy laboratory-reared crayfish (body weight: 20–25 g) were selected. Prior to the experiment, 3 crayfish were randomly sampled for muscle tissue, and WSSV copy number was detected via qPCR to confirm they were WSSV-free (The WSSV strain comes from the laboratory archive). The remaining 9 crayfish were randomly divided into 3 groups (n = 3 per group): Positive control group: Injected with 0.1 mL WSSV only; dsRNA-TS group: Injected with 0.1 mL of a mixture containing 1.5 µg/g (body weight) dsRNA-TS and 105 WSSV copies; dsRNA-RR group: Injected with 0.1 mL of a mixture containing 1.5 µg/g (body weight) dsRNA-RR and 105 WSSV copies.
All injections were administered into the second abdominal segment muscle. Based on the WSSV inhibition results observed in the dsRNA-TS and dsRNA-RR groups, the efficacy of dsRNA-TS and dsRNA-RR in suppressing WSSV replication was evaluated. In this experiment, WSSV was administered via intramuscular injection to induce infection. The virus spreads from the injection site inward, thereby preventing false-negative results in other tissues caused by slow viral spread. This experiment selected muscle tissue for WSSV testing, and tail muscle tissue is also within the range of WSSV infection [21].
Table 1. Primer sequences.
|
Name |
Sequence |
Note |
|---|---|---|
|
WSSV-F |
ATGGATCTTTCTTTCACTCTTTCGG |
|
|
WSSV-R |
TCCGCATCTTCTTCCTTCATC |
|
|
dsRNA-TS-F |
TAATACGACTCACTATAGGG-ACATCTGGAACGCTAATGGG |
T7 promoter upstream of “-” |
|
dsRNA-TS-R |
TAATACGACTCACTATAGGG-AAATGTGTGCGTCACCAAGA |
T7 promoter upstream of “-” |
|
dsRNA-RR-F |
TAATACGACTCACTATAGGG-GCGAAGAGTTTAAGGCGTTG |
T7 promoter upstream of “-” |
|
dsRNA-RR-R |
TAATACGACTCACTATAGGG-CTAGTCCCTGCACACCCAAT |
T7 promoter upstream of “-” |
|
18S-F |
TATACGCTAGTGGAGCTGGAA |
|
|
18S-R |
GGGGAGGTAGTGACGAAAAAT |
|
|
dsRNA-TS2-F |
CCCAAGCTTGGG-ATGGAGGGAGAACATCAAT |
Restriction enzyme site upstream of “-” |
|
dsRNA-TS2-R |
GGGGTACCCC-TCAAACAGCCATTTTCATC |
Restriction enzyme site upstream of “-” |
2.3. RNAi Assay
Twenty crayfish (weight: 15–20 g) were randomly assigned to two groups: (i) the control group, which was injected with WSSV suspension (105 copies); and (ii) the dsRNA-TS group, which was injected with a mixture of WSSV (105 copies) and dsRNA-TS (1.5 μg/g body weight). All injection volumes were adjusted to 0.1 mL with PBS. Survival rates were observed and recorded daily. On days 1, 3, 5, 7, and 14 post-injection, three crayfish were randomly selected from each group to collect appendage samples. These were placed in anhydrous ethanol and stored at −20 °C for subsequent analysis.
2.4. RNAi Assay Under Simulated Transport Stress
Sixty crayfish (weight: 15–20 g) were evenly divided into two groups (control and dsRNA-TS group), with each group comprising three subgroups (n = 10 per subgroup). Injection methods and dosages followed Section 2.3. 2 h post-injection, all six subgroups were transferred to six separate tanks. Subsequently, three subgroups from each group were transferred to identical plastic baskets for simulated dry transport: horizontal shaking for 3 min every 20 min under water-free conditions. After 2 h of simulated transport, crayfish were returned to their original tanks for 14 days of rearing. Survival rates were recorded daily, and appendage samples were collected on days 1, 4, 7, and 14 post-transport.
2.5. Prokaryotic dsRNA Extraction and RNA Interference
Recombinant plasmids were constructed by cloning the TS fragment into the L4440 plasmid vector, kindly provided by Professor Donghai Peng’s laboratory. The TS gene was amplified using primers dsRNA-TS2-F/R (Table 1). The recombinant plasmid was transformed into E. coli HT115 (DE3) competent cells.
After sequencing verification of the recombinant plasmid, positive clones were inoculated into LB medium and cultured to the logarithmic growth phase (OD600 ≈ 0.6). Isopropyl-β-D-thiogalactoside (IPTG) was added to a final concentration of 0.5 mmol/L. Cultures were shaken continuously at 37 °C for 4 h to induce dsRNA expression. Bacterial cells were collected and extracted for dsRNA (dsRNA-TS2) using the TRIzol method. Residual single-stranded RNA and DNA were digested with RNase A and DNase I, respectively, to obtain high-purity dsRNA. RNAi experiments were conducted as described in Section 2.2 to validate the inhibitory efficacy against WSSV.
2.6. WSSV Copy Number Determination via Absolute qPCR
To quantitatively determine WSSV content in crayfish appendages, absolute quantitative PCR was performed using primers WSSV-F/R (Table 1). A standard curve was established by performing a 10-fold serial dilution of the pMD18-T plasmid (Takara, Otsu, Japan) containing the target WSSV DNA fragment. The number of WSSV genomic copies per 1 μg of muscle template DNA was calculated.
2.7. Data Processing and Bioinformatics Analysis
Experimental data were statistically analyzed using SPSS 23.0 software. The qPCR results are expressed as mean ± standard deviation. Differences between groups at the same time point were analyzed using t-tests; multiple comparisons of data from different time points within the same group were performed using Duncan’s multiple range test. Statistical significance was defined as p ≤ 0.05 (*), and high significance as p ≤ 0.01 (**). Graphs were generated using GraphPad Prism 9.0 software (GraphPad Software, San Diego, CA, USA). For bioinformatics analysis, WSSV and P. clarkii TS gene sequences were retrieved from the NCBI database. Sequence-to-protein alignment was performed using NCBI BLAST, and the TS phylogenetic tree was constructed with MEGA 11 software.
3. Results
3.1. Bioinformatics Analysis of TS and RR
Preliminary homology analysis of WSSV TS and RR with crayfish TS and RR via BLAST indicates homology exists only between WSSV TS and crayfish TS. The TS gene sequences and RR sequences are shown in Figure 1a,b. To further explore potential functional interactions between the virus and host TS, a series of bioinformatics analyses compared WSSV TS with TS from common farmed shrimp and crabs. Phylogenetic tree construction assessed the evolutionary relationship between WSSV TS and crustacean TS (Figure 1c). The phylogenetic tree constructed based on TS proteins clearly shows that WSSV TS diverges significantly from shrimp and crab TS, forming an independent branch, indicating a distant evolutionary relationship between the two. However, similarities persist—for instance, BLAST analysis revealed 64% homology and 77% sequence similarity between P. clarkii TS and WSSV TS, with localized homologous regions. These localized homologous regions may represent functionally relevant domains, laying the groundwork for investigating potential virus-host interactions (e.g., competitive binding of substrates or cofactors).
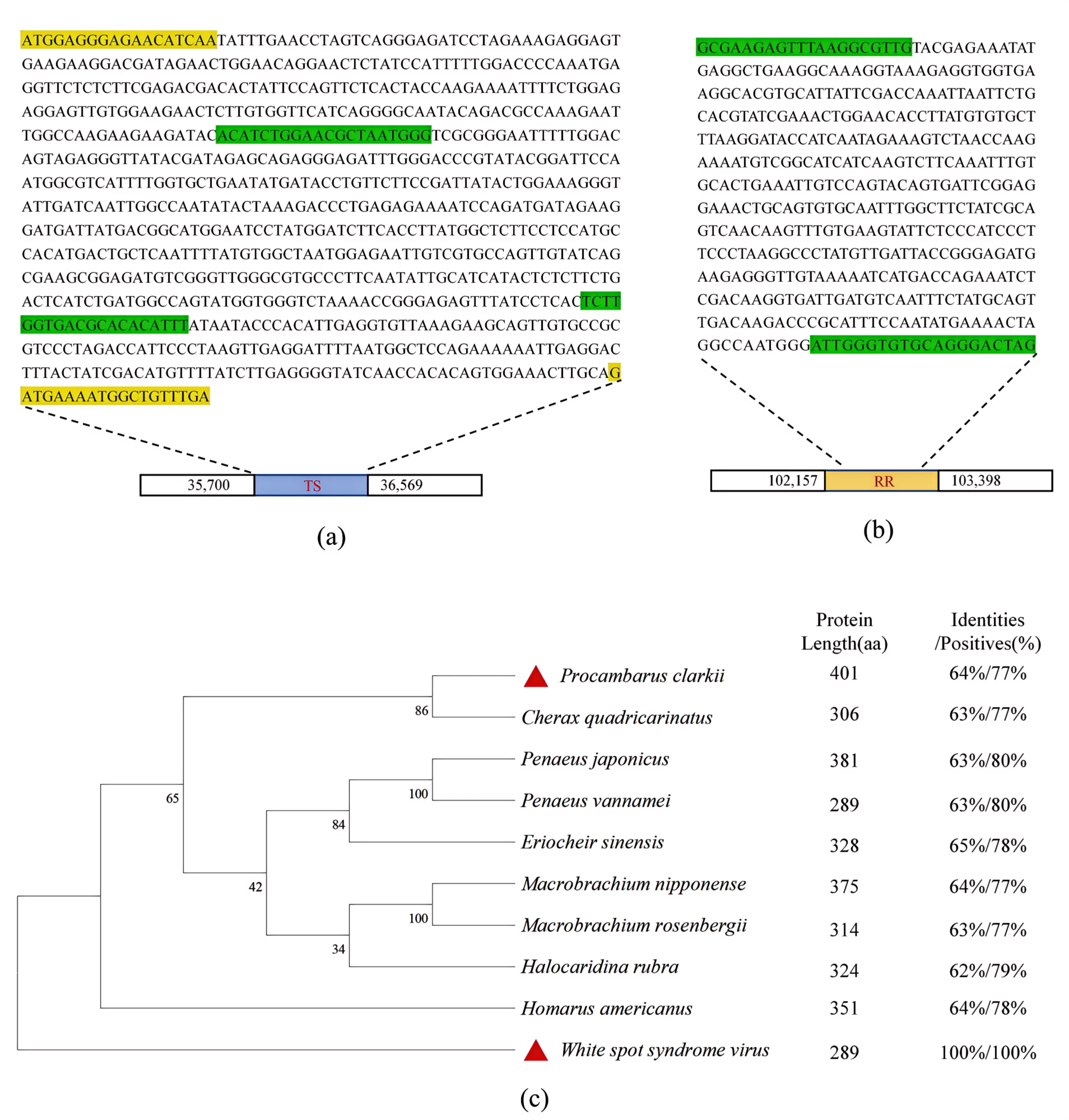
Figure 1. Bioinformatics Analysis of the TS and RR Gene. Note: (a) Nucleotide sequences and genomic locations of the WSSV TS and RR genes (genome reference: NCBI RefSeq: GCF_003024735.1), with sequences amplified using the T7 kit highlighted in green and those amplified for prokaryotic expression highlighted in yellow; (b) Nucleotide sequence of WSSV RR1 subunit gene, with the T7 kit amplification sequence highlighted in green; (c) Phylogenetic tree analysis of TS.
3.2. Validation of dsRNA-TS and dsRNA-RR Efficacy
To further examine the impact of RR and TS gene silencing on WSSV replication, dsRNA synthesis was performed using the TS and RR1 subunit sequences (Figure 1a,b). The experimental workflow for evaluating dsRNA-TS&dsRNA-RR’s inhibitory effect on WSSV is shown in Figure 2a. The efficacy of dsRNA-TS and dsRNA-RR in inhibiting the replication of WSSV was evaluated using qPCR. Results demonstrated that dsRNA-TS and dsRNA-RR exhibited highly significant inhibitory effects on WSSV replication compared to the control group injected with WSSV alone (Figure 1c). The RR2 subunit in the RR gene has been demonstrated to play a crucial role in WSSV replication [22]. And the RR1 subunit in this study exhibits consistent functional effects [23]. Furthermore, the TS gene shares significant homology with WSSV and commonly farmed shrimp and crabs, offering greater potential for virus-host interactions and compatibility with molecular breeding approaches—an advantage not present in the RR gene. Considering factors such as these, the TS gene was subsequently selected for further research in this study.
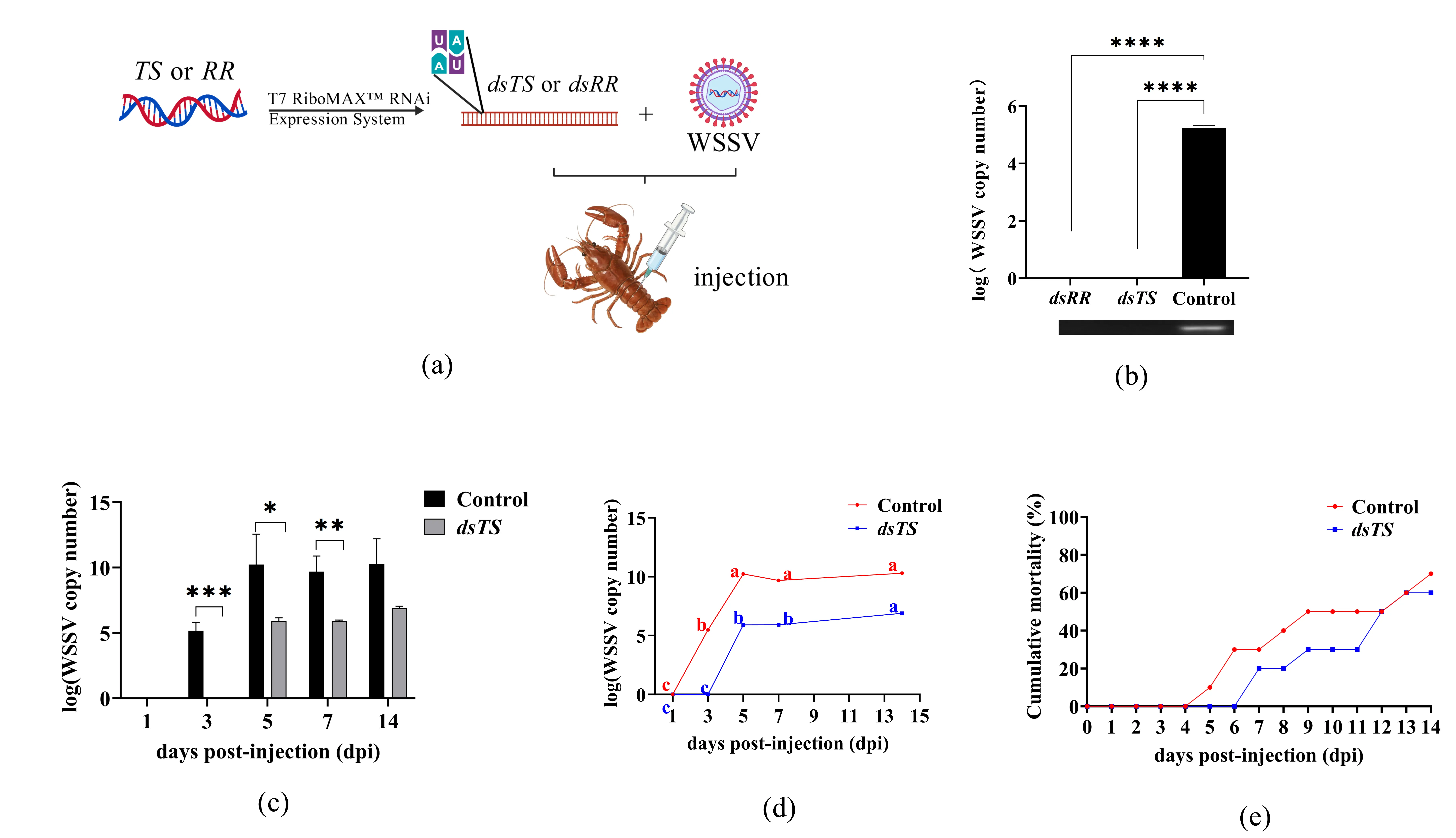
Figure 2. Effect of dsRNA-TS on WSSV replication in appendage tissue of P. clarkii. Note: (a) Experimental workflow for WSSV inhibition by dsRNA-TS and dsRNA-RR; (b) Interference effects of dsRNA-TS and dsRNA-RR detected by qPCR; the control group was injected with WSSV only (**** p ≤ 0.0001). (c) WSSV copy number in appendage tissue of the control and dsRNA-TS groups at different time points (*** p ≤ 0.001; ** p ≤ 0.01; * p ≤ 0.05); (d) Dynamic changes in WSSV copy number in appendages of the control and dsRNA-TS groups on days 1–14 post-injection (different letters indicate significant differences among time points within the same group); (e) Cumulative mortality of P. clarkii within 14 dpi.
3.3. Effect of dsRNA-TS on WSSV Replication
The experimental procedure for evaluating the inhibitory effect of dsRNA-TS on WSSV is also consistent with that shown in Figure 2a. The number of WSSV copies in the appendage tissues of P. clarkii was quantified at different time points post-injection using qPCR technology. Results revealed significant differences in viral copy numbers between the dsRNA-TS group and the control group on days 3, 5, and 7 post-injection (Figure 2c). Notably, no WSSV was detected in the dsRNA-TS group at day 3—a key finding indicating potent early-stage viral suppression (Figure 2c). Dynamic monitoring of viral load over 14 days further confirmed sustained suppression in the dsRNA-TS group, with consistently lower viral copy numbers than the control group (Figure 2d; different letters denote significant differences between time points within the same group).
Regarding survival rates, the cumulative mortality in the dsRNA-TS group after 14 dpi was 60%, representing a 10% reduction compared to the 70% mortality in the control group (Figure 2e). These results indicate that dsRNA-TS not only suppresses viral replication but also enhances survival rates in crayfish during WSSV infection.
3.4. Effects of dsRNA-TS on WSSV After Transportation Stress
To simulate real aquaculture conditions, this study evaluated the efficacy of dsRNA-TS under simulated transport stress conditions. The experimental design is shown in Figure 3a. Real-time quantitative PCR analysis revealed no significant differences in WSSV copy numbers between the dsRNA-TS group and the control group at most early time points, except on Day 1. However, significant differences emerged at 7 and 14 dpi, with markedly reduced viral loads in the dsRNA-TS group—confirming that dsRNA-TS maintains the inhibitory activity against the replication of WSSV under stress conditions (Figure 3b). The 14-day viral load trajectory further supported this conclusion: the dsRNA-TS group sustained consistently lower levels of viral replication compared to the control group (Figure 3c).
Notably, survival trends under stress differed from non-stress conditions: the cumulative mortality rate in the dsRNA-TS group reached 66.67% by day 14, significantly higher than the control group’s 36.67% mortality rate (Figure 3d). This contradictory result indicates that while dsRNA-TS retains its inhibitory activity against the replication of WSSV, This contradictory result indicates that it is not feasible to reduce the mortality rate during transportation merely by inhibiting the replication of WSSV in the body. Some scholars have demonstrated that the mortality of Procambarus clarkii after transport stress is caused by oxidative stress, the imbalance of reactive oxygen species regulation, and decreased resistance to WSSV, which eventually leads to irreversible tissue damage [4]. It may also be necessary to reduce other influencing factors, such as stress, that affect the survival rate after transportation.
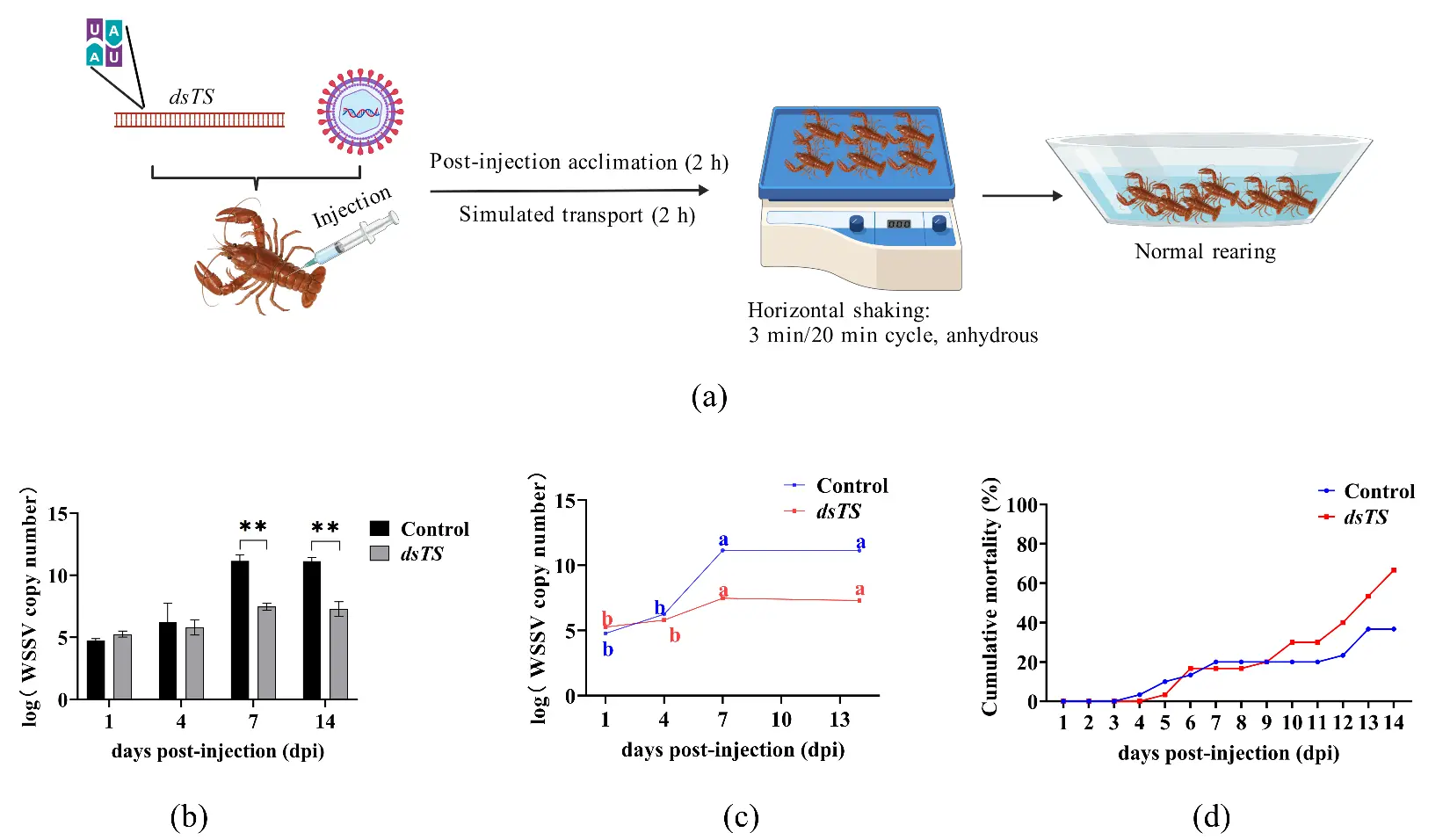
Figure 3. WSSV copy number and its changes in control and dsRNA-TS groups after transport stress. Note: (a) Experimental workflow for WSSV inhibition by dsRNA-TS; (b) WSSV copy number in appendage tissue of the control and dsRNA-TS groups at different time points (** p ≤ 0.01); (c) Dynamic changes in WSSV copy number in appendages of the control and dsRNA-TS groups on days 1–14 post-injection (different letters indicate significant differences among time points within the same group); (d) Cumulative mortality of P. clarkii within 14 dpi.
3.5. RNAi of Prokaryotically Expressed dsRNAs
To achieve low-cost application of dsRNA-TS, a cost-effective production pathway for double-stranded RNA (dsRNA) was explored by expressing dsRNA-TS in the E. coli HT115 (DE3) strain. The experimental workflow for prokaryotic expression of dsRNA is shown in Figure 4a. Gel electrophoresis confirmed successful dsRNA expression: dsRNA-TS was detected in lanes 1–2 (arrows indicate target bands; Figure 4b). However, qPCR analysis revealed no significant suppression of WSSV replication in the dsRNA-TS-treated group compared to the control group (Figure 4c). This lack of efficacy may be attributed to two key factors: (1) low concentration and insufficient purity of the extracted dsRNA (residual bacterial nucleic acids may interfere with RNAi activity) [24]; (2) insufficient response time for dsRNA to exert its effects of inhibiting the replication of WSSV [10].
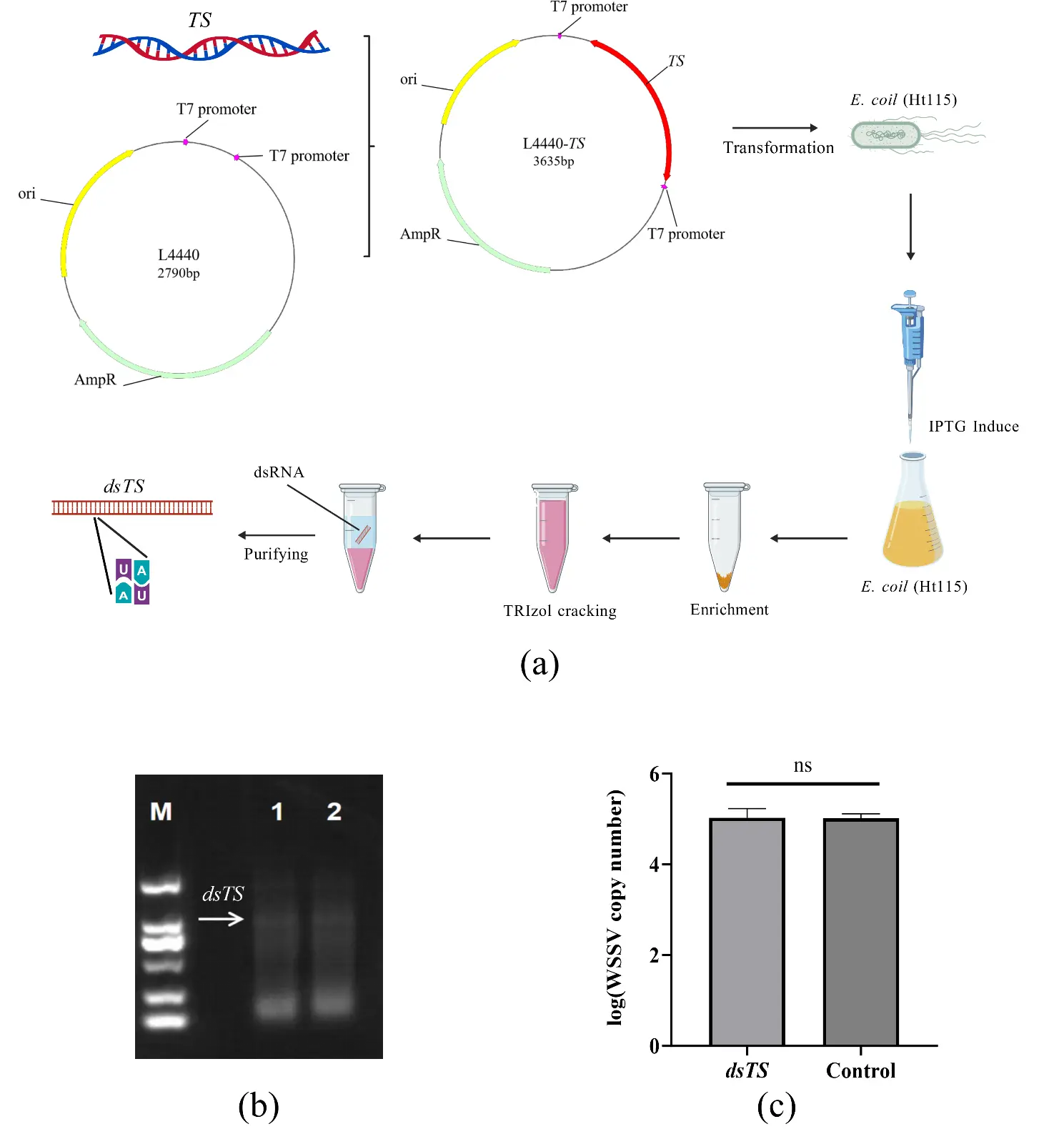
Figure 4. Workflow for dsRNA-TS Expression in a Prokaryotic System and Its Interference Effect on WSSV. Note: (a) The experimental workflow for prokaryotic expression of dsRNA; (b) Gel electrophoresis detection of dsRNA-TS (Lane M: DL2000 DNA Marker; Lanes 1–2: dsRNA-TS-containing mixture; arrow indicates dsRNA-TS); (c) Interference of dsRNA-TS Expressed in a Prokaryotic System on WSSV in P. clarkii (ns, not significant).
4. Discussion
RNAi technology has been extensively employed as a strategy to combat WSSV, with most studies targeting either host immune-related genes or viral envelope proteins [25,26,27,28]. However, viral nucleoside/nucleotide transferases, which serve as core regulators of viral DNA replication, have received relatively little attention as potential therapeutic targets against WSSV. Our findings demonstrate that in vitro transcribed dsRNA-TS effectively suppresses WSSV replication, thereby validating TS as a high-priority candidate target for antiviral intervention. As an early-expressed WSSV gene, silencing TS via dsRNA directly disrupts dTMP synthesis, providing a basis for the complete absence of detectable WSSV in the dsRNA-TS-treated group by 3 dpi [7]. This observation is consistent with previous studies indicating that RNAi-mediated silencing of viral metabolic genes impairs replication [11]. Notably, by targeting a virus-encoded enzyme, this approach minimizes potential off-target effects on host cellular processes. Furthermore, dsRNA-TS treatment reduced cumulative mortality from 70% to 60% and delayed mortality onset by two days. During the survival rate experiment, the random sampling of the crayfish’s appendages each time caused some damage to the crayfish, which in turn had a certain impact on their survival rate. Nevertheless, the survival rate in the experimental group continued to be higher than that in the control group. It is important to highlight that this window of time can be crucial at the field level, as it would provide time for an emergency harvest, thus reducing economic losses [10].
Under transport stress conditions, dsRNA-TS maintained significant inhibitory activity against WSSV from 7 to 14 dpi. However, it failed to improve survival rates, instead increasing mortality (66.67% vs. 36.67% in the control group). This paradoxical outcome stems from two factors: first, the presence of an anomalous control group exhibiting delayed mortality (13 days vs. 4–5 days) and lower viral loads, potentially linked to elevated baseline immunity. Second, transport stress reduces Dicer and Argonaute expression (key RNAi components) by 30–40% in crayfish [4]. Although dsRNA-TS suppressed viral replication, the combined effects of stress and dsRNA-induced hepatopancreatic damage increased mortality [19], underscoring the need to optimize dosages of dsRNA under stressed conditions This study confirmed that dsRNA-TS expressed in E. coli HT115 did not exhibit any inhibitory activity against WSSV replication, which may be related to dsRNA dysfunction. Three technical limitations explain this phenomenon: (1) Low purity and concentration of dsRNA obtained via TRIzol extraction—residual bacterial nucleic acids degrade dsRNA [24]; subsequent studies should quantify intact dsRNA using RNase If-qPCR [29]; (2) Prokaryotically expressed dsRNA often shows low activity due to production defects rather than host misrecognition. Unlike eukaryotic systems, prokaryotes lack protective RNA modifications (e.g., A-to-I) [30] and often yield heterogeneous, misfolded duplexes with cytotoxic contaminants (e.g., LPS). These impurities mask specific immune activation, while the uncontrolled, ultra-long length triggers non-specific effects (e.g., PKR activation) [31] that obscure RNAi or precise immune induction. Synthetic approaches enabling rational length design and chemical modification are therefore essential [32,33]. (3) Lack of protective delivery—free dsRNA is degraded by hemolymph ribonuclease, whereas encapsulated dsRNA (e.g., in recombinant baculovirus or virusoids) remains unaffected [22,34]. Alternatively, dsRNA can be delivered directly using microalgae or probiotics as delivery vehicles [35,36]. But the use of bacteria (E. coli) for dsRNA synthesis intended for use in the agri-food sector may be problematic in some jurisdictions due to risks to human health. Also consider the limitation that the dsRNA has been administered via injection (a method that is completely unfeasible economically under real-world industrial production conditions). Therefore, the challenge remains of finding a method of biological synthesis using another microorganism that can be administered directly, such as microalgae and probiotics, for example [10,35,36,37,38].
The conserved nature of WSSV TS and its irreplaceable role in replication further highlight its potential application in P. clarkii breeding programs. Molecular marker-assisted breeding (MAB)—accelerating trait selection through genotype-phenotype linkage—has successfully enhanced WSSV resistance in crustaceans: for instance, P. clarkii strains selected based on immune gene-associated SNPs exhibited a 60% higher WSSV survival rate than unselected populations [39]. Local sequence similarity between crayfish TS and WSSV TS suggests competitive inhibition, identifying the host TS gene as a potential regulator of viral replication. Screening natural crayfish populations for functional SNPs in host TS—particularly variants altering WSSV TS binding affinity or enhancing competitive inhibition—holds promise for developing naturally WSSV-resistant strains via MAB. Advances in gene editing technologies (e.g., CRISPR-Cas9) offer novel avenues for antiviral breeding utilizing WSSV TS. Arnon Pudgerd et al. [40] constructed a chimeric vector containing the Litopenaeus vannamei U6 promoter, WSSV-targeting gRNA, WSSV nucleoside reductase promoter, and the Cas9 protein from the pAC-sgRNA-Cas9 vector. Integration into the L. vannamei genome enables specific cleavage of WSSV DNA and replication inhibition. Applying this strategy to Litopenaeus vannamei (via engineered vectors targeting WSSV TS) achieves sequence-specific cleavage and replication suppression of the viral TS. Unlike RNAi technologies requiring repeated applications, gene-edited traits are heritable, making them a cost-effective solution for long-term aquaculture. However, this approach also presents the challenge of the legal difficulties involved in the development and use of genetically edited animals used for direct human consumption [10].
In summary, this study confirms three core conclusions: (1) In vitro transcribed dsRNA-TS effectively inhibits WSSV replication in P. clarkii; (2) Transport stress and individual immune variation mask the survival benefits of dsRNA, despite its retention of the activity to suppress WSSV replication; (3) dsRNA production in prokaryotes requires improved purity, activity, and delivery efficiency. Future research will aim to overcome the limitations of prokaryotic dsRNA expression and delivery methods, such as rationally designing the length of dsRNA fragments, performing specific chemical modifications, and selecting appropriate delivery vectors; optimizing application strategies in stress-prone environments, such as choosing the appropriate timing for RNA interference and combining treatments with other anti-stress agents; and utilizing key WSSV-resistant genes, such as heat shock protein genes, in selective breeding programs. Efforts will focus on integrating these short-term interventions with long-term breeding strategies—such as molecularly assisted selection using TS-interaction markers and CRISPR-based gene editing—to establish a multi-layered defense system against WSSV. This is crucial for fully realizing its potential and effectively reducing the global economic losses caused by WSSV in the crayfish aquaculture industry.
Statement of the Use of Generative AI and AI-Assisted Technologies in the Writing Process
AI solely assisted in completing refining academic language and designing the figure and table formats for the initial draft. The authors independently conducted all core research work and bear full responsibility for the scientific validity and originality of the study.
Acknowledgments
In this study, we would like to express our gratitude to Donghai Peng, College of Life Sciences, Huazhong Agricultural University, for his free donation of the L4440 plasmid.
Author Contributions
Z.Y.: Methodology, Investigation, Writing–original draft, Formal analysis. T.Z.: Methodology, Investigation. K.Y.: Methodology, Investigation. Y.C.: Methodology, Investigation. Y.X.: Methodology, Investigation. Y.Y.: Resources. Y.L.: Project administration, Supervision, Funding acquisition, Writing–review & editing, Resources, Conceptualization.
Ethics Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The authors declare that the data supporting the findings of this study are available within the paper.
Funding
This work was supported by the Fundamental Research Funds for the Central Universities (2662025FW008), and Fishery Seed Industry Joint Breeding Program of Jiangxi Province (sczy202501).
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
-
Yuan D, Wang L, Wang H, Miao R, Wang Y, Jin H, et al. Application of microalgae Scenedesmus acuminatus enhances water quality in rice-crayfish culture. Front. Bioeng. Biotechnol. 2023, 11, 1143622. DOI:10.3389/fbioe.2023.1143622 [Google Scholar]
-
Bao W, Tang K, Alcivar-Warren A. The complete genome of an endogenous nimavirus (nimav-1_LVa) from the pacific whiteleg shrimp Penaeus (litopenaeus) vannamei. Genes 2020, 11, 94–114. DOI:10.3390/genes11010094 [Google Scholar]
-
Liu WJ, Shiung HJ, Lo CF, Leu JH, Lai YJ, Lee TL, et al. Characterization and interactome study of white spot syndrome virus envelope protein VP11. PLoS ONE 2014, 9, e85779. DOI:10.1371/journal.pone.0085779 [Google Scholar]
-
Shi R, Yang S, Wang Q, Zhang L, Li Y. Effects of transport stress on immune response, physiological state, and WSSV concentration in the red swamp crayfish Procambarus clarkii. Aquac. Fish. 2023, 8, 498–508. DOI:10.1016/j.aaf.2022.01.001 [Google Scholar]
-
De-la-Re-Vega E, Garcia-Orozco KD, Arvizu-Flores AA, Yepiz-Plascencia G, Muhlia-Almazan A, Hernández J, et al. White spot syndrome virus Orf514 encodes a bona fide DNA polymerase. Molecules 2011, 16, 532–542. DOI:10.3390/molecules16010532 [Google Scholar]
-
Tsai MF, Lo CF, Van Hulten MCW, Tzeng HF, Chou CM, Huang CJ, et al. Transcriptional analysis of the ribonucleotide reductase genes of shrimp white spot syndrome virus. Virology 2000, 277, 92–99. DOI:10.1006/viro.2000.0596 [Google Scholar]
-
Li Q, Pan D, Zhang J, Yang F. Identification of the thymidylate synthase within the genome of white spot syndrome virus. J. Gen. Virol. 2004, 85, 2035–2044. DOI:10.1099/vir.0.80048-0 [Google Scholar]
-
Su MA, Huang YT, Chen IT, Lee DY, Hsieh YC, Li CY, et al. An invertebrate warburg effect: A shrimp virus achieves successful replication by altering the host metabolome via the PI3K-akt-mTOR pathway. PLoS Pathog. 2014, 10, e1004196. DOI:10.1371/journal.ppat.1004196 [Google Scholar]
-
Hough J, Howard JD, Brown S, Portwood DE, Kilby PM, Dickman MJ. Strategies for the production of dsRNA biocontrols as alternatives to chemical pesticides. Front. Bioeng. Biotechnol. 2022, 10, 980592. DOI:10.3389/fbioe.2022.980592 [Google Scholar]
-
Fajardo C, De Donato M, Macedo M, Charoonnart P, Saksmerprome V, Yang L, et al. RNA Interference Applied to Crustacean Aquaculture. Biomolecules 2024, 14, 1358. DOI:10.3390/biom14111358 [Google Scholar]
-
Li CY, Wang YJ, Huang SW, Cheng CS, Wang HC. Replication of the shrimp virus WSSV depends on glutamate-driven anaplerosis. PLoS ONE 2016, 11, e0146902. DOI:10.1371/journal.pone.0146902 [Google Scholar]
-
Thamizhvanan S, Nafeez Ahmed A, Vinoth Kumar D, Vimal S, Majeed SA, Taju G, et al. Silencing of prophenoloxidase (proPO) gene in freshwater prawn, Macrobrachium rosenbergii, makes them susceptible to white spot syndrome virus (WSSV). J. Fish Dis. 2021, 44, 573–584. DOI:10.1111/jfd.13297 [Google Scholar]
-
Xiong H, Jiang Y, Ji T, Zhang Y, Wei W, Yang H. The identification of a nuclear factor akirin with regulating the expression of antimicrobial peptides in red swamp crayfish (Procambarus clarkii). Int. J. Biol. Macromol. 2021, 183, 707–717. DOI:10.1016/j.ijbiomac.2021.04.153 [Google Scholar]
-
Yang F, Li S, Li F, Xiang J. A cuticle protein from the pacific white shrimp litopenaeus vannamei involved in WSSV infection. Dev. Comp. Immunol. 2018, 81, 303–311. DOI:10.1016/j.dci.2017.12.018 [Google Scholar]
-
Yang H, Liu Z, Jiang Q, Xu J, An Z, Zhang Y, et al. A novel ferritin gene from Procambarus clarkii involved in the immune defense against Aeromonas hydrophila infection and inhibits WSSV replication. Fish Shellfish Immunol. 2019, 86, 882–891. DOI:10.1016/j.fsi.2018.12.022 [Google Scholar]
-
Fajardo C, De Donato M, Charoonnart P, Saksmerprome V, Jackson HO, Mancera JM, et al. Rab proteins as potential therapeutic targets applied to crustacean aquaculture: Rab7 Case Analysis. Rev. Aquac. 2025, 17, e13005. DOI:10.1111/raq.13005 [Google Scholar]
-
Joo Hong S, Hong Kim K. Effects of length and sequence of long double-stranded RNAs targeting ribonucleotide reductase 2 of white spot syndrome virus (WSSV) on protective efficacy against WSSV. J. Invertebr. Pathol. 2023, 196, 107869. DOI:10.1016/j.jip.2022.107869 [Google Scholar]
-
Mu X, Greenwald E, Ahmad S, Hur S. An origin of the immunogenicity of in vitro transcribed RNA. Nucleic Acids Res. 2018, 46, 5239–5249. DOI:10.1093/nar/gky177 [Google Scholar]
-
Silver K, Cooper AM, Zhu KY. Strategies for enhancing the efficiency of RNA interference in insects. Pest Manag. Sci. 2021, 77, 2645–2658. DOI:10.1002/ps.6277 [Google Scholar]
-
De-Souza EA, Camara H, Salgueiro WG, Moro RP, Knittel TL, Tonon G, et al. RNA interference may result in unexpected phenotypes in caenorhabditis elegans. Nucleic Acids Res. 2019, 47, 3957–3969. DOI:10.1093/nar/gkz154 [Google Scholar]
-
Usman A, Andayani S, Herawati EY. White spot syndrome virus (WSSV) detection at traditional ponds of Lithopenaeus vannamei in Pasuruan district. J. Exp. Life Sci. 2018, 8, 173–176. DOI:10.21776/ub.jels.2018.008.03.07 [Google Scholar]
-
Rattanarojpong T, Khankaew S, Khunrae P, Vanichviriyakit R, Poomputsa K. Recombinant baculovirus mediates dsRNA specific to rr2 delivery and its protective efficacy against WSSV infection. J. Biotechnol. 2016, 229, 44–52. DOI:10.1016/j.jbiotec.2016.05.007 [Google Scholar]
-
Kolberg M. Structure, function, and mechanism of ribonucleotide reductases. Biochim. Et Biophys. Acta (BBA)—Proteins Proteom. 2004, 1699, 1–34. DOI:10.1016/S1570-9639(04)00054-8 [Google Scholar]
-
Smirnov A. How global RNA-binding proteins coordinate the behaviour of RNA regulons: An information approach. Comput. Struct. Biotechnol. J. 2022, 20, 6317–6338. DOI:10.1016/j.csbj.2022.11.019 [Google Scholar]
-
Wuthisathid K, Chaijarasphong T, Chotwiwatthanakun C, Somrit M, Sritunyalucksana K, Itsathitphaisarn O. Co-expression of double-stranded RNA and viral capsid protein in the novel engineered escherichia coli DualX-B15(DE3) strain. BMC Microbiol. 2021, 21, 88–99. DOI:10.1186/s12866-021-02148-8 [Google Scholar]
-
Shekhar MS, Lu Y. Application of nucleic-acid-based therapeutics for viral infections in Shrimp Aquaculture. Mar. Biotechnol. 2009, 11, 1–9. DOI:10.1007/s10126-008-9155-0 [Google Scholar]
-
Wang S, Song X, Zhang Z, Li H, Lǚ K, Yin B, et al. Shrimp with knockdown of LvSOCS2, a negative feedback loop regulator of JAK/STAT pathway in Litopenaeus vannamei, exhibit enhanced resistance against WSSV. Dev. Comp. Immunol. 2016, 65, 289–298. DOI:10.1016/j.dci.2016.07.021 [Google Scholar]
-
Alenton RRR, Kondo H, Hirono I, Maningas MBB. Gene silencing of VP9 gene impairs WSSV infectivity on Macrobrachium rosenbergii. Virus Res. 2016, 214, 65–70. DOI:10.1016/j.virusres.2016.01.013 [Google Scholar]
-
Wang P-H, Schulenberg G, Whitlock S, Worden A, Zhou N, Novak S, et al. RNase if -treated quantitative PCR for dsRNA quantitation of RNAi trait in genetically modified crops. BMC Biotech. 2018, 18, 3–15. DOI:10.1186/s12896-018-0413-6 [Google Scholar]
-
Liddicoat BJ, Piskol R, Chalk AM, Ramaswami G, Higuchi M, Hartner JC, et al. RNA editing by ADAR1 prevents MDA5 sensing of endogenous dsRNA as nonself. Science 2015, 349, 1115–1120. DOI:10.1126/science.aac7049 [Google Scholar]
-
Peisley A, Hur S. Multi-level regulation of cellular recognition of viral dsRNA. Cell Mol. Life Sci. 2012, 70, 1949–1963. DOI:10.1007/s00018-012-1149-4 [Google Scholar]
-
Sajeesh S, Lee TY, Hong SW, Dua P, Choe JY, Kang A, et al. Long dsRNA-mediated RNA interference and immunostimulation: A targeted delivery approach using polyethyleneimine based nano-carriers. Mol. Pharm. 2014, 11, 872–884. DOI:10.1021/mp400541z [Google Scholar]
-
Da Silva R, Viana VE, Avila LA, Zotti MJ, Smagghe G, Junior AM, et al. Advances on polymeric nanocarriers for sustainable agriculture: Enhancing dsRNA/siRNA delivery to combat agricultural pests. Int. J. Biol. Macromol. 2024, 282, 137000. DOI:10.1016/j.ijbiomac.2024.137000 [Google Scholar]
-
Jariyapong P, Chotwiwatthanakun C, Pooljun C, Weerachatyanukul W. Infectious hypodermal and hematopoietic necrosis virus-like particles encapsulating VP28 double-stranded RNA protect shrimp from white spot syndrome virus. Aquaculture 2019, 504, 260–266. DOI:10.1016/j.aquaculture.2019.02.001 [Google Scholar]
-
Charoonnart P, Worakajit N, Zedler JAZ, Meetam M, Robinson C, Saksmerprome V. Generation of microalga Chlamydomonas reinhardtii expressing shrimp antiviral dsRNA without supplementation of antibiotics. Sci. Rep. 2019, 9, 3164. DOI:10.1038/s41598-019-39539-x [Google Scholar]
-
Riet J, Costa-Filho J, Dall’Agno L, Medeiros L, Azevedo R, Nogueira LF, et al. Bacillus subtilis expressing double-strand RNAs (dsRNAs) induces RNA interference mechanism (RNAi) and increases survival of WSSV-challenged Litopenaeus vannamei. Aquaculture 2021, 541, 736834. DOI:10.1016/j.aquaculture.2021.736834 [Google Scholar]
-
Charoonnart P, Purton S, Saksmerprome V. Applications of microalgal biotechnology for disease control in aquaculture. Biology 2018, 7, 24. DOI:10.3390/biology7020024 [Google Scholar]
-
Saelim H, Loprasert S, Phongdara A. Bacillus subtilis expressing dsVP28 improved shrimp survival from WSSV challenge. ScienceAsia 2020, 46S, 19. DOI:10.2306/scienceasia1513-1874.2020.S003 [Google Scholar]
-
Ren X, Peng G, Peng B, Tan Y, Bai X. Robust strategy for disease resistance and increasing production breeding in red swamp crayfish (Procambarus clarkii). Fish Shellfish Immunol. 2022, 122, 57–66. DOI:10.1016/j.fsi.2022.01.032 [Google Scholar]
-
Pudgerd A, Saedan S, Santimanawong W, Weerachatyanukul W, Jariyapong P, Chaijarasphong T, et al. Genome editing of WSSV CRISPR/Cas9 and immune activation extends the survival of infected Penaeus vannamei. Sci. Rep. 2024, 14, 26306. DOI:10.1038/s41598-024-78277-7 [Google Scholar]